External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G04070 43 / 2e-05
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT1G69490 42 / 6e-05
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT1G61110 41 / 0.0001
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT3G15510 41 / 0.0001
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
AT5G07680 40 / 0.0002
NAC
ANAC079, ATNAC4, ANAC080
Arabidopsis NAC domain containing protein 79, NAC domain containing protein 80 (.1.2)
AT3G10480 40 / 0.0002
NAC
ANAC050
NAC domain containing protein 50 (.1.2.3)
AT5G14000 39 / 0.0005
NAC
ANAC084
NAC domain containing protein 84 (.1)
AT5G39820 39 / 0.0005
NAC
ANAC094
NAC domain containing protein 94 (.1)
AT5G61430 39 / 0.0006
NAC
ANAC100, ATNAC5
NAC domain containing protein 100 (.1)
AT3G29035 39 / 0.0008
NAC
ORS1, AtNAC3, ANAC059
ORE1 SISTER1, Arabidopsis NAC domain containing protein 59, NAC domain containing protein 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036959
186 / 4e-61
AT3G04070 82 / 8e-19
NAC domain containing protein 47 (.1.2)
Lus10037106
171 / 4e-55
AT3G04070 78 / 3e-17
NAC domain containing protein 47 (.1.2)
Lus10037103
79 / 4e-19
AT4G39830 121 / 2e-32
Cupredoxin superfamily protein (.1)
Lus10009924
79 / 5e-19
AT3G04070 62 / 1e-11
NAC domain containing protein 47 (.1.2)
Lus10022914
54 / 1e-09
AT1G61110 54 / 7e-09
NAC domain containing protein 25 (.1)
Lus10024907
53 / 4e-09
AT5G62380 52 / 3e-08
VASCULAR-RELATED NAC-DOMAIN 6, NAC-domain protein 101 (.1)
Lus10037939
45 / 4e-06
AT3G10490 415 / 2e-143
Arabidopsis NAC domain containing protein 51, NAC domain containing protein 52 (.1.2)
Lus10038670
45 / 5e-06
AT3G10490 441 / 5e-143
Arabidopsis NAC domain containing protein 51, NAC domain containing protein 52 (.1.2)
Lus10043095
45 / 5e-06
AT3G15510 351 / 1e-119
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10036955 pacid=23171090 polypeptide=Lus10036955 locus=Lus10036955.g ID=Lus10036955.BGIv1.0 annot-version=v1.0
ATGAAGATCAAGAACCTAGCAGTAGGGTACCGATTCAAACCATCGGACCAAGAACTAATCCTCACATACCTCTACCCGTTAGCCATCAGAGCCACCGTGA
ACCTCAACCACCCTCCTCCGCCCTTCGACGGTATAATCGAGTGCGACCTCTACTCCCCCGACGAGCCATGGAAGAGACACTTCAAGGACGAGGTCAACGA
GTCGGAGCTCTACTTCTTCACCAAGCAGAAGAAGAAGAAGAACGAGAAGGGCAAGCGAGTCGATAGGAGCTTCGGAAACTGCATGTGGAAGTCGCAGAAG
GAGGAGCCTATAGTCTACAGCACCGACGGCACACCCCCGACGGCGAGTCCACCACCATCGGTTTCGATAGGTCTTTCTCTTACAAGGTCATCAATCACGG
TGGTGGTTCGAGTTCGAGTTCTTCTTCGGAGGCTGGGGAATGGGTGA
AA sequence
>Lus10036955 pacid=23171090 polypeptide=Lus10036955 locus=Lus10036955.g ID=Lus10036955.BGIv1.0 annot-version=v1.0
MKIKNLAVGYRFKPSDQELILTYLYPLAIRATVNLNHPPPPFDGIIECDLYSPDEPWKRHFKDEVNESELYFFTKQKKKKNEKGKRVDRSFGNCMWKSQK
EEPIVYSTDGTPPTASPPPSVSIGLSLTRSSITVVVRVRVLLRRLGNG
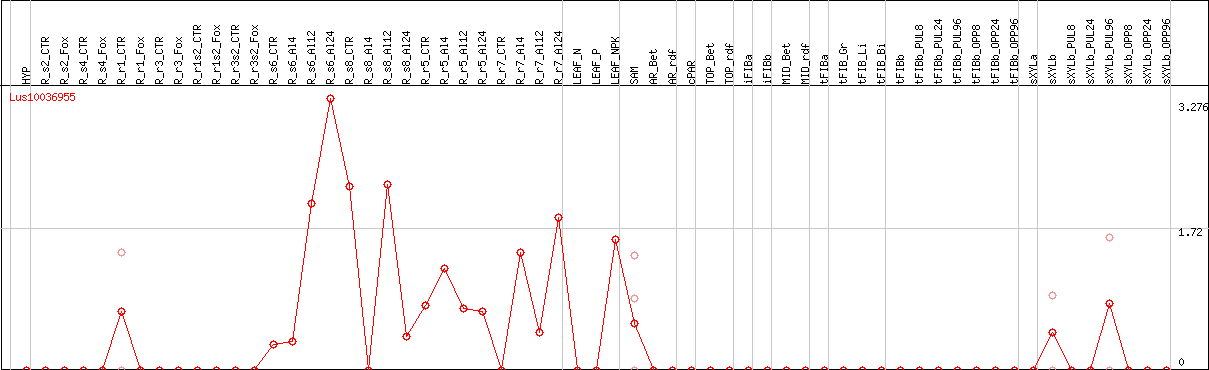
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036955 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.