External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G02960 129 / 1e-36
Heavy metal transport/detoxification superfamily protein (.1)
AT5G50740 100 / 4e-25
Heavy metal transport/detoxification superfamily protein (.1.2.3.4)
AT5G63530 91 / 2e-21
ATFP3
ARABIDOPSIS THALIANA FARNESYLATED PROTEIN 3, farnesylated protein 3 (.1.2)
AT5G24580 65 / 3e-12
Heavy metal transport/detoxification superfamily protein (.1.2.3)
AT5G03380 63 / 2e-11
Heavy metal transport/detoxification superfamily protein (.1.2)
AT5G60800 61 / 9e-11
Heavy metal transport/detoxification superfamily protein (.1.2)
AT2G28090 56 / 2e-09
Heavy metal transport/detoxification superfamily protein (.1)
AT4G35060 52 / 1e-08
HIPP25
heavy metal associated isoprenylated plant protein 25, Heavy metal transport/detoxification superfamily protein (.1)
AT2G18196 50 / 1e-07
Heavy metal transport/detoxification superfamily protein (.1)
AT4G38580 50 / 1e-07
HIPP26, ATFP6
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015762
344 / 2e-121
AT3G02960 151 / 3e-45
Heavy metal transport/detoxification superfamily protein (.1)
Lus10012867
86 / 5e-19
AT2G40770 1843 / 0.0
zinc ion binding;DNA binding;helicases;ATP binding;nucleic acid binding (.1)
Lus10038785
80 / 7e-18
AT5G50740 183 / 6e-57
Heavy metal transport/detoxification superfamily protein (.1.2.3.4)
Lus10039074
80 / 9e-18
AT5G50740 182 / 8e-57
Heavy metal transport/detoxification superfamily protein (.1.2.3.4)
Lus10030514
76 / 6e-16
AT5G50740 221 / 1e-70
Heavy metal transport/detoxification superfamily protein (.1.2.3.4)
Lus10016133
67 / 1e-12
AT5G60800 175 / 3e-52
Heavy metal transport/detoxification superfamily protein (.1.2)
Lus10021432
66 / 1e-12
AT5G60800 158 / 1e-46
Heavy metal transport/detoxification superfamily protein (.1.2)
Lus10021516
62 / 5e-11
AT5G03380 202 / 2e-61
Heavy metal transport/detoxification superfamily protein (.1.2)
Lus10015583
61 / 6e-11
AT5G24580 314 / 6e-107
Heavy metal transport/detoxification superfamily protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G017600
125 / 1e-35
AT3G02960 199 / 2e-64
Heavy metal transport/detoxification superfamily protein (.1)
Potri.016G006600
100 / 2e-25
AT5G50740 153 / 5e-45
Heavy metal transport/detoxification superfamily protein (.1.2.3.4)
Potri.006G006100
95 / 2e-23
AT5G50740 143 / 2e-41
Heavy metal transport/detoxification superfamily protein (.1.2.3.4)
Potri.015G099500
92 / 7e-22
AT5G50740 214 / 7e-68
Heavy metal transport/detoxification superfamily protein (.1.2.3.4)
Potri.012G101400
90 / 5e-21
AT5G63530 204 / 2e-63
ARABIDOPSIS THALIANA FARNESYLATED PROTEIN 3, farnesylated protein 3 (.1.2)
Potri.004G214700
77 / 2e-16
AT5G60800 142 / 7e-40
Heavy metal transport/detoxification superfamily protein (.1.2)
Potri.009G007600
75 / 9e-16
AT5G60800 157 / 1e-45
Heavy metal transport/detoxification superfamily protein (.1.2)
Potri.012G007300
59 / 4e-10
AT5G24580 251 / 5e-82
Heavy metal transport/detoxification superfamily protein (.1.2.3)
Potri.016G093100
57 / 2e-09
AT2G36950 123 / 5e-32
Heavy metal transport/detoxification superfamily protein (.1)
Potri.004G175400
50 / 9e-08
AT4G38580 254 / 2e-88
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00403
HMA
Heavy-metal-associated domain
Representative CDS sequence
>Lus10037044 pacid=23171215 polypeptide=Lus10037044 locus=Lus10037044.g ID=Lus10037044.BGIv1.0 annot-version=v1.0
ATGGCAAAGGGAGATGTCATCTTCAAGGCTCTTATTCACTGCGATGGATGCGTCACAAAGATCAATAAATCCCTATCAGGATTTGGCGGGGTGGAAGATG
TTTCGATCGAGAGGGAGAATGACATAGTAGTAGCCAACGGGATTAACCTCGACCCATTGAAGTTGCTAGCCAGGCTACAAAGCAAGTACAGTCGAAATGT
CGAGTTGATTTCACCTATACCCAAACCCAAGCCCCAAATCTCAGACAATAAACAGCCACCATCAAAGCCTGAGTCGAAGATAAAGACACTGGTTCTTAGA
GTGTTGATGTTCTGCCAAGGATGTGCAGACGACATCGAGAGAACGGTGGGGAAGATACAAGGAGTGGTGAGTGTGGAAGCAGAGATGGAGAGTTCGACGA
CAGTGGTGATTGGGTTCTTTGATGCAGAGAGATTGGTTGAGAAGGTGAGGACGCAATTAACGAAAAAAGTTGAGGTTTTGAAATTAGTTGAAGAATCATC
TAAGGGAAAGGTGAGGCTAAAGTGGAGAAACCTCCTCCTTCTGCACCTCCAGCTCCAGATGCAGTATGTTTGCTTTTGTACTGCTCATCCGCATCAACCT
TTTCTAACCAGGGGATCGGAAGGGGGCATCCATCTGTCGCTCGTTTTTAACGACGACAATGTATTTGCTTGCTCAATCATGTGA
AA sequence
>Lus10037044 pacid=23171215 polypeptide=Lus10037044 locus=Lus10037044.g ID=Lus10037044.BGIv1.0 annot-version=v1.0
MAKGDVIFKALIHCDGCVTKINKSLSGFGGVEDVSIERENDIVVANGINLDPLKLLARLQSKYSRNVELISPIPKPKPQISDNKQPPSKPESKIKTLVLR
VLMFCQGCADDIERTVGKIQGVVSVEAEMESSTTVVIGFFDAERLVEKVRTQLTKKVEVLKLVEESSKGKVRLKWRNLLLLHLQLQMQYVCFCTAHPHQP
FLTRGSEGGIHLSLVFNDDNVFACSIM
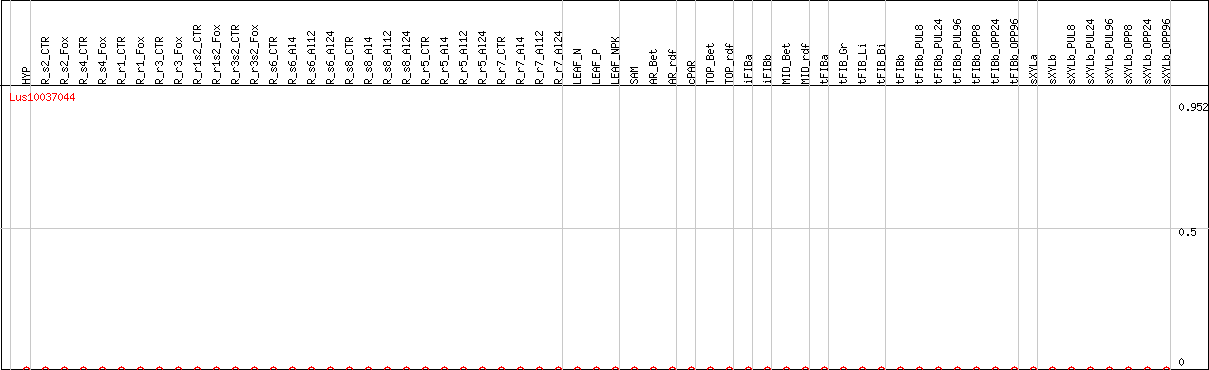
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037044 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.