External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G61240 544 / 0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
AT1G13910 478 / 2e-171
Leucine-rich repeat (LRR) family protein (.1)
AT3G11080 150 / 3e-40
AtRLP35
receptor like protein 35 (.1)
AT1G35710 147 / 4e-39
Protein kinase family protein with leucine-rich repeat domain (.1)
AT4G20140 143 / 8e-38
GSO1
GASSHO1, Leucine-rich repeat transmembrane protein kinase (.1)
AT3G12610 137 / 2e-37
DRT100
DNA-DAMAGE REPAIR/TOLERATION 100, Leucine-rich repeat (LRR) family protein (.1)
AT5G46330 140 / 6e-37
FLS2
FLAGELLIN-SENSITIVE 2, Leucine-rich receptor-like protein kinase family protein (.1)
AT4G36180 140 / 7e-37
Leucine-rich receptor-like protein kinase family protein (.1)
AT2G25790 139 / 2e-36
Leucine-rich receptor-like protein kinase family protein (.1)
AT4G08850 139 / 2e-36
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026651
595 / 0
AT5G61240 549 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10004662
578 / 0
AT5G61240 543 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10036961
561 / 0
AT5G61240 488 / 1e-175
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Lus10008712
152 / 7e-41
AT4G08850 1009 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Lus10020935
151 / 1e-40
AT4G08850 1011 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Lus10038381
143 / 7e-39
AT4G20270 457 / 5e-152
BARELY ANY MERISTEM 3, Leucine-rich receptor-like protein kinase family protein (.1)
Lus10006045
144 / 4e-38
AT2G25790 858 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Lus10036242
144 / 4e-38
AT4G20270 1248 / 0.0
BARELY ANY MERISTEM 3, Leucine-rich receptor-like protein kinase family protein (.1)
Lus10036321
142 / 2e-37
AT5G51350 957 / 0.0
MORE LATERAL GROWTH1, Leucine-rich repeat transmembrane protein kinase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G093400
553 / 0
AT5G61240 513 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Potri.010G160700
550 / 0
AT5G61240 517 / 0.0
Leucine-rich repeat (LRR) family protein (.1), Leucine-rich repeat (LRR) family protein (.2)
Potri.012G124300
158 / 6e-43
AT4G08850 727 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Potri.012G124100
155 / 8e-42
AT4G08850 711 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Potri.018G045500
152 / 7e-41
AT2G25790 925 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.006G237400
151 / 1e-40
AT2G25790 885 / 0.0
Leucine-rich receptor-like protein kinase family protein (.1)
Potri.006G014232
147 / 3e-39
AT1G53440 504 / 5e-161
Leucine-rich repeat transmembrane protein kinase (.1)
Potri.019G060400
147 / 3e-39
AT4G08850 728 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
Potri.005G117700
145 / 8e-39
AT3G24240 240 / 1e-67
Leucine-rich repeat receptor-like protein kinase family protein (.1)
Potri.001G075000
146 / 9e-39
AT4G20140 1371 / 0.0
GASSHO1, Leucine-rich repeat transmembrane protein kinase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0022
LRR
PF00560
LRR_1
Leucine Rich Repeat
Representative CDS sequence
>Lus10037108 pacid=23152428 polypeptide=Lus10037108 locus=Lus10037108.g ID=Lus10037108.BGIv1.0 annot-version=v1.0
ATGGCGCGTCATCTCCTCCTCTTCCTCTTCGTCGGTCTCCTCAACACTCTCGCCCTCTCCAAAACCCTCAAACGCGACGTGAAGGCACTGAACGAGATCA
AGTCGTCGCTGGGATGGAGAGTGGTGTACGCCTGGGTAGGAGACGACCCTTGCGGCGGCGACGGCCTCCAGCCTTGGTCCGGCGTCACCTGCTCCACTCA
GGGAGATTACAGAGTCGTCACCGAATTGGAAGTGTATGCAGTGTCGATTGTCGGTCCATTTCCTACTGCGGTGACGAATCTGCTGGATCTCACCAGATTG
GATTTTCATAACAACAAGCTGACAGGGCCGATCCCTCCTCAAATTGGGAGGTTGAAGCGTCTCAAGATACTGAATTTGAGGTGGAACAAGTTGCAAGATT
CAATTCCTCCTGAAATCGGCGAACTTAAGAGTCTGACACATTTGTATTTGAGCTTCAATAACTTCAAAGGAGAGATTCCAAGGGAGCTAGCTAATCTTCC
CGAGCTTCGGTATCTATACTTGCAAGAGAATAGGTTCTCGGGGAGAGTTCCTGCAGAATTGGGAACCTTGCAGAATCTTCGTCACTTAGATGTTGGCAAC
AATCATTTGGTGGGTACTATAAGAGAACTAATCCGGATTGAGGGTTGTTTCCCTGCGCTTCGTAACCTCTATCTGAACAATAATTATTTTACTGGAGGAG
TCCCTACGCAGCTAGCTAACTTGACAAACTTAGAAATCTTGCACCTATCCTATAACAAGATGACTGGAGTAATACCTATTGGACTTGCTTCCATCCCGAA
GTTGACATATCTGTATTTGGATCACAATCAATTCACAGGAAGAATCCCTGAAGCCTTATATAAACATCAGTTCTTGAAAGAAATGTACATCGAAGGAAGC
TCGTTCAAGCCTGGCGTAAACCCAATAGGCATGCACAAAGTCTTCGAAGTATCTGATGCAGATTTTGTAGTTTAA
AA sequence
>Lus10037108 pacid=23152428 polypeptide=Lus10037108 locus=Lus10037108.g ID=Lus10037108.BGIv1.0 annot-version=v1.0
MARHLLLFLFVGLLNTLALSKTLKRDVKALNEIKSSLGWRVVYAWVGDDPCGGDGLQPWSGVTCSTQGDYRVVTELEVYAVSIVGPFPTAVTNLLDLTRL
DFHNNKLTGPIPPQIGRLKRLKILNLRWNKLQDSIPPEIGELKSLTHLYLSFNNFKGEIPRELANLPELRYLYLQENRFSGRVPAELGTLQNLRHLDVGN
NHLVGTIRELIRIEGCFPALRNLYLNNNYFTGGVPTQLANLTNLEILHLSYNKMTGVIPIGLASIPKLTYLYLDHNQFTGRIPEALYKHQFLKEMYIEGS
SFKPGVNPIGMHKVFEVSDADFVV
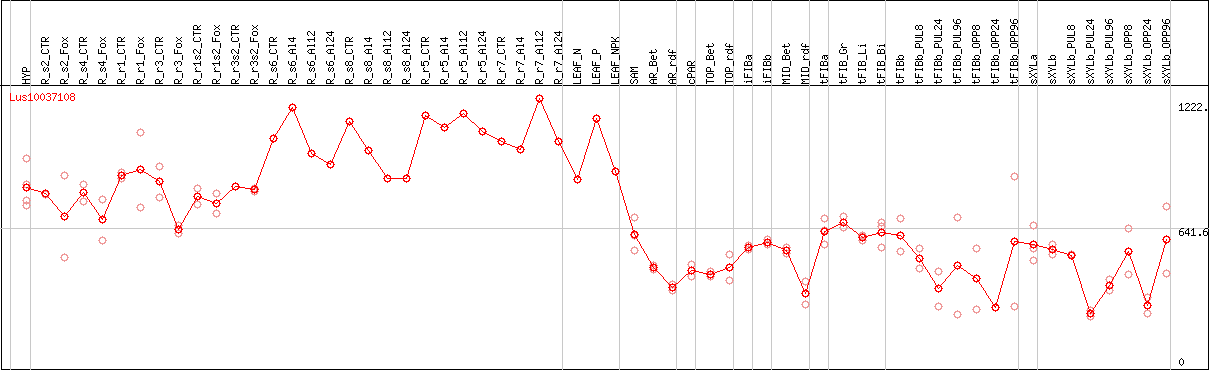
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037108 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.