External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G26940 367 / 2e-130
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT2G36130 68 / 5e-14
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT3G44600 69 / 3e-13
AtCYP71, CYP71
cyclophilin 71, cyclophilin71 (.1)
AT1G01940 64 / 9e-13
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT4G33060 63 / 2e-11
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT1G53720 60 / 3e-10
ATCYP59, CYP59
cyclophilin 59 (.1)
AT5G67530 58 / 7e-10
ATPUB49
plant U-box 49 (.1)
AT3G63400 56 / 3e-09
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
AT2G15790 56 / 6e-09
CYP40, SQN
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
AT3G62030 48 / 2e-06
CYP20-3, ROC4
cyclophilin 20-3, rotamase CYP 4 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036733
177 / 5e-57
AT1G26940 151 / 5e-47
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Lus10009237
69 / 3e-14
AT1G01940 317 / 2e-112
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Lus10020573
66 / 2e-12
AT3G44600 1115 / 0.0
cyclophilin 71, cyclophilin71 (.1)
Lus10006271
66 / 2e-12
AT3G44600 1110 / 0.0
cyclophilin 71, cyclophilin71 (.1)
Lus10018746
63 / 3e-11
AT3G63400 306 / 4e-98
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
Lus10024831
63 / 3e-11
AT3G63400 308 / 8e-99
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
Lus10021335
59 / 1e-10
AT2G36130 286 / 1e-100
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Lus10027924
60 / 2e-10
AT2G15790 534 / 0.0
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
Lus10012059
59 / 4e-10
AT2G15790 596 / 0.0
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0475
Cyclophil-like
PF00160
Pro_isomerase
Cyclophilin type peptidyl-prolyl cis-trans isomerase/CLD
Representative CDS sequence
>Lus10037197 pacid=23152484 polypeptide=Lus10037197 locus=Lus10037197.g ID=Lus10037197.BGIv1.0 annot-version=v1.0
ATGACGAGAATGTGGATCTTAGTCGTAGGTTTGGCAATTGCAGCGGCGGCCGGCGTCTCATTAGCGGAGCCTGAGCTCAGCTCAGCTCGTGTTGTCTTCC
AGACCAATTACGGGGATATCGAGTTCGGGTTCTATCCATCGGTGGCGCCGATAACGGTGGAGCACATTTTCAAGCTTGTTCGGCTTGGAGGATACAACAC
TAATCACTTCTTCAGGGTTGACAAGGGATTTGTGGCTCAAGTAGCGGATGTTGCTGGTGGGCGATCGGCTCCGATGAACGAAGAGCAAAGACTGGAGGCA
GAGAAGACTGTGATTGGTGAATTTAGTGACGTTAAGCATGTTCGAGGCATCCTGTCCATGGGGAGGTATGCTGATCCAAACAGTGGATCATCCTCGTTCT
CGATCCTTCTTGGGAATGCACCTCATCTTGACGGCCAATATGCTATATTTGGTAAAGTTACCAAAGGTGATGACACATTAACTAAGCTGGAGCAGCTTCC
TACCCGGACTGAAGGAATCTTTGTGATGCCAACAGAACGGATCACTATTTTGTCATCATATTACTACGATACTAACATGGAGAAGTGCGATGAGGAGAGG
TCGATCTTGAAACGCAGACTCGCCGCATCTGCTATTGAAGTCGAAAGACAGAGAATGAAATGCTTCCCGTGA
AA sequence
>Lus10037197 pacid=23152484 polypeptide=Lus10037197 locus=Lus10037197.g ID=Lus10037197.BGIv1.0 annot-version=v1.0
MTRMWILVVGLAIAAAAGVSLAEPELSSARVVFQTNYGDIEFGFYPSVAPITVEHIFKLVRLGGYNTNHFFRVDKGFVAQVADVAGGRSAPMNEEQRLEA
EKTVIGEFSDVKHVRGILSMGRYADPNSGSSSFSILLGNAPHLDGQYAIFGKVTKGDDTLTKLEQLPTRTEGIFVMPTERITILSSYYYDTNMEKCDEER
SILKRRLAASAIEVERQRMKCFP
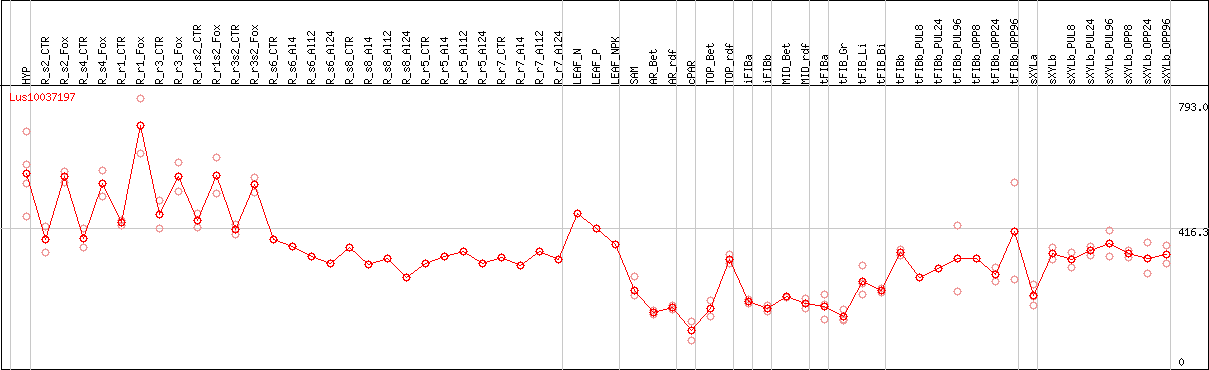
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037197 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.