External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G31790 82 / 3e-18
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G74400 62 / 3e-11
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G18520 62 / 5e-11
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G34400 60 / 2e-10
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT4G14820 58 / 1e-09
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G55740 57 / 3e-09
CRR21
chlororespiratory reduction 21, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G66500 56 / 3e-09
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G11290 56 / 5e-09
CRR22
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G27610 55 / 1e-08
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G19191 54 / 2e-08
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035697
201 / 8e-63
AT1G31790 226 / 1e-69
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10000349
87 / 1e-19
AT3G09510 49 / 5e-06
Ribonuclease H-like superfamily protein (.1)
Lus10012727
65 / 5e-12
AT5G65530 457 / 8e-157
Protein kinase superfamily protein (.1)
Lus10036375
61 / 9e-11
AT2G46050 435 / 7e-146
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10004751
59 / 7e-10
AT5G55740 577 / 0.0
chlororespiratory reduction 21, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10006174
57 / 3e-09
AT1G71460 459 / 2e-157
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10041049
56 / 4e-09
AT1G71460 796 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10006123
56 / 6e-09
AT4G14850 923 / 0.0
lovastatin insensitive 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10004987
56 / 8e-09
AT3G46790 715 / 0.0
CHLORORESPIRATORY REDUCTION 2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10037287 pacid=23152362 polypeptide=Lus10037287 locus=Lus10037287.g ID=Lus10037287.BGIv1.0 annot-version=v1.0
ATGGACGATGGTGGAAGCACTGCTCGACAAGTCCATGCAATTGCCATCAAATTACAGGGGGGATTGAATAGGAAGGTTGTTATGATCCAGTGTGGATTGA
TTAAGATGTACGGCAGGTGTGGGTTGGTCCGGTATTCAGAACAGGTATTCGACAGGACGCTCACTGACAAGCAGAGATGTAATGCGCGTTGGAATACGAT
GCTAATGTCCTATTTGCAGAATGGGTTATATGTGGAGGCGTTCAAGCTTCTTTGCAAGATGAAAGCAGCTGCAGTAGAGGTGACTGAGTCATTGGTAAAC
AAAGTCAGAATTGCCTGTGGCTCTGTAAAGGCTGAAGAACAGGGGAAACTGGTGGCCAAACCATCTGTTCTTGTTGCAGGGGAATTAGCATCCTCAATCC
GATGCCCTGTTCTTGTCATGACCGACTGCGTCTCCCTTGTGAGGCTCCTGCATGATGACTCGAGCTCTTTCCACTGGGAGGGAGTGCCTTTAACTACTAA
GATCTCGCGGAGTTTACGTACATCCTCTTATATCACGATTATGTTCCTCCCGCGCAAGGAGAATTTCAAAGTTGATTGGGTTGCTCGTAACTCCTCTGCT
GAATCTGTAGCGTTGGATTGGATTGTTATGTTACCGTCCTCTAAGGTGTAA
AA sequence
>Lus10037287 pacid=23152362 polypeptide=Lus10037287 locus=Lus10037287.g ID=Lus10037287.BGIv1.0 annot-version=v1.0
MDDGGSTARQVHAIAIKLQGGLNRKVVMIQCGLIKMYGRCGLVRYSEQVFDRTLTDKQRCNARWNTMLMSYLQNGLYVEAFKLLCKMKAAAVEVTESLVN
KVRIACGSVKAEEQGKLVAKPSVLVAGELASSIRCPVLVMTDCVSLVRLLHDDSSSFHWEGVPLTTKISRSLRTSSYITIMFLPRKENFKVDWVARNSSA
ESVALDWIVMLPSSKV
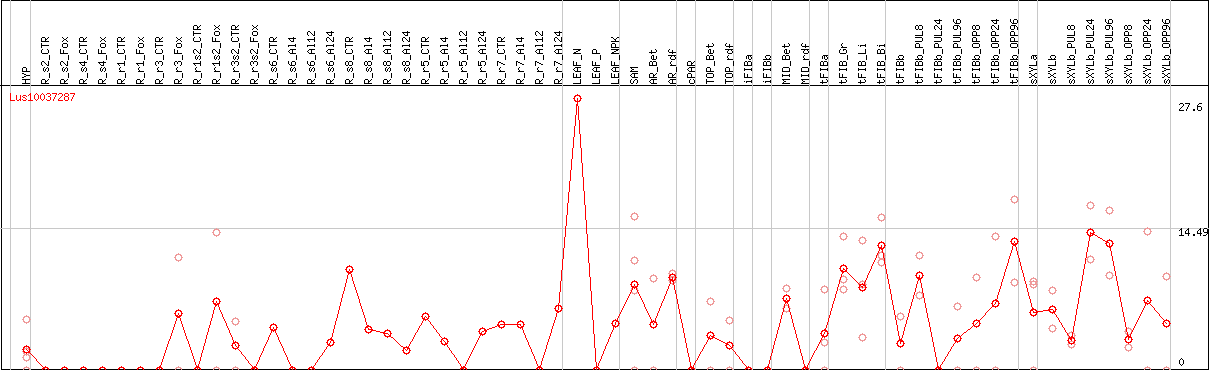
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037287 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.