Lus10037444 [FLAX]

| External link |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||
|
Arabidopsis homologues
|
| ||||||||||
|
Paralogs
|
|
||||||||||
|
Poplar homologues |
|
||||||||||
| PFAM info |
| ||||||||||
|
Representative CDS sequence |
>Lus10037444 pacid=23167271 polypeptide=Lus10037444 locus=Lus10037444.g ID=Lus10037444.BGIv1.0 annot-version=v1.0
ATGGCGACAAAGAAGCAGAAGAAGTCCTCCGCCGCCGACGGCGGTAGAAAGGAGACGGACGGCGACATCGACCTTCTGAAATCAGAAGTCGCCTCTTTTG
CTTCTACGCTAGGCCTTTCTTCTTCAGCTCCATCCTCTTCCTCCGGCTTCAGCGATGCCGATTTCCGGAAAACCGGCCGTCTCAAGCCAATTCAGAAATC
GAAACCCCAATCCGATACCAAAACCAAGCCTCCTCAGAAGCAGGACAAGAAACCCAGAAACGAGAAATCCAATACCAAACCGGAGATCAAGCCGAAGCCA
AAGCCTCCAGTTTTGACGCTGGATGATGATGATTCAGCTAACCCTGCCAGGGCGAGAGCGGCTTACAAGTACAAGGACCTCCCCAAGCTGCCGCTGATGA
AGGCAAATGCGACTGGCGTGTGGCACGTGGATGCGGAGGAATTGGAGGTGAAAGTGTTGGGAGAAGCTCACAAGAAGGTGGATGTTAAGATGAGAGTGGA
GGAGTGGAAGCAGTTTGTAGAGAAGAAGAGGGAATTGGCTGAGAGGTTGATGTGGCAGTATGTGCAGGATTATGAACAGGGGAGAGGGCAGAGTGGAGAT
ATTAAGATGCTTGTTACTACTCAGAGGTCTGGTACTGCTGCTGATAAGGTGTCAGCATTCGCGGTTCTTATTGGAGATAATCCTATGGCTAATCTGCGAT
CAGTGGATGCTCTTCTTGGAATGGTGGCATCAAAGGTGGGAAAGCGCCATGCATTGTCAGGATTTGAAGCATTGCGAGAGCTGTTTGTTGAAAGCTTGCT
ACCTGATCGGAAGCTGAAGACACTCTTACAGAGGCCTCTGAATAATCTTCCTGAGACAAAAGATGGTTACTCTCTTCTACTATTTTGGTATTGGGAGGAT
TGCCTAAAACAGAGGTATCAACGTTTTGTTTCGGCACTTGAGGAAGCATCCAAAGATATGCTACCCATATTAAAAGATAAGGCGTTGAAGAGCATGTATT
TGCTGTTGAAGAGCAAATCAGAGCAGGAGCGCAGGTTGCTCTCTTCACTAGTTAACAAACTTGGCGATCCGCAAAGTAAGGGTGCATCTAATGCTGAGTA
TTACTTGTCAAACCTGTTGTCCAATCATCCCAGTATGAAGGCCGTAGTCATTGATGAAGTGGACACCTTTCTCTTTCGGCCTCACTTGGGGATGCGAGCT
AAATACTATGCCATTAACTTTTTGAGTCAAGTTCGACTGAGCAATAGAGATGATGGACCTAAGGTGGCAAAACGATTGATAGATGTGTACTTTGCATTAT
TCAAGGTGCTAATTACTGAGGCAGGTGGTCAGAAAACGAACAAAGATGCCAAAGCTGAAGATGAAAGTACTCCTGGTGCTTCTAAGAATGAAAAAGCCAA
AGCTTCAGCAGAATCTCATATTGAGATGGATTCACGAATTCTGTCAACTCTTTTAACAGGAGTAAATAGGGCATTTCCTTACGTTTCAAGTACTGAGGCT
GACGACATCATTCAGAGTCAAACACCAATGCTCTTCCAATTGGTCCACTCCAATAACTTTAATGTAGGAGTTCAAGCCTTGATGCTCCTCGATAAAATCT
CATCCAAGAATCAGATCGTGGAAATGTTCATTGGACTTATACTCAGAGCCATGAAAAACGACGTGAACCTAAAGCGTGTTGCTGCCTTCGCAAAGCGCCT
ATTGCAAGTTGCTATGCAGCAGCCGCCTCAATATGCTTGTGGATCTCTGTTTCTGCTTTCTGAAGTGCTGAAAGCAAGGCCCCCTCTATGGAATATGGTG
CTTCAGAGTGAAGCTTTGGATGACGAAGATGAACATTTTGAGGATGTTCTGGAAGAGACCATAACCAAGGAAGATCATGGTGATATTTCAACAGATGAAC
GTAATGAGAATGGTGACCCTGAATCTGACTCTTCTGATGATGAAGAAGTCAATGCTGCCGCATCGAGTTCCGAGGATGATGATTCAGATGACGACGAGCA
ACTGATAAGAGAAGATCATTCAGAAGAGGAGATATTGATACCGAAACCTAGCGACGAGAAAGCTGAAGCATGTTCTGCAAAATCTTCCCTGCCTGGGGGA
TACGATCCTCGTCATAGGGAGCCGTCTTACTGCAATGCAGACCGTGCTAGCTGGTGGGAGCTGAGAGTGCTTGCATCACATGCACACCCTTCGGTTGCTA
CCATGGCTGGGACTCTTCTCTCTGGAGCAAATATAGTTTACAATGGAAACCCACTAAACGACCTGTCGTTGAATGCTTTCCTGGACAAGTTCATGGAGAA
GAAAGTCAAGCAGAAGGAATGGCACGGAGGCTCCCAAATCGAACCCGCAAAGAAGCTCGACATGGACAACAACCTGATCGGACCCGAGATTGTATCGTTG
GCCGAGGAGGATGTCGCTCCCGAAGATGTCGTCTTCCACAAGTTCTACACGAACAAAATGAACAGCACGAATTCGAAGAAGCAAAAGAAGAAGAAGAAGA
AGACTGCCGACGAAGAGGATGCAGAAGAGCTCTTTGATGCAGGTGACGGTGGCGCCGATGATGATGTGGCAGGAGTTGACGAGAGCGATAACGAAGAGAT
CGACAACTTGCTGGATGCGACATTAGAAGAAGACCATGACGGGGAGTACAACTACGATGACCTGGACCAGATCGCTGACGAGGATGACGATGAGCTCCTG
GCGGATGTGAGTGACACCGAAATCGGAGCATTTTCAGATATTGGCGATGATGAGAGCGATATCGAGGTAGGAGATGTGGACGATGGCGATGATGATGAGG
TGACGGTTATGGATACAAAGAAAAGGAAGCGTCCGAAGAAGATTGGTGGTGGTAAGGGTGCTTCGCCATTTGCGAGTTTCGAGGAGTACGAGCATTTGAT
GGAAGAAGGGGAAGGGGAAGGAGAGGAGGAGGAGGATAGTCCTAGTCCGAAGAAGGACAAGAAGAAGAAGCCGAAATCACGAAACAAGAAGCAGAAGAGG
AAGAAGTCATTGTCAGACTAA
|
||||||||||
|
AA sequence
|
>Lus10037444 pacid=23167271 polypeptide=Lus10037444 locus=Lus10037444.g ID=Lus10037444.BGIv1.0 annot-version=v1.0
MATKKQKKSSAADGGRKETDGDIDLLKSEVASFASTLGLSSSAPSSSSGFSDADFRKTGRLKPIQKSKPQSDTKTKPPQKQDKKPRNEKSNTKPEIKPKP
KPPVLTLDDDDSANPARARAAYKYKDLPKLPLMKANATGVWHVDAEELEVKVLGEAHKKVDVKMRVEEWKQFVEKKRELAERLMWQYVQDYEQGRGQSGD
IKMLVTTQRSGTAADKVSAFAVLIGDNPMANLRSVDALLGMVASKVGKRHALSGFEALRELFVESLLPDRKLKTLLQRPLNNLPETKDGYSLLLFWYWED
CLKQRYQRFVSALEEASKDMLPILKDKALKSMYLLLKSKSEQERRLLSSLVNKLGDPQSKGASNAEYYLSNLLSNHPSMKAVVIDEVDTFLFRPHLGMRA
KYYAINFLSQVRLSNRDDGPKVAKRLIDVYFALFKVLITEAGGQKTNKDAKAEDESTPGASKNEKAKASAESHIEMDSRILSTLLTGVNRAFPYVSSTEA
DDIIQSQTPMLFQLVHSNNFNVGVQALMLLDKISSKNQIVEMFIGLILRAMKNDVNLKRVAAFAKRLLQVAMQQPPQYACGSLFLLSEVLKARPPLWNMV
LQSEALDDEDEHFEDVLEETITKEDHGDISTDERNENGDPESDSSDDEEVNAAASSSEDDDSDDDEQLIREDHSEEEILIPKPSDEKAEACSAKSSLPGG
YDPRHREPSYCNADRASWWELRVLASHAHPSVATMAGTLLSGANIVYNGNPLNDLSLNAFLDKFMEKKVKQKEWHGGSQIEPAKKLDMDNNLIGPEIVSL
AEEDVAPEDVVFHKFYTNKMNSTNSKKQKKKKKKTADEEDAEELFDAGDGGADDDVAGVDESDNEEIDNLLDATLEEDHDGEYNYDDLDQIADEDDDELL
ADVSDTEIGAFSDIGDDESDIEVGDVDDGDDDEVTVMDTKKRKRPKKIGGGKGASPFASFEEYEHLMEEGEGEGEEEEDSPSPKKDKKKKPKSRNKKQKR
KKSLSD
|
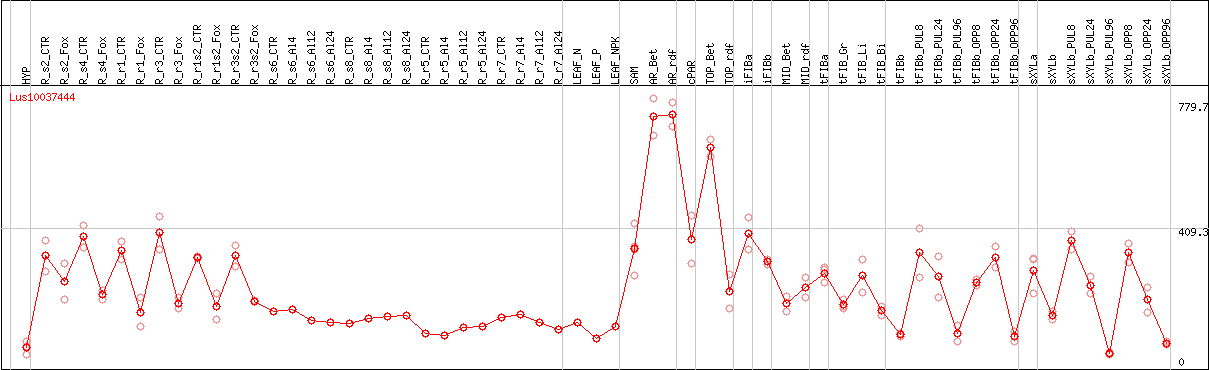
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037444 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
