Lus10037469 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10037469 pacid=23167084 polypeptide=Lus10037469 locus=Lus10037469.g ID=Lus10037469.BGIv1.0 annot-version=v1.0
ATGTCGAGTAGGGGAGGGTTTGACCAGCAGCAGCTGCCGCCAAGGCGGATTCTGCGGACTCAAACTGCTGGTAATTTGGGGGAGACCATGCTGGACAGTG
AGGTTGTGCCGTCTTCGTTGGTCGAGATTGCCCCAATTCTTCGGGTTGCTAACCAAGTAGAGCTTAGTAACCCAAGAGTTGCTTATCTATGCCGGTTCTA
TGCTTTCGAGAAAGCTCACGGACTAGATCCTACATCCAGTGGGCGTGGTGTTCGACAATTCAAAACGGCACTCTTACAACGTCTCGAAAAGGAAAATGAG
ACCACTTTACGAGGAAGGACAATGAGTGATGCTCGGGAAATGCAGAAATTTTATCGAGATTACTATAAGAAGTATATTCAAGCTTTGCAAAATGCTGCTG
ACAAGGCTGATCGTGCACAGCTTACTAAAGCATATCAAACTGCTGCAGTCCTCTTTGAAGTTCTCAAGGCTGTCAACCAGACAGAAGCTGTTCCTGAAGA
GATTCTGGAAGCTCATACCAAAGTTGAAGAGAAAAGAGAGATATATGTCCCTTACAATGTCCTCCCACTCGACCCCGACAGCAAAAATCAGGCTATCATG
AGATACCCAGAGATTCAAGCTGCTGTTTCTGCACTGAGGAATATCAGGGGATTGCCGTGGTCAAAAAATCACAAGAAGAAGGAAAATGAAGACATATTAG
ATTGGCTTCAATCTATGTTTGGGTTTCAGAAAGATAATGTGGCGAATCAAAGGGAACATTTGATTCTACTGCTTGCAAATGTGCACATGCGCCGGTTTCC
TAAGCCTGAACTACAACCAAAGTTGGACGATCATACACTAACTGAGGTGATGAAGAAATTATTCAAGAACTACAAAAGATGGTGCAAGTTTCTGGGGAGG
AAAAGTAGCCTTTGGCTGCCTACCATTCAACAAGAAGTGCAGCAAAGGAAACTACTATATATGGGTCTATATCTACTGATATGGGGAGAAGCAGCAAACT
TGAGATTCCTGCCAGAGTGCCTTTGCTACATATATCATCATATGGCTTTTGAATTGTACGGGCTACTAGCTCGGAGTGTAAGTCCATTGACAGGGGAACA
TATACAGCCAGCTTATCGTGGCGAAGAAGAGGCTTTCCTTTGGAAAGTCGTCAAGCCAATTTACGACACCATTGCAAAGGAAGCGGGGAGGAGTAAGGGT
GGAAGGTCAAAGCATTCTGAATGGAGAAACTATGATGACTTAAATGAGTACTTCTGGTCGGTCGATTGCTTTCGATTGGGTTGGCCTATGCGTGCTGATG
CCGATTTCTTTCGCCTGCCTCCTGAAGAGCTTCACCAAAACCGAGACGTGGAGAAGAAATCTGCTACCGTGAATCGCTGGATTGGCAAAGTCAATTTTGT
TGAAACGAGATCGTTTTGGCACATTTTCCGGAGCTTCGACAGACTCTGGAGTTTTTTTATATTGTGTTTACAGGCAATGATAATCATTGCTTGGAATGGA
TCGGGGAATTTGAGTTCTGTATTCGGCAGTGATGTTCTCAAGAAAGTTCTGAGCATCTTCATTACCTCTGCAATATTGAATTTCCTGCAAGCTGTTCTTG
ACGTAATTTTTAGCTTTAAGGCAAGGCAGATAATGCCATACTACGTAAAGCTGAGATATATTCTGAAGGTTATAACAGCTGCTGCATGGGTCATAATTCT
ACCAGTAACTTACGCATATAGTTGGAACGATCCTACTGGATTCGGAAGAACCATTCGGGGCTGGTTCGGCAACAGCCCTAGCACGCCTTCTTTGTTCATT
ATGGCTCCTCGACTTTATGTCGGAAGGGGGATGCATGAGAGCTCTATTTCACTTTTCAAATATACTCTGTTCTGGGTTCTTCTCTTGGTGTCAAAATTGG
CGTTCAGTTATTATATCGAGATTAAGCCTCTTGTCGGTCCGACGAAAGCCATAATGAGTGTTCACATAAGAACCTACCAATGGCATGAATTCTTTCCCCA
AGCAAAGAACAACATTGGCGTTGTGATTGCGCTTTGGGCTCCTATTGTTCTTGTATATTTCATGGATACCCAGATTTGGTATGCTATTTACTCCACTATT
TTCGGTGGTTTATATGGTGCTTTTCGCCGCCTTGGAGAGATTCGAACATTAGGAATGCTGAGGTCTCGGTTCCAATCACTACCTGGTGCTTTCAATGCCT
GCTTGGTGCCTGTAGAAAAAAGCGAGAGTAACAAAAAGAAGGGGTTGAAAGCTACTTTTTCACGTAGATTCCAAGAGAGCCCCTCCAGCAAAGAAAAGCA
GGAAGCAAGGTTTGCTCAAATGTGGAATAAAATCATAAGTAGCTTCAGGGAGGAGGATCTGATAAATGACAGGGAGATGAATTTGATGCTTGTTCCTTAC
TGGGCCGACCGTGAACTAGAACTCATTCAATGGCCTCCATTTTTACTTGCTAGCAAGATCCCTATAGCAGTCGATATGGCTAAAGATAGCAACGGGAAAG
ACCGCGAATTGAAGAAGCGATTGGTTTCAGATAACTATATGTTGTGTGCAGTTCGTGAATGCTACGCTTCGATTAAAAGTATCGCCAATTATTTGATTCT
AGGAGAGCGCGAAATATTGGTAATAAATGAAATCTTTTCCAAAGTTGACGAGTACATAGACAATGAAACCATTATAAAAGAACTGAATATGAGCGCTCTT
CCGATCCTCAACGAACAATTCGTGAAGCTCGTCGAGTATTTGTTGGAGAATAAGAGAGAGGACAAGGATCAAGTTGTTATTCTGTTGCTTGACATGCTGG
AAGTTGTGACAAGAGACATATTGGAGGATGAGGTTCCCAGTTTGTTGGAATCAAGCCATGGGGGATTCTATGGGAAACAAGATGGAATGATCTCACTTGA
TCAGCATCAAAAGCATCAAATCTTTGGCGAGCTTAGGTTTCCTGTTCCTCAATCGGATGCTTGGAACGAGAAAATCAGAAGACTTCATCTATTACTCACA
GTCAAGGAATCTGCCATGGATGTTCCATCAAACTTGGAAGCTAGGAGGCGCATTTCTTTCTTCAGCAATTCATTGTTCATGGACATGCCTGATGCCCCCA
AGGTTCGCAGCATGCTTTCATTCTGTGTTCTCACTCCTTATTACACGGAGGATGTACTTTATTCGCTTAATACCCTCGAAAAACCGAACGAAGATGGTGT
TTCCGTCCTCTTCTACTTGCAGAAAATTTTCCCAGATGAATGGACAAATTTTCTTCAAAGGGTTGGATGCAGTAGTGAAGAGGAACTTAGATCGACCGAA
GAGCTCGAAGAAGAGCTACGCCTATGGGCATCATACAGAGGCCAAACGCTGACAAAAACAGTGCGTGGCATGATGTATTACCGGAAAGCATTGGAACTGC
AGGCCTTCCTCGACCTGGCAACAGACGAAGAATTGATGAAGGGATATAAGGCTGCTGAAGCTAACAGTGAGCAACATTCGAAAAGGCAGAGGTCTTTGTG
GGCACAATGCCAAGCAATAACTGATATGAAGTTCACTTATGTCGTCTCCTGCCAGAACTATGGTATTCATAAGCGATCCGGTGATACTCGTGCTAACGAC
ATTTTGAGGCTCATGACTACGCATCCATCGCTGCGTGTAGCTTATATCGACGAGGTTGAAGAGACGAGCAAAGATAGAGCGAAAAGATCTGTGGAGAAGG
TGTATTATTCAGCTCTTGTAAAAGCTGCTCCTCCGACCACTCCTATTGATTCTTCCGAGCGTGTTCAAAATCTCGATCAGGTAATTTACCGTATAAAGCT
ACCTGGACCGGCATTGTTGGGTGAGGGCAAGCCGGAAAATCAGAACCATGCAATCATTTTCACACGTGGGGAAGGTTTGCAAACAATAGATATGAATCAG
GATAACTACATGGAAGAAGCATTCAAAATGAGGAACTTGCTAGAAGAATTTCTAGTAAAGCATGGTGGTGTTAGATGCCCTACAATTTTAGGCCTTAGGG
AGCATATCTTTACTGGCAGTGTTTCCTCTCTTGCCTGGTTCATGTCGAATCAAGAGAACAGTTTCGTGACCATCGGGCAGCGATTGTTGGCTCATCCTTT
GAAAGTGAGATTCCATTATGGTCACCCTGATGTGTTCGATAGACTATTTCACCTTACTAGAGGGGGAATCAGCAAAGCTTCGAGGGGTATCAATTTAAGT
GAGGACATATTTGCAGGATTCAATTCCACGTTGCGAGAAGGCAATGTCACTCACCACGAATATATTCAAGTCGGTAAAGGGAGAGATGTGGGCCTTAACC
AGATATCCATGTTCGAGGCGAAGATTGCTAACGGGAACGGCGAACAGACACTGAGTCGTGACATTTACAGGCTTGGACATCGATTCGACTTTTTCCGAAT
GTTGTCTTGCTACTTCACTACAATTGGCTTCTATTTCAGTACCTTGATCACTGTGCTCACGGTCTATGTTTTCCTCTATGGCCGTCTTTATCTAGCCCTT
AGCGGACTGGAAGAAGGGCTGATTAACCAACGAGCTATACGGGATAACAAGCCTCTTCAAGTAGCTCTTGCTTCGCAGTCCGTCGTACAGATAGGGATTT
TGATGGCTTTACCTATGATGATGGAAATCGGTCTAGAAAGGGGGTTTCGAAACGCGCTGAGCGATTTCGTTCTGATGCAATTGCAACTTGCACCTTTATT
CTTCACATTCTCCCTCGGCACGAAGACTCACTACTACGGTAGAACGTTGCTCCACGGAGGGGCGGAATACAGAGGAACTGGTCGTGGGTTCGTAGTGTTC
CATGCAAAGTTTGCCGAGAACTACCGGCTCTACTCTCCCATCCACTTCCTAACAGGCATCGAACTGATGATCTTGCTTCTCGTGTACCATATTTTCGGCC
ACTCGTACCGAGGCGTACTTGCAGGAGTCTTGATCACGATTTCGATATGGTTCATGGTTGTGACGTGGCTCTTTGCACCGTTCCTGTTCAATCCGTCCGG
GTTCGAGTGGCAGAAAATACTCGATGACTGGACAGATTGGCATAAGTGGATAAATAGCCGCGGAGGTATCGGGGTGCCTCCTGAAAAGAGCTGGGAGTCG
TGGTGGGAGAGCGAGCATGCTCATCTCCGCCATTCGGGAATCAGAGGCATTATCGCGGAGATATTGTTAGCGTTGCGGTTTTTCATCTTTCAGTATGGCC
TCGTGTATCATCTCAGCATAATCAACAAAGCCAAAACATTTCTGGTATATGGCGTTTCGTGGCTTGTGATCGCACTGATACTGTCTATAGTGAAGGCCAT
ATCAGTTGGACGAAGACGACTGAGCGCGAATTTCCAGCTCGTGTTTCGACTGATCAAAGGGCTCATATTCCTGACGTTCCTTGCCACATTCGCTACTTTG
ATAGCGGTCCTCCATATGACACTTCAGGATGTAGTAGTGTGCATTCTAGCATTCATGCCGACCGGATGGGGACTTCTACTGATAGCACAGGCCTGCAAGC
CTGTAATACAACGAGCTGGATTCTGGGGATCAGTGCGAGTTCTGGCTCGCGGGTACGAGATAGTGATGGGGTTGCTTCTATTCATTCCAGTCGCGTTCTT
GGCATGGTTCCCGTTTGTATCCGAGTTCCAGACGCGTATGCTGTTCAACCAAGCATTCAGCAGAGGTCTGCAGATCTCGCGTATCCTGGGCGGACCGAGG
AAAGATCGCTCTACCAGAAGCAACAAGGAATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10037469 pacid=23167084 polypeptide=Lus10037469 locus=Lus10037469.g ID=Lus10037469.BGIv1.0 annot-version=v1.0
MSSRGGFDQQQLPPRRILRTQTAGNLGETMLDSEVVPSSLVEIAPILRVANQVELSNPRVAYLCRFYAFEKAHGLDPTSSGRGVRQFKTALLQRLEKENE
TTLRGRTMSDAREMQKFYRDYYKKYIQALQNAADKADRAQLTKAYQTAAVLFEVLKAVNQTEAVPEEILEAHTKVEEKREIYVPYNVLPLDPDSKNQAIM
RYPEIQAAVSALRNIRGLPWSKNHKKKENEDILDWLQSMFGFQKDNVANQREHLILLLANVHMRRFPKPELQPKLDDHTLTEVMKKLFKNYKRWCKFLGR
KSSLWLPTIQQEVQQRKLLYMGLYLLIWGEAANLRFLPECLCYIYHHMAFELYGLLARSVSPLTGEHIQPAYRGEEEAFLWKVVKPIYDTIAKEAGRSKG
GRSKHSEWRNYDDLNEYFWSVDCFRLGWPMRADADFFRLPPEELHQNRDVEKKSATVNRWIGKVNFVETRSFWHIFRSFDRLWSFFILCLQAMIIIAWNG
SGNLSSVFGSDVLKKVLSIFITSAILNFLQAVLDVIFSFKARQIMPYYVKLRYILKVITAAAWVIILPVTYAYSWNDPTGFGRTIRGWFGNSPSTPSLFI
MAPRLYVGRGMHESSISLFKYTLFWVLLLVSKLAFSYYIEIKPLVGPTKAIMSVHIRTYQWHEFFPQAKNNIGVVIALWAPIVLVYFMDTQIWYAIYSTI
FGGLYGAFRRLGEIRTLGMLRSRFQSLPGAFNACLVPVEKSESNKKKGLKATFSRRFQESPSSKEKQEARFAQMWNKIISSFREEDLINDREMNLMLVPY
WADRELELIQWPPFLLASKIPIAVDMAKDSNGKDRELKKRLVSDNYMLCAVRECYASIKSIANYLILGEREILVINEIFSKVDEYIDNETIIKELNMSAL
PILNEQFVKLVEYLLENKREDKDQVVILLLDMLEVVTRDILEDEVPSLLESSHGGFYGKQDGMISLDQHQKHQIFGELRFPVPQSDAWNEKIRRLHLLLT
VKESAMDVPSNLEARRRISFFSNSLFMDMPDAPKVRSMLSFCVLTPYYTEDVLYSLNTLEKPNEDGVSVLFYLQKIFPDEWTNFLQRVGCSSEEELRSTE
ELEEELRLWASYRGQTLTKTVRGMMYYRKALELQAFLDLATDEELMKGYKAAEANSEQHSKRQRSLWAQCQAITDMKFTYVVSCQNYGIHKRSGDTRAND
ILRLMTTHPSLRVAYIDEVEETSKDRAKRSVEKVYYSALVKAAPPTTPIDSSERVQNLDQVIYRIKLPGPALLGEGKPENQNHAIIFTRGEGLQTIDMNQ
DNYMEEAFKMRNLLEEFLVKHGGVRCPTILGLREHIFTGSVSSLAWFMSNQENSFVTIGQRLLAHPLKVRFHYGHPDVFDRLFHLTRGGISKASRGINLS
EDIFAGFNSTLREGNVTHHEYIQVGKGRDVGLNQISMFEAKIANGNGEQTLSRDIYRLGHRFDFFRMLSCYFTTIGFYFSTLITVLTVYVFLYGRLYLAL
SGLEEGLINQRAIRDNKPLQVALASQSVVQIGILMALPMMMEIGLERGFRNALSDFVLMQLQLAPLFFTFSLGTKTHYYGRTLLHGGAEYRGTGRGFVVF
HAKFAENYRLYSPIHFLTGIELMILLLVYHIFGHSYRGVLAGVLITISIWFMVVTWLFAPFLFNPSGFEWQKILDDWTDWHKWINSRGGIGVPPEKSWES
WWESEHAHLRHSGIRGIIAEILLALRFFIFQYGLVYHLSIINKAKTFLVYGVSWLVIALILSIVKAISVGRRRLSANFQLVFRLIKGLIFLTFLATFATL
IAVLHMTLQDVVVCILAFMPTGWGLLLIAQACKPVIQRAGFWGSVRVLARGYEIVMGLLLFIPVAFLAWFPFVSEFQTRMLFNQAFSRGLQISRILGGPR
KDRSTRSNKE
|
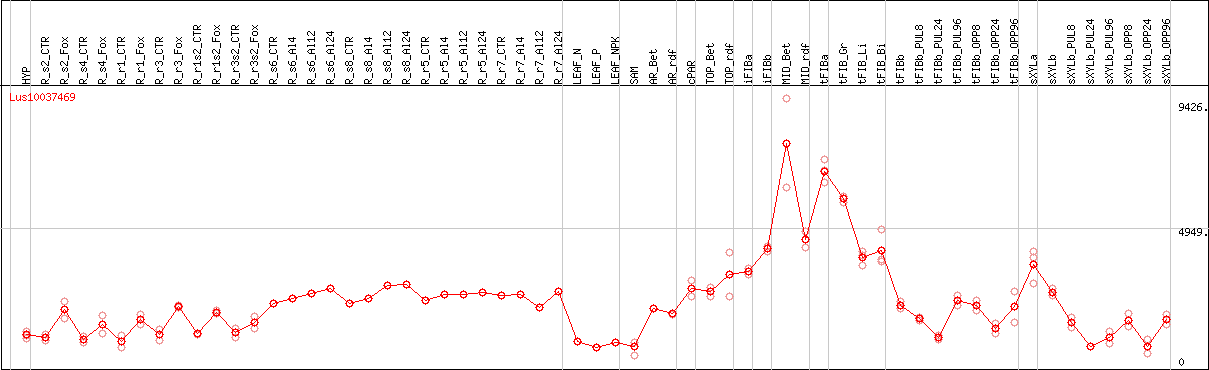
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037469 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
