External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G02500 449 / 2e-159
ATXT2, XXT2
XYG XYLOSYLTRANSFERASE 2, ARABIDOPSIS THALIANA UDP-XYLOSYLTRANSFERASE 2, UDP-xylosyltransferase 2 (.1)
AT3G62720 429 / 2e-151
ATXT1, XXT1
XYG XYLOSYLTRANSFERASE 1, xylosyltransferase 1 (.1.2)
AT1G74380 321 / 4e-109
XXT5
xyloglucan xylosyltransferase 5 (.1)
AT1G18690 320 / 6e-108
Galactosyl transferase GMA12/MNN10 family protein (.1)
AT5G07720 317 / 2e-107
Galactosyl transferase GMA12/MNN10 family protein (.1)
AT4G37690 157 / 7e-46
Galactosyl transferase GMA12/MNN10 family protein (.1)
AT2G22900 155 / 7e-45
Galactosyl transferase GMA12/MNN10 family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037519
473 / 6e-169
AT4G02500 786 / 0.0
XYG XYLOSYLTRANSFERASE 2, ARABIDOPSIS THALIANA UDP-XYLOSYLTRANSFERASE 2, UDP-xylosyltransferase 2 (.1)
Lus10002870
466 / 5e-166
AT4G02500 779 / 0.0
XYG XYLOSYLTRANSFERASE 2, ARABIDOPSIS THALIANA UDP-XYLOSYLTRANSFERASE 2, UDP-xylosyltransferase 2 (.1)
Lus10037516
465 / 7e-166
AT4G02500 786 / 0.0
XYG XYLOSYLTRANSFERASE 2, ARABIDOPSIS THALIANA UDP-XYLOSYLTRANSFERASE 2, UDP-xylosyltransferase 2 (.1)
Lus10019076
321 / 2e-109
AT5G07720 744 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Lus10021631
320 / 2e-108
AT5G07720 766 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Lus10015721
318 / 2e-108
AT5G07720 741 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Lus10000563
319 / 5e-108
AT5G07720 759 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Lus10037514
279 / 2e-93
AT3G62720 615 / 0.0
XYG XYLOSYLTRANSFERASE 1, xylosyltransferase 1 (.1.2)
Lus10011550
163 / 8e-48
AT2G22900 571 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G025300
446 / 2e-158
AT4G02500 775 / 0.0
XYG XYLOSYLTRANSFERASE 2, ARABIDOPSIS THALIANA UDP-XYLOSYLTRANSFERASE 2, UDP-xylosyltransferase 2 (.1)
Potri.008G208000
446 / 2e-158
AT4G02500 773 / 0.0
XYG XYLOSYLTRANSFERASE 2, ARABIDOPSIS THALIANA UDP-XYLOSYLTRANSFERASE 2, UDP-xylosyltransferase 2 (.1)
Potri.010G025100
356 / 2e-123
AT3G62720 674 / 0.0
XYG XYLOSYLTRANSFERASE 1, xylosyltransferase 1 (.1.2)
Potri.012G068300
325 / 7e-111
AT5G07720 755 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Potri.015G061800
322 / 3e-109
AT5G07720 730 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Potri.007G005900
151 / 2e-43
AT2G22900 593 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Potri.014G006100
147 / 6e-42
AT2G22900 592 / 0.0
Galactosyl transferase GMA12/MNN10 family protein (.1)
Potri.010G025400
68 / 6e-15
AT4G02500 68 / 8e-15
XYG XYLOSYLTRANSFERASE 2, ARABIDOPSIS THALIANA UDP-XYLOSYLTRANSFERASE 2, UDP-xylosyltransferase 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF05637
Glyco_transf_34
galactosyl transferase GMA12/MNN10 family
Representative CDS sequence
>Lus10037520 pacid=23167258 polypeptide=Lus10037520 locus=Lus10037520.g ID=Lus10037520.BGIv1.0 annot-version=v1.0
ATGGCATTTGAATTGCCTTGGGAGAGGTATAAGGACGTCAATCTGGTGATGCACGGTTGGAACGAGATGGTTTACGATCAGAAGAACTGGATTGGGCTGA
ACACTGGAAGCTTTCTGCTGAGGAACAGCCAATGGGCATTGGATCTTCTGGACGCATGGGCGCCAATGGGACCAAAAGGGAAGATTAGAGACGAAGCTGG
GAAAGTACTCACTCGGGAGCTGAAAGACAGACCTGTTTTCGAAGCGGACGATCAATCTGCCATGGTTTATCTATTGGCTACACAGAGGGAAATGTGGGGG
GAGAAGGTGTATCTAGAGAGCGCTTACTATTTGCACGGTTACTGGGGGATATTGGTGGACCGGTATGAGGAAATGATCGAGAATTATCATCCGGGACTGG
GCGATCATCGGTGGCCGTTGGTTACTCATTTCGTGGGTTGCAAACCTTGCGGGAAGTTCGGTGATTACCCAGTGGAAAGATGCTTGAAGCAAATGGATCG
AGCTTTCAACTTCGGTGACAATCAAATTCTGCAGATTTATGGGTTCACACACAAAACTTTGGCTAGTAGGAGAGTTAAGCGGGTTAGAAACGAGACGAGC
AATCCACTGGAAGCTAAAGACGAGCTTGGGTTGCTTCATCCTGCTTTCAAAGCTGTCAAAGTATCAACGACATCGTCTTGA
AA sequence
>Lus10037520 pacid=23167258 polypeptide=Lus10037520 locus=Lus10037520.g ID=Lus10037520.BGIv1.0 annot-version=v1.0
MAFELPWERYKDVNLVMHGWNEMVYDQKNWIGLNTGSFLLRNSQWALDLLDAWAPMGPKGKIRDEAGKVLTRELKDRPVFEADDQSAMVYLLATQREMWG
EKVYLESAYYLHGYWGILVDRYEEMIENYHPGLGDHRWPLVTHFVGCKPCGKFGDYPVERCLKQMDRAFNFGDNQILQIYGFTHKTLASRRVKRVRNETS
NPLEAKDELGLLHPAFKAVKVSTTSS
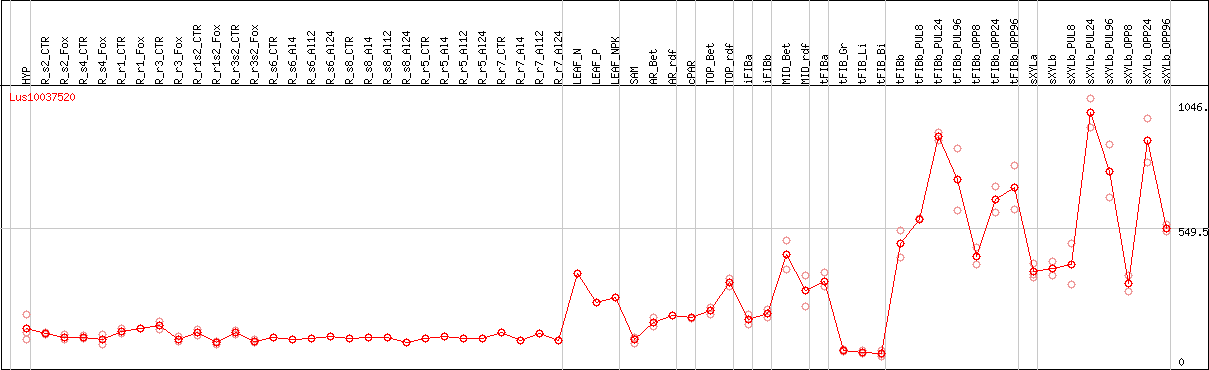
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037520 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.