External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G17420 241 / 2e-80
NTR2, ATNTRA, NTRA
NADPH-DEPENDENT THIOREDOXIN REDUCTASE 2, NADPH-dependent thioredoxin reductase A (.1)
AT4G35460 236 / 1e-78
ATNTRB, TRB1, NTR1
NADPH-DEPENDENT THIOREDOXIN REDUCTASE 1, NADPH-dependent thioredoxin reductase B (.1)
AT2G41680 139 / 7e-40
NTRC
NADPH-dependent thioredoxin reductase C (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006853
285 / 2e-92
AT4G33270 729 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10038256
256 / 7e-88
AT2G17420 442 / 1e-157
NADPH-DEPENDENT THIOREDOXIN REDUCTASE 2, NADPH-dependent thioredoxin reductase A (.1)
Lus10025845
255 / 5e-87
AT2G17420 437 / 7e-155
NADPH-DEPENDENT THIOREDOXIN REDUCTASE 2, NADPH-dependent thioredoxin reductase A (.1)
Lus10018051
129 / 3e-36
AT2G41680 845 / 0.0
NADPH-dependent thioredoxin reductase C (.1)
Lus10042048
129 / 8e-36
AT2G41680 840 / 0.0
NADPH-dependent thioredoxin reductase C (.1)
Lus10019550
99 / 2e-27
AT2G17420 202 / 5e-65
NADPH-DEPENDENT THIOREDOXIN REDUCTASE 2, NADPH-dependent thioredoxin reductase A (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G456800
254 / 2e-85
AT2G17420 582 / 0.0
NADPH-DEPENDENT THIOREDOXIN REDUCTASE 2, NADPH-dependent thioredoxin reductase A (.1)
Potri.011G145100
251 / 1e-84
AT2G17420 568 / 0.0
NADPH-DEPENDENT THIOREDOXIN REDUCTASE 2, NADPH-dependent thioredoxin reductase A (.1)
Potri.006G049100
130 / 1e-36
AT2G41680 785 / 0.0
NADPH-dependent thioredoxin reductase C (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF00070
Pyr_redox
Pyridine nucleotide-disulphide oxidoreductase
Representative CDS sequence
>Lus10037596 pacid=23167094 polypeptide=Lus10037596 locus=Lus10037596.g ID=Lus10037596.BGIv1.0 annot-version=v1.0
ATGGAGGAGGCGAATTTCTTGACGAAGTACGGGAGCAAGGTTTACATCATTCATCGGAGGGACACCTTTAGGGCATCGAAGATTATGCAGAACAGGGCGA
TGAAGAACCCTAAGATTGAGGTGATTTGGAATTCGGTAACGGAGGAGGCCTATGGTGAGAAGTCGCTAAAAGGATTGAAGGTGAAGAACATTGTTAGCGG
TGAGGTTTCTGATTTGACAGTTTCCGGGTTGTTCTTCGCCATTGGACATGAGCCTGCTACTAAGTTCTTGGATGGGCAGCTTGAGGTTCACTCTGATGGA
TACGTTGTGACGAAGCCTGGAACTACGCAGACTAGTGTTAGAGGTGTTTTTGCTGCTGGTGATGTTCAGGATAAGAAGTATAGGCAGGCCATTACTGCTG
CTGGCACTGGTTAG
AA sequence
>Lus10037596 pacid=23167094 polypeptide=Lus10037596 locus=Lus10037596.g ID=Lus10037596.BGIv1.0 annot-version=v1.0
MEEANFLTKYGSKVYIIHRRDTFRASKIMQNRAMKNPKIEVIWNSVTEEAYGEKSLKGLKVKNIVSGEVSDLTVSGLFFAIGHEPATKFLDGQLEVHSDG
YVVTKPGTTQTSVRGVFAAGDVQDKKYRQAITAAGTG
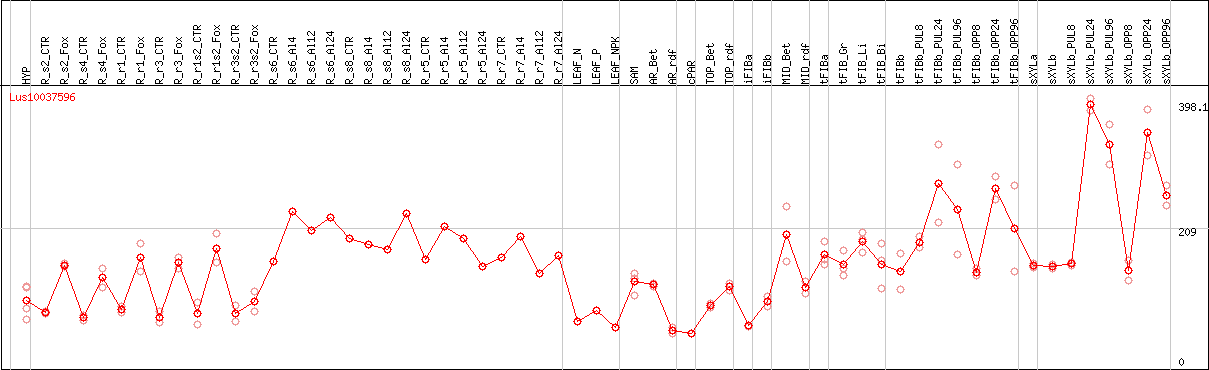
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037596 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.