External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G63980 421 / 4e-148
SUPO1, RON1, ALX8, HOS2, FRY1, ATSAL1, SAL1
suppressors of PIN1 overexpression 1, ROTUNDA 1, HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 2, FIERY1, ALTERED EXPRESSION OF APX2 8, Inositol monophosphatase family protein (.1)
AT5G64000 326 / 1e-111
ATSAL2, SAL2
Inositol monophosphatase family protein (.1)
AT5G09290 326 / 1e-111
Inositol monophosphatase family protein (.1)
AT5G63990 320 / 4e-109
Inositol monophosphatase family protein (.1.2)
AT5G54390 206 / 2e-64
ATAHL, AHL
HAL2-like (.1)
AT4G05090 110 / 9e-28
Inositol monophosphatase family protein (.1)
AT1G31190 54 / 4e-08
IMPL1
myo-inositol monophosphatase like 1 (.1)
AT4G39120 47 / 9e-06
HISN7, IMPL2
HISTIDINE BIOSYNTHESIS 7, myo-inositol monophosphatase like 2 (.1)
AT3G02870 46 / 1e-05
VTC4
Inositol monophosphatase family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015663
498 / 1e-178
AT5G63980 454 / 2e-159
suppressors of PIN1 overexpression 1, ROTUNDA 1, HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 2, FIERY1, ALTERED EXPRESSION OF APX2 8, Inositol monophosphatase family protein (.1)
Lus10037888
218 / 7e-69
AT5G54390 497 / 6e-177
HAL2-like (.1)
Lus10038597
218 / 8e-69
AT5G54390 505 / 2e-180
HAL2-like (.1)
Lus10018453
192 / 8e-58
AT5G54390 442 / 2e-154
HAL2-like (.1)
Lus10011231
190 / 3e-57
AT5G54390 447 / 4e-156
HAL2-like (.1)
Lus10007817
118 / 2e-30
AT4G05090 436 / 4e-152
Inositol monophosphatase family protein (.1)
Lus10006739
117 / 3e-30
AT4G05090 469 / 8e-166
Inositol monophosphatase family protein (.1)
Lus10041968
48 / 5e-06
AT4G39120 447 / 6e-158
HISTIDINE BIOSYNTHESIS 7, myo-inositol monophosphatase like 2 (.1)
Lus10017976
46 / 2e-05
AT4G39120 429 / 9e-151
HISTIDINE BIOSYNTHESIS 7, myo-inositol monophosphatase like 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G105100
433 / 6e-153
AT5G63980 535 / 0.0
suppressors of PIN1 overexpression 1, ROTUNDA 1, HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 2, FIERY1, ALTERED EXPRESSION OF APX2 8, Inositol monophosphatase family protein (.1)
Potri.005G063900
426 / 3e-150
AT5G63980 529 / 0.0
suppressors of PIN1 overexpression 1, ROTUNDA 1, HIGH EXPRESSION OF OSMOTICALLY RESPONSIVE GENES 2, FIERY1, ALTERED EXPRESSION OF APX2 8, Inositol monophosphatase family protein (.1)
Potri.011G124700
215 / 1e-67
AT5G54390 489 / 4e-174
HAL2-like (.1)
Potri.011G044900
215 / 7e-67
AT5G54390 502 / 2e-177
HAL2-like (.1)
Potri.004G036400
211 / 3e-65
AT5G54390 497 / 7e-176
HAL2-like (.1)
Potri.004G033200
109 / 2e-27
AT4G05090 522 / 0.0
Inositol monophosphatase family protein (.1)
Potri.010G156500
49 / 1e-06
AT3G02870 421 / 7e-151
Inositol monophosphatase family protein (.1.2.3)
Potri.010G156300
46 / 1e-05
AT3G02870 438 / 4e-157
Inositol monophosphatase family protein (.1.2.3)
Potri.009G120600
46 / 1e-05
AT4G39120 466 / 2e-165
HISTIDINE BIOSYNTHESIS 7, myo-inositol monophosphatase like 2 (.1)
Potri.006G014900
44 / 6e-05
AT3G02870 442 / 8e-159
Inositol monophosphatase family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0171
Phospoesterase
PF00459
Inositol_P
Inositol monophosphatase family
Representative CDS sequence
>Lus10037677 pacid=23167176 polypeptide=Lus10037677 locus=Lus10037677.g ID=Lus10037677.BGIv1.0 annot-version=v1.0
ATGCAGTCAGATGTTCAATCCAAATCAGATAAAAGCCCTGTTACTGTGGCTGACTATGGATCACAAGCTGTTGTTAGTCTTGTTCTACAGAGGGAGCTCC
CGTCTGTTCCGTTCTCGTTAGTCGCTGAGGAGGACTCGGGCGATCTTCGGAATGATAGTGCCCGAGACATCACAGCCCGCATTACGGAACTTGTTAATGA
TACCTTACGCACTGACGGCTCTTCTTCAACTCTGTCAGTGGAACAAGTGCTGAAGGCCATCGATTGTGGTATATCTGAAGGAGGCTCTACTGGAAGGCAT
TGGGTTTTGGACCCTATTGATGGTACTAAAGGGTTTGTAAGAGGAGAACAGTATGCAATAGCATTGGCTCTGCTGGATGAAGGGAAAGTTGTCTTGGTCG
GTGCTGGAACTTATATGCAGTCGTTAGAAGGTGATTCATTAGTGCAAGTGAATGTTACTGCTACTGAGAACCCAGAAGAAGCGTCCTTCTTCGAGTCATA
TGCAAAAGCACACTCCTTGCATGGCTTAACCAGCTCTATTGCTGAGAAACTTGGTGTTAAGGCACCACCAGTTAGATTTGATAGCCAGGCGAAATATGGG
GCTCTTTCGAGGGGTGATGGAGCCATATACATGCGTTTTCCCCACCAAGGATACCGCGAGAAAATATGGGATCATGCTGCTGGTTGCATTGTCGTGACAG
AAGCCGGTGGACTTGTAACTGATGCCGCGGGGAATGAATTGGATTTTTCGAAGGGAAGATACTTGGATGTGGACACAGGGATCATTTCCACTAATCCCAA
ATTGATGCCATTGCTCCTCAAGGCCATCAGTGAAGCTAAGGACAGCAGTACTTCTTCCCTCTGA
AA sequence
>Lus10037677 pacid=23167176 polypeptide=Lus10037677 locus=Lus10037677.g ID=Lus10037677.BGIv1.0 annot-version=v1.0
MQSDVQSKSDKSPVTVADYGSQAVVSLVLQRELPSVPFSLVAEEDSGDLRNDSARDITARITELVNDTLRTDGSSSTLSVEQVLKAIDCGISEGGSTGRH
WVLDPIDGTKGFVRGEQYAIALALLDEGKVVLVGAGTYMQSLEGDSLVQVNVTATENPEEASFFESYAKAHSLHGLTSSIAEKLGVKAPPVRFDSQAKYG
ALSRGDGAIYMRFPHQGYREKIWDHAAGCIVVTEAGGLVTDAAGNELDFSKGRYLDVDTGIISTNPKLMPLLLKAISEAKDSSTSSL
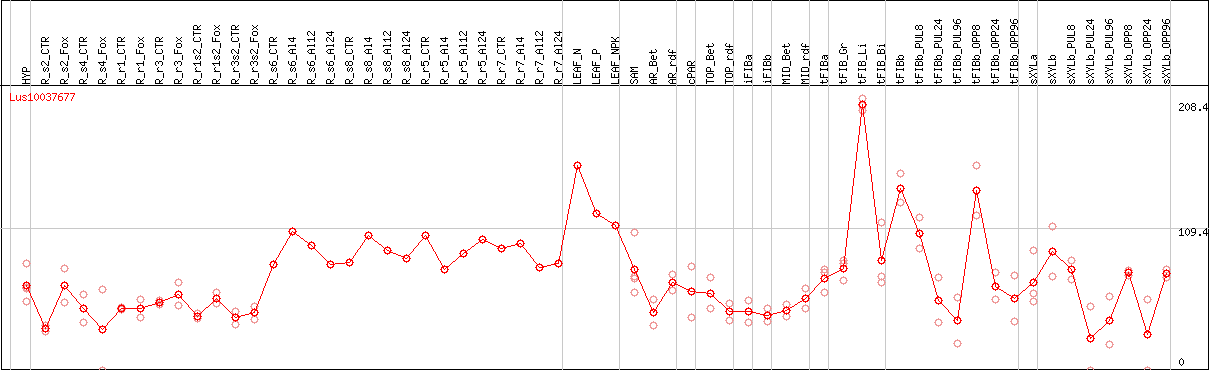
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037677 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.