External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G10400 215 / 2e-69
U11/U12-31K
U11/U12-31K, RNA recognition motif and CCHC-type zinc finger domains containing protein (.1)
AT4G20030 75 / 2e-16
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G23830 64 / 1e-12
AtGRP4, GR-RBP4, GRP4
glycine-rich RNA-binding protein 4 (.1.2)
AT5G64200 64 / 1e-11
ATSC35, At-SC35
ARABIDOPSIS THALIANA ORTHOLOG OF HUMAN SPLICING FACTOR SC35, ortholog of human splicing factor SC35 (.1.2)
AT1G23860 60 / 9e-11
SRZ21, SRZ-21, RSZP21, RSZ21, At-RSZ21
RS-containing zinc finger protein 21 (.1.2.3.4)
AT1G53720 61 / 2e-10
ATCYP59, CYP59
cyclophilin 59 (.1)
AT4G13860 56 / 2e-10
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G60650 60 / 3e-10
AtRZ-1b
AtRZ-1b, RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.1), RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.2)
AT3G46020 56 / 3e-10
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT4G19610 59 / 1e-09
nucleotide binding;nucleic acid binding;RNA binding (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038693
368 / 8e-130
AT3G10400 244 / 6e-81
U11/U12-31K, RNA recognition motif and CCHC-type zinc finger domains containing protein (.1)
Lus10002699
62 / 5e-11
AT3G55460 138 / 3e-39
SC35-like splicing factor 30 (.1)
Lus10006936
60 / 6e-11
AT4G20030 137 / 4e-42
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10015146
58 / 2e-10
AT3G08000 145 / 5e-46
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10026925
58 / 3e-10
AT4G31580 184 / 2e-59
RS-containing zinc finger protein 22, serine/arginine-rich 22 (.1.2)
Lus10000953
61 / 4e-10
AT2G39670 588 / 0.0
Radical SAM superfamily protein (.1.2)
Lus10003957
59 / 4e-10
AT5G64200 217 / 9e-70
ARABIDOPSIS THALIANA ORTHOLOG OF HUMAN SPLICING FACTOR SC35, ortholog of human splicing factor SC35 (.1.2)
Lus10002222
59 / 4e-10
AT3G53460 263 / 1e-87
chloroplast RNA-binding protein 29 (.1.2.3.4)
Lus10031534
57 / 5e-10
AT3G08000 146 / 2e-46
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G227500
255 / 1e-85
AT3G10400 223 / 7e-73
U11/U12-31K, RNA recognition motif and CCHC-type zinc finger domains containing protein (.1)
Potri.003G074900
68 / 6e-14
AT4G20030 164 / 2e-52
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.003G023800
62 / 5e-11
AT5G64200 207 / 4e-66
ARABIDOPSIS THALIANA ORTHOLOG OF HUMAN SPLICING FACTOR SC35, ortholog of human splicing factor SC35 (.1.2)
Potri.019G050600
60 / 6e-11
AT1G23860 131 / 3e-39
RS-containing zinc finger protein 21 (.1.2.3.4)
Potri.005G097500
62 / 8e-11
AT1G53720 667 / 0.0
cyclophilin 59 (.1)
Potri.014G157300
58 / 1e-10
AT3G08000 129 / 2e-39
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.008G033000
60 / 2e-10
AT1G60650 172 / 3e-52
AtRZ-1b, RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.1), RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.2)
Potri.008G022280
57 / 2e-10
AT3G20930 203 / 2e-65
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.010G228700
60 / 3e-10
AT5G04280 172 / 7e-52
AtRZ-1c, RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.1)
Potri.008G022200
59 / 5e-10
AT3G20930 427 / 3e-149
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0511
Retroviral_zf
PF00098
zf-CCHC
Zinc knuckle
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Lus10037960 pacid=23163890 polypeptide=Lus10037960 locus=Lus10037960.g ID=Lus10037960.BGIv1.0 annot-version=v1.0
ATGGGCAAGGGCCGGGGGAGCGACAGCGATGAGGACGACACCTTCTACTACCGATATGCCTCCGTCTCAGCCCCACCTCCACCGTCAACATCCACTTCCG
CCTCCAACCGAAACTCAAAGCCGAGCAATTCAGGCGGCCTAGCTCCGTCCAAGTCCACCGTTTACGTATCGAATCTGGATTTCACCCTCACAAACTCTGA
TCTACACACCCTCTTCTCCACCTTCGGCAAGATAGCTCGAGTCACCGTCCTCAAGGACAGGGCCACCCGCCAGTCCCGCGGCGTGGCCTTCATTCAGTTC
GTCTCCCGCGACGACGCCGTCTCCGCCGCCCGCCAGATGGACAAGAAGATCCTCAACGGGCGAACCCTATCAGCGAACATCGCCAACGACAACGGCCGGG
CGACTGAGTTCATCAAGAAGAGGGTTTATCTGGATAAGAGCAGGTGCTACGAGTGTGGAGAGGGCGGTCATCTTTCGTACGAGTGCCCTAAGAATCAGCT
GGGCCCTAGGGAGAGGCCCGCGCCGAAGCGGACCCGAAGAGGCGGCGGCGGGGGTGGGGGAGGGGTTCATAGGAGGAAGGAGGTGGAGGCGGCGGCGGAG
GATGACGATGACGAAGGGGTGAATGATGGGGGAGGATTTGAAGAGGAGAATTGGGCTTCGGTGGTTGATGGTAGGGCGGATGAGAGATTGTTGACAGGAG
ATGAAGATGGAAACGGCGTAGGGAAGAAGAAGGTGAAGAAAGCCAGTTATTTCAGCGACGAGAGTGACGACGAGGATTGA
AA sequence
>Lus10037960 pacid=23163890 polypeptide=Lus10037960 locus=Lus10037960.g ID=Lus10037960.BGIv1.0 annot-version=v1.0
MGKGRGSDSDEDDTFYYRYASVSAPPPPSTSTSASNRNSKPSNSGGLAPSKSTVYVSNLDFTLTNSDLHTLFSTFGKIARVTVLKDRATRQSRGVAFIQF
VSRDDAVSAARQMDKKILNGRTLSANIANDNGRATEFIKKRVYLDKSRCYECGEGGHLSYECPKNQLGPRERPAPKRTRRGGGGGGGGVHRRKEVEAAAE
DDDDEGVNDGGGFEEENWASVVDGRADERLLTGDEDGNGVGKKKVKKASYFSDESDDED
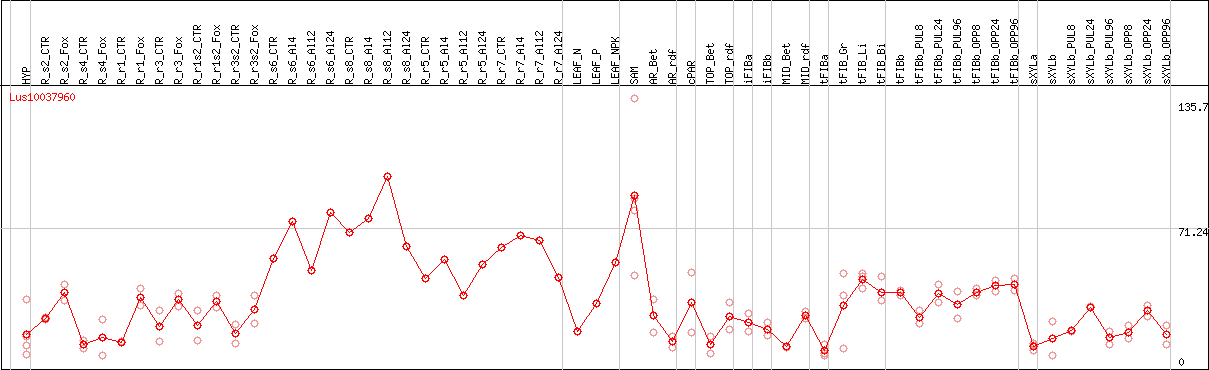
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037960 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.