External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G04260 149 / 3e-47
WCRKC2
WCRKC thioredoxin 2 (.1)
AT5G06690 118 / 8e-35
WCRKC1
WCRKC thioredoxin 1 (.1.2)
AT3G15360 49 / 4e-08
ATHM4, ATM4, TRX-M4
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
AT1G19730 45 / 4e-07
ATTRX4, ATH4
thioredoxin H-type 4, Thioredoxin superfamily protein (.1)
AT2G35010 45 / 6e-07
ATO1
thioredoxin O1 (.1.2)
AT1G03680 45 / 1e-06
ATHM1, ATM1, TRX-M1
ARABIDOPSIS THIOREDOXIN M-TYPE 1, thioredoxin M-type 1 (.1)
AT4G03520 44 / 4e-06
ATHM2
Thioredoxin superfamily protein (.1.2)
AT3G53220 42 / 1e-05
Thioredoxin superfamily protein (.1)
AT5G42980 40 / 3e-05
ATTRXH3, ATTRX3, ATH3
THIOREDOXIN H3, thioredoxin H-type 3, thioredoxin 3 (.1)
AT1G11530 40 / 3e-05
ATCXXS1
C-terminal cysteine residue is changed to a serine 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038703
182 / 2e-59
AT5G04260 191 / 2e-61
WCRKC thioredoxin 2 (.1)
Lus10021067
120 / 9e-36
AT5G06690 215 / 3e-71
WCRKC thioredoxin 1 (.1.2)
Lus10017244
73 / 2e-17
AT5G06690 156 / 2e-48
WCRKC thioredoxin 1 (.1.2)
Lus10007630
47 / 2e-07
AT1G11530 143 / 4e-45
C-terminal cysteine residue is changed to a serine 1 (.1)
Lus10018383
47 / 2e-07
AT1G11530 145 / 3e-46
C-terminal cysteine residue is changed to a serine 1 (.1)
Lus10041799
45 / 4e-07
AT3G51030 182 / 8e-61
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10028349
45 / 6e-07
AT3G51030 179 / 1e-59
ARABIDOPSIS THALIANA THIOREDOXIN H-TYPE 1, thioredoxin H-type 1 (.1)
Lus10013827
45 / 6e-07
AT3G53220 172 / 2e-56
Thioredoxin superfamily protein (.1)
Lus10042784
45 / 1e-06
AT3G15360 165 / 4e-52
ARABIDOPSIS THIOREDOXIN M-TYPE 4, thioredoxin M-type 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF00085
Thioredoxin
Thioredoxin
Representative CDS sequence
>Lus10037975 pacid=23163668 polypeptide=Lus10037975 locus=Lus10037975.g ID=Lus10037975.BGIv1.0 annot-version=v1.0
ATGTCAGAGCCTAATAGCGTTGTTTACTCTTCCAGGATGGCCAATTGGTGCAGAAAATGTATATATTTGAAACCAAAGCTGGAAAGATTAGCAGCTGACT
ATCATCCCAGTCTGAGATTCTACTGCGTAGACGTCAACAACGTGCCTCACAAGCTAGTAGCTCGAGCAGGAGTCACTAAAATGCCAACAATACAGCTGTG
GAGAGATGGGGAGAAGCAAGGAGAAGTAATTGGCGGGCACAAGGCTTACCAGGTGATCAATGAAGTTAGACAAATGATTGAGACTAGTAGTATCGAGGGT
GGTGACATCTGA
AA sequence
>Lus10037975 pacid=23163668 polypeptide=Lus10037975 locus=Lus10037975.g ID=Lus10037975.BGIv1.0 annot-version=v1.0
MSEPNSVVYSSRMANWCRKCIYLKPKLERLAADYHPSLRFYCVDVNNVPHKLVARAGVTKMPTIQLWRDGEKQGEVIGGHKAYQVINEVRQMIETSSIEG
GDI
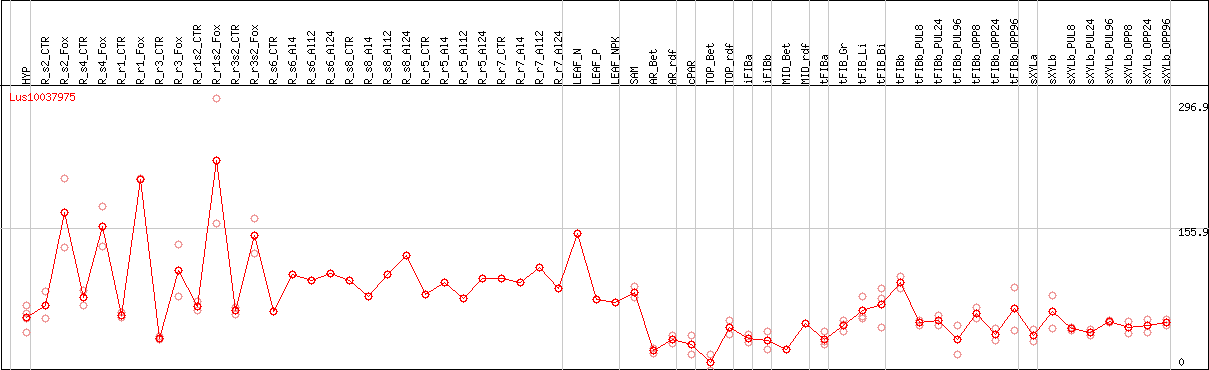
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037975 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.