External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G46210 377 / 8e-134
Ribosomal protein S5 domain 2-like superfamily protein (.1.2.3.4.5.6)
AT2G07110 78 / 7e-18
unknown protein
AT3G61620 80 / 1e-17
RRP41
3'-5'-exoribonuclease family protein (.1.2)
AT4G27490 64 / 9e-12
3'-5'-exoribonuclease family protein (.1)
AT3G03710 51 / 3e-07
PDE326, PNP, RIF10
resistant to inhibition with FSM 10, POLYNUCLEOTIDE PHOSPHORYLASE, PIGMENT DEFECTIVE 326, polyribonucleotide nucleotidyltransferase, putative (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009221
468 / 1e-169
AT3G46210 387 / 4e-138
Ribosomal protein S5 domain 2-like superfamily protein (.1.2.3.4.5.6)
Lus10005148
67 / 6e-13
AT3G61620 376 / 4e-133
3'-5'-exoribonuclease family protein (.1.2)
Lus10003036
66 / 3e-12
AT3G61620 370 / 9e-131
3'-5'-exoribonuclease family protein (.1.2)
Lus10037395
61 / 2e-10
AT4G27490 409 / 2e-146
3'-5'-exoribonuclease family protein (.1)
Lus10041318
52 / 9e-08
AT4G27490 414 / 4e-148
3'-5'-exoribonuclease family protein (.1)
Lus10013636
44 / 8e-05
AT3G03710 1148 / 0.0
resistant to inhibition with FSM 10, POLYNUCLEOTIDE PHOSPHORYLASE, PIGMENT DEFECTIVE 326, polyribonucleotide nucleotidyltransferase, putative (.1)
Lus10001332
44 / 0.0001
AT3G03710 954 / 0.0
resistant to inhibition with FSM 10, POLYNUCLEOTIDE PHOSPHORYLASE, PIGMENT DEFECTIVE 326, polyribonucleotide nucleotidyltransferase, putative (.1)
Lus10001173
40 / 0.001
AT3G07750 466 / 6e-168
3'-5'-exoribonuclease family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G043600
418 / 8e-150
AT3G46210 404 / 2e-144
Ribosomal protein S5 domain 2-like superfamily protein (.1.2.3.4.5.6)
Potri.006G239100
284 / 2e-97
AT3G46210 295 / 4e-102
Ribosomal protein S5 domain 2-like superfamily protein (.1.2.3.4.5.6)
Potri.014G094300
65 / 4e-12
AT3G61620 379 / 6e-135
3'-5'-exoribonuclease family protein (.1.2)
Potri.004G212500
64 / 8e-12
AT4G27490 392 / 2e-139
3'-5'-exoribonuclease family protein (.1)
Potri.009G010600
62 / 4e-11
AT4G27490 394 / 3e-140
3'-5'-exoribonuclease family protein (.1)
Potri.019G040100
52 / 3e-07
AT3G03710 1197 / 0.0
resistant to inhibition with FSM 10, POLYNUCLEOTIDE PHOSPHORYLASE, PIGMENT DEFECTIVE 326, polyribonucleotide nucleotidyltransferase, putative (.1)
Potri.013G065700
48 / 5e-06
AT3G03710 1205 / 0.0
resistant to inhibition with FSM 10, POLYNUCLEOTIDE PHOSPHORYLASE, PIGMENT DEFECTIVE 326, polyribonucleotide nucleotidyltransferase, putative (.1)
Potri.014G166200
42 / 0.0003
AT3G07750 422 / 3e-150
3'-5'-exoribonuclease family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0329
S5
PF01138
RNase_PH
3' exoribonuclease family, domain 1
Representative CDS sequence
>Lus10037991 pacid=23163846 polypeptide=Lus10037991 locus=Lus10037991.g ID=Lus10037991.BGIv1.0 annot-version=v1.0
ATGGAGATAGACAGAAACGATGGCCGCACTGCGAACCAGCTGAGGCCGTTGGGTTGCTCTCGCAGCATCCTCCACCGCGCGCACGGATCCGCCAGTTGGT
CTCAAGGAGAAACGAAGGTTATAGCTGCTGTTTATGGACCTAAGGCTGGAGTTAAGAAGAATGAGGACCCTGAGAAAGCTTGCATTGAAGTCATTTGGAA
GCCCAAAACTGGACAAATTGGGAAGGAGGAAAAAGAGTGTGAGATGATAGTGAAGAAGACTATACAGAGCATTTGTCTTTTGACAATCAACCCAAACACA
ACGACGTCAATTATAGTTCAGATGATAAAATGCTGTAGCAGCACTGCTCCTGGTGAGACTTATATGGTGTTTGTCACGGTTGTTAATGATGATGGTGCTC
TTCTACCATGTGCTATAAATGCAGCATGTGCTGCCCTTGTGGATGCTGGAATTCCCATGAAGCACCTTGCTGTAGCAATTTGTTGTTGTGTTGCAGAAGG
GGGATCCGTAGTACTCGACCCCACAAAGTTGGAGGAGCAGCAAATGAGAGGGTTTGCATATCTGGTATTTCCAAACTCGGGTCAATCGGCTCTACCAGAA
GGACCATCTCTCGTGGATGGCGAGCCGTTAGAACATGGGATCATCACCTCTGTCACCCATGGCATTATGTCAGTGGAAGAATACTTGAAATGTGTTGAAA
GAGGACGTGCAGCATGCTCGAAGCTGTCTTGTTTTGTCAGGAGGAACTTGCAATCACAACTTCCCAGTGACTCTTCCAAAGCTGCATAG
AA sequence
>Lus10037991 pacid=23163846 polypeptide=Lus10037991 locus=Lus10037991.g ID=Lus10037991.BGIv1.0 annot-version=v1.0
MEIDRNDGRTANQLRPLGCSRSILHRAHGSASWSQGETKVIAAVYGPKAGVKKNEDPEKACIEVIWKPKTGQIGKEEKECEMIVKKTIQSICLLTINPNT
TTSIIVQMIKCCSSTAPGETYMVFVTVVNDDGALLPCAINAACAALVDAGIPMKHLAVAICCCVAEGGSVVLDPTKLEEQQMRGFAYLVFPNSGQSALPE
GPSLVDGEPLEHGIITSVTHGIMSVEEYLKCVERGRAACSKLSCFVRRNLQSQLPSDSSKAA
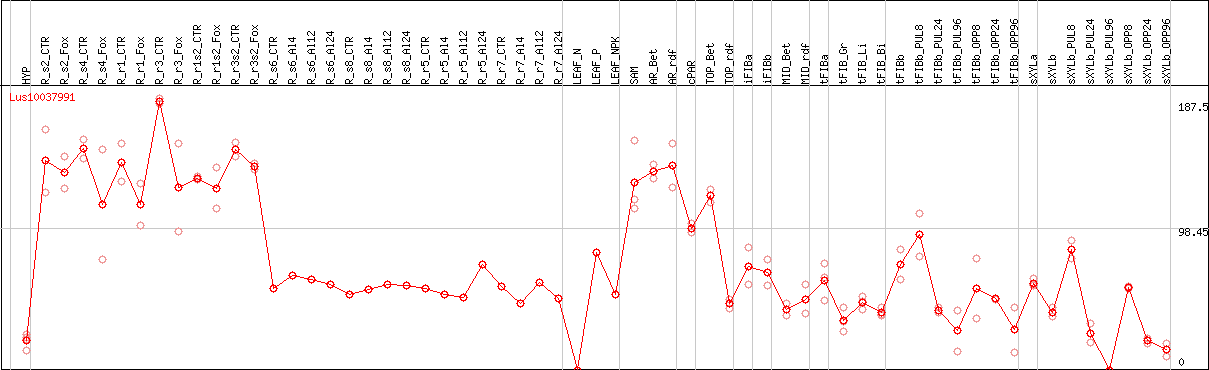
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10037991 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.