External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G62100 118 / 3e-34
AUX_IAA
IAA30
indole-3-acetic acid inducible 30 (.1)
AT2G46990 118 / 4e-34
AUX_IAA
IAA20
indole-3-acetic acid inducible 20 (.1)
AT3G17600 107 / 8e-30
AUX_IAA
IAA31
indole-3-acetic acid inducible 31 (.1)
AT2G33310 94 / 1e-23
AUX_IAA
IAA13
auxin-induced protein 13 (.1.2.3)
AT4G28640 90 / 2e-22
AUX_IAA
IAA11
indole-3-acetic acid inducible 11 (.1.2.3)
AT1G04550 90 / 3e-22
AUX_IAA
BDL, IAA12
indole-3-acetic acid inducible 12, BODENLOS, AUX/IAA transcriptional regulator family protein (.1.2)
AT5G25890 84 / 1e-20
AUX_IAA
IAR2, IAA28
IAA-ALANINE RESISTANT 2, indole-3-acetic acid inducible 28 (.1)
AT3G16500 84 / 6e-20
AUX_IAA
IAA26, PAP1
indole-3-acetic acid inducible 26, phytochrome-associated protein 1 (.1)
AT5G65670 85 / 7e-20
AUX_IAA
IAA9
indole-3-acetic acid inducible 9 (.1.2)
AT1G51950 82 / 2e-19
AUX_IAA
IAA18
indole-3-acetic acid inducible 18 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009967
228 / 9e-78
AT3G62100 128 / 2e-38
indole-3-acetic acid inducible 30 (.1)
Lus10010081
166 / 5e-53
AT2G46990 115 / 4e-33
indole-3-acetic acid inducible 20 (.1)
Lus10007193
166 / 8e-53
AT3G62100 117 / 4e-34
indole-3-acetic acid inducible 30 (.1)
Lus10014464
93 / 6e-23
AT2G33310 231 / 4e-75
auxin-induced protein 13 (.1.2.3)
Lus10023719
93 / 7e-23
AT2G33310 223 / 4e-72
auxin-induced protein 13 (.1.2.3)
Lus10022868
89 / 1e-21
AT4G28640 198 / 1e-62
indole-3-acetic acid inducible 11 (.1.2.3)
Lus10039413
84 / 3e-20
AT1G04240 250 / 2e-85
SHORT HYPOCOTYL 2, indole-3-acetic acid inducible 3, AUX/IAA transcriptional regulator family protein (.1)
Lus10039488
83 / 3e-20
AT5G43700 247 / 3e-84
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Lus10038285
84 / 2e-19
AT3G16500 224 / 3e-72
indole-3-acetic acid inducible 26, phytochrome-associated protein 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G111700
144 / 3e-44
AT3G62100 150 / 7e-47
indole-3-acetic acid inducible 30 (.1)
Potri.002G186400
136 / 3e-41
AT3G62100 147 / 3e-45
indole-3-acetic acid inducible 30 (.1)
Potri.010G065200
91 / 3e-22
AT2G33310 243 / 4e-80
auxin-induced protein 13 (.1.2.3)
Potri.008G172400
91 / 3e-22
AT2G33310 246 / 4e-81
auxin-induced protein 13 (.1.2.3)
Potri.013G041300
87 / 1e-21
AT5G43700 239 / 5e-81
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Potri.006G255200
87 / 2e-21
AT4G32280 137 / 7e-40
indole-3-acetic acid inducible 29 (.1)
Potri.002G045000
84 / 2e-20
AT5G43700 236 / 5e-80
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Potri.005G218200
84 / 2e-20
AT5G43700 243 / 2e-82
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Potri.005G053800
83 / 3e-20
AT5G43700 239 / 7e-81
indole-3-acetic acid inducible 4, AUXIN INDUCIBLE 2-11, AUX/IAA transcriptional regulator family protein (.1)
Potri.003G048100
86 / 4e-20
AT3G16500 224 / 4e-72
indole-3-acetic acid inducible 26, phytochrome-associated protein 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0072
Ubiquitin
PF02309
AUX_IAA
AUX/IAA family
Representative CDS sequence
>Lus10038025 pacid=23163681 polypeptide=Lus10038025 locus=Lus10038025.g ID=Lus10038025.BGIv1.0 annot-version=v1.0
ATGGGAAGAGGAGCGGCTAGTAATCATCATTACAACTCTACTTGCCCTTCTTCGTTTTCATCTTCTTCTTCTTCTTCTTCAATGTCTCATCACAAAAGAC
ACCGGAGAGACCTCACCACCGATCTCCGGCTCGGCCTCAGCAACAACTCCGGCGGCCGCGATGAAATGGAGTATGGAAACAACAACAACAACAACAACAA
CAACCATGGCGGCCGTGGTGGTGCTACTTTCTTTGTGAAGGTGTACATGGAAGGCATCCCAATTGGGAGGAAGTTGGATTTGTTGGCTTATGCTTCTTAT
GAAGACATGATCTCCACTCTTGATGCCATGTTTGGCACCAACATTCTTTGGGATGAGATGGAAAGTGACCAGAGCAATTACAATGGACAAGTGCAATACC
ATGTGTTGACTTATGAAGACAAAGAAGGAGATTGGTTGATTGTTGGAGATGTTCCATGGGAGGTGTTTGTGTCCTGCGTGAAGAGACTGAAGATAACTAG
AGCAGATAGCTTGTGA
AA sequence
>Lus10038025 pacid=23163681 polypeptide=Lus10038025 locus=Lus10038025.g ID=Lus10038025.BGIv1.0 annot-version=v1.0
MGRGAASNHHYNSTCPSSFSSSSSSSSMSHHKRHRRDLTTDLRLGLSNNSGGRDEMEYGNNNNNNNNNHGGRGGATFFVKVYMEGIPIGRKLDLLAYASY
EDMISTLDAMFGTNILWDEMESDQSNYNGQVQYHVLTYEDKEGDWLIVGDVPWEVFVSCVKRLKITRADSL
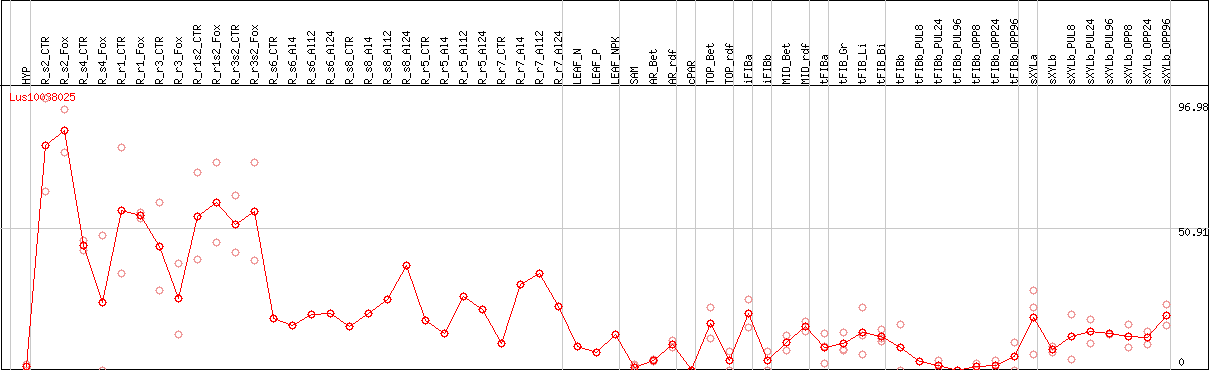
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038025 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.