Lus10038129 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10038129 pacid=23163815 polypeptide=Lus10038129 locus=Lus10038129.g ID=Lus10038129.BGIv1.0 annot-version=v1.0
ATGGGCTCCCCCCACGACAGGCTCCCTGATTTTGTTCGAAAAGTTAAATCCTTGATTTCTGGGAGGAGTGACCCTACAGATCTGTCTAGGGATTTTTGGA
TGCCTGATCAGAGCTGTAGGGTATGCTACGATTGTGATTCTCAGTTCACCGTATTTAACAGAAGACATCACTGCCGAGTCTGTGGCCGGATTTTTTGTGC
TAAGTGTACGTCAAATGCAGTTATCCACCCGTCTGATAAGCAAAAGACTTCTTGTGAAGACAAAGAGAGAGTGAGGGTCTGTAATTATTGCTTTAAGCAA
TATGAGGGTAATGGTTTAGCTATTGATAATGAGTCTAATGTGGCTGCTCCAGGCCTCAATTCATCACCTTCACCGATATTGGCCGACAGCAGTTCTAGCT
CTATCTGTAACAGTAGTAGCAGCACAGTTGGATCAGGTCCATATTCAACTGGGCCATACCTCTGTGTGTCGCATAGTTCTTGTCTTAGTCCTCAGAAGGC
TGCACCAATGGATGCAGCAACTGTTGAGCAAGAAACTGAGGAATACAGAGGTAGTATAGACCCCTCCAGAGCTGCATTTCATTCTTCTTCCACCCAGAAT
GCATATTGCATGAACAGCGGGCCAAGTGATGATGATGACGATGATGATGATTGCTATGGTATCTACGATGCAGGCTTGCAGGCGAGGCATTTCTCTTATG
TTGATAAATATTGTGGGTCTGCATCTGTTCATGACTTCGACCATGAATGTGGGATACATGATGTACATGACAGAGGAGGCCTTGTTGATGATCAAATTGT
AAGTTTCTCACCATTAACAGAGGATTTTAATTCAAAGAGTTTGGATGGAATCGAGGGTACTGCAGAAGCTGAATCTTATGGTGATGATTGTGATGCTACT
CCATATGACGTGGAGAATGAAGATAATGAGCCTGTTGATTTTGAGAACAACGGTCTTCTTTGGGTACCTCCTGAACCAGAAGATGAAGAGGATGAAAGGG
AAACTCCTTTGTTTGATGATGATGAAGATGAAGAAGATGATGGGACTACAGGTGAGTGGGGATATTTACGTCCTTCCAATAGCTTTGGGAGCGGGGAACT
CAGGAGCAAGGAAAAATCAAGCGAGGAGCACAGGAATGCTATGAAGAATGTGGTGGAAGGACACTTTAGAGCTTTAGTTGCACAGCTCTTACAAGTTGAA
AACCTCCCAATTGGTGTGGATGATGATAATGCCGGTTGGCTGGATATAATTACTAATCTGTCATGGGAAGCAGCTAGTCATCTTAAACCAGATACAAGCA
ATGGCGGTGGAATGGATCCTGGTGGATATGTGAAGGTCAAGTGCCTAGCTTGTGGGAGTCGCACTGAAAGCATGGTGGTGAAAGGAGTTGTTTGCAAAAA
GAATGTTGCCCATCGGAGAATGATGTCAAAAATAAATAAACCTCGTTTTCTAATCCTTAGCGGTGCTTTAGAGTATCAGCGAGTTCCTAACCTTCTGTCA
AGTGTTGATACGTTGCTTCAGCAGGAAAAGGATCATTTGAAGATGGCAGTTGCAAAGATTGAAGCTCATCACCCTAATGTTCTTTTAGTAGAGAAGTCAG
TGTCCCGATTTGCTCAGGAATGCCTTCTTGTGAAGGACATTTCACTGGTCCTAAACATTAAGAAACCACTTTTGGATCGTATAGCCCGCTGCACTGGTGC
ACAAATAGTACCTTCAGTTGATCATCTCACGACTCAGAAGACGGGACACTGTGATTCATTCCATGTGGAGAAATTTTTCGAGGAGCATGGTAGCGCTGGC
CAAGGAGGAAAGAAGTTGACCAAGACTCTAATGTATTTTGAGGGTTGCCCAAGACCTTTTGGTTGCACTATTTTGTTGAAAGGTGCCAGTGGGGATGAAT
TGAAAAAAGTAAAACATGTGGTACAGTATGGAGTTTTCGCAGCTTATCACCTAGCTCTGGAGACATCTTTCCTGGCAGATGAGGGAGCGACACTGCCAGA
ACTTCCTTTGAGGTCTCCACTGACTGTGGCACTACCTGATAAATCAAGTATGGATAGATCCATTTCTGCTATACCTGGTTTCATAGTTTTGGGTGCTGAA
AAGCCTGTGAGTCCTCAGACACCTGTTGAACTTGTGAAGTCTGATATTGGGGCAGGGGACCTAAAGAAATCCGAGAATTGCATCACGGCTGATACAGCAT
CTTCTGCTATTGATAAACCTGCTTGTGCAGAAGAAGATGGTCATTCGAATTCGTTGTCTGAAAATCATCCTTGTCAAACAGTGTCCCAAAATGTTGGGCT
AAATGGAAATGAATCTCCTTATTTGAGTTCAATTTCTCCAACTTTAATTGGGGATAATTTGTTGGATTCTCACATTGACATACACCCAAGGAATAATATC
TCAGGGCTGTCCTATGGAAATGATTCTAAAGACACTGTGTTGGGTGAGGAAGAGGATGTTAATACTTCTGCCACTGATATCCTCTTAGCATTTTCAGGAT
TGCCTCAACCACAAGAATTTGTCAGTAGCCACGCTAACAATGATCTCTTGCCTGCTAATGGTTATATGACCCATGGGTCCTTCAACAAAGATGGTATAGA
TCAGAATGAGGATATTGGTCCCTCAAAAGAGGATTTCCCTCCATCACCTTCAGATCACCAGAGCATTCTGGTGTCATCATCAACACGATGTGTTTGGAAG
GGGACTGTATGTGAAAGGGCCCATCTCTGCCGGATTAAGTACTATGGGAGCTTTGATAGGCCTCTTGGAAGTTTCTTACGAGATCAGCTGTTTGATCAGA
ACTATGAATGCCGTTCTTGTGAGATGCCATCAGAAGCACATGTTCACTGCTATATTCACCAACAAGGAAGCCTTACCATATCAGTTCGGAAGCTGCCCGA
ATTTCTTTTAACTGGGGAGAGAGAAGGGAAAATCTGGATGTGGCATAGGTGTCTTCGTTGTCCTCGGATTAATGGCTTCCCACCTGCTACTCGTAGAGTG
GTTATGTCTGACGCTGCCTGGGGCTTATCTTTTGGTAAGTTTTTAGAGCTTAGCTTTTCAAATCATGCAGCAGCAAGCAGGGTTGCGAGCTGTGGTCACT
CTCTACACAGAGATTGTCTTCGATTCTATGGTTTTGGGAGAATGGTTGCTTGCTTCCGGTATGCTTCAATTAATGTTAACTCTGTTTGTCTGCCACCTTC
AAAGCTTGAATTCAACTTTGATAGTCAGGAGTGGATTCAGACAGAAGCAAATGAGGTAAGACAAATGGCAGAACTTCTGTTTTCTGAAGTTGGGGCTGCT
TTCCGTAAGATTTTGGAGAAGGCTTCATCTGCTGGGTTAGATGATGATAGTGTCAAGGCATCATCAAGACTTCACAGTTTGGAGTTGGAGCAAGCATTAC
TGAAAGAAAAAGCAGAATTTGAGTACGATCCTCAACATTACTTGTACTTTACAGGTTCACAGAGGTATAAGCCAATCACTGGAATAGGGATGCCACTTCC
AAATTTTAACACTTCTTTTGATGGGAGTCCTTCCATTCCTAAAAATCTTATTCTAAACGAGTACGATCCAGTTTATATTCCACTTTTGAAGGAAGTGGAA
CAGCAAAGTGGCGCTAGGCTGTTTCTGCCTGTTGGAGTTAATGATACAGTTGTGCCGATATATGATGACGAGCCAACAAGCATTGTAGCATATGCTCTAG
TCTCATCGGCTTATCAGGCCCAAATTTATGAGCCTGAAACCACAAAAAATGCAGGGGACTCGATGCCTCAACATTTGAATTCTGTAAATCCTCTCTCTCA
GTCATTTGATGACTCTATTTCTGATACCCTTAAAAGTATTGGATCAATGGACGAGAGTATCCTATCATTGACTAGTTCTAAAAGTTCTCTGGTTTCAGAC
CTAGTTTCTCACACGAAGGATTTGCATGCCAGAATTTCTTTCAATGCTGAGGGACCACTTGGTAAGGTGAAGTATACAGTAACATGTTACTACGCAAAGA
GGTTTGAGGCTTTGAAAAACATCTGTTGCCCTCGTGATTTAGACTTTGTAAGATCTCTTAGTCGTTGTAAGAAGTGGGGAGCCCAGGGTGGCAAGAGCAA
CGTCTTCTTTGCTAAAACTCTGGACGATCGGTTTATCATCAAGCAAGTTACAAAAACAGAGCTGGAGTCATTCATCAAGTTTGGACCTTCATATTTCAAG
TACTTAACGGAATCAATTATTACCCGAAGCCCAACATGCTTGGCAAAGATCCTTGGAATATATCAGGTCTCAAAGCAGGTTAAAGGAGGAAAAGAAACCA
AGATGGATATTCTAGTGATGGAGAATCTTCTATTCCGTCGAGATGTGTCTCGACTTTATGACCTCAAAGGATCATCTCGGTCTCGTTATAATGCAGATAC
AAGCGGGAGCAACAAGGTTTTGCTGGACCAAAATTTGATCGAAGCGATGCCAACATCTCCAATTTTTGTAGGGAACAAGGCAAAGCGGTTACTAGAGAGA
GCCGTGTGGAATGATACTGCGTTTCTTGCCTCGATCGATGTAATGGACTACTCTCTACTGGTTGGAGTGGACGAGGGGAAACACGAGTTGGTGCTTGGAA
TCATTGATTTCATGAGACAGTATACGTGGGACAAGCATTTGGAGACTTGGGTGAAGGCTTCAGGGATCCTTGGCGGGTCGAAGAACTCGATGCCAACTGT
GATATCTCCCCAGCAGTACAAGAAACGGTTTAGGAAAGCTATGACTGCTTATTTTGTTATGGTTCCTGATCATTGGTCTCCGACTACGATGATTGTACCC
AGTGGCACCCAATCCGATGCTGCTTTTGATGAAAATAGCTTGCAAGCCCAATCTTTATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10038129 pacid=23163815 polypeptide=Lus10038129 locus=Lus10038129.g ID=Lus10038129.BGIv1.0 annot-version=v1.0
MGSPHDRLPDFVRKVKSLISGRSDPTDLSRDFWMPDQSCRVCYDCDSQFTVFNRRHHCRVCGRIFCAKCTSNAVIHPSDKQKTSCEDKERVRVCNYCFKQ
YEGNGLAIDNESNVAAPGLNSSPSPILADSSSSSICNSSSSTVGSGPYSTGPYLCVSHSSCLSPQKAAPMDAATVEQETEEYRGSIDPSRAAFHSSSTQN
AYCMNSGPSDDDDDDDDCYGIYDAGLQARHFSYVDKYCGSASVHDFDHECGIHDVHDRGGLVDDQIVSFSPLTEDFNSKSLDGIEGTAEAESYGDDCDAT
PYDVENEDNEPVDFENNGLLWVPPEPEDEEDERETPLFDDDEDEEDDGTTGEWGYLRPSNSFGSGELRSKEKSSEEHRNAMKNVVEGHFRALVAQLLQVE
NLPIGVDDDNAGWLDIITNLSWEAASHLKPDTSNGGGMDPGGYVKVKCLACGSRTESMVVKGVVCKKNVAHRRMMSKINKPRFLILSGALEYQRVPNLLS
SVDTLLQQEKDHLKMAVAKIEAHHPNVLLVEKSVSRFAQECLLVKDISLVLNIKKPLLDRIARCTGAQIVPSVDHLTTQKTGHCDSFHVEKFFEEHGSAG
QGGKKLTKTLMYFEGCPRPFGCTILLKGASGDELKKVKHVVQYGVFAAYHLALETSFLADEGATLPELPLRSPLTVALPDKSSMDRSISAIPGFIVLGAE
KPVSPQTPVELVKSDIGAGDLKKSENCITADTASSAIDKPACAEEDGHSNSLSENHPCQTVSQNVGLNGNESPYLSSISPTLIGDNLLDSHIDIHPRNNI
SGLSYGNDSKDTVLGEEEDVNTSATDILLAFSGLPQPQEFVSSHANNDLLPANGYMTHGSFNKDGIDQNEDIGPSKEDFPPSPSDHQSILVSSSTRCVWK
GTVCERAHLCRIKYYGSFDRPLGSFLRDQLFDQNYECRSCEMPSEAHVHCYIHQQGSLTISVRKLPEFLLTGEREGKIWMWHRCLRCPRINGFPPATRRV
VMSDAAWGLSFGKFLELSFSNHAAASRVASCGHSLHRDCLRFYGFGRMVACFRYASINVNSVCLPPSKLEFNFDSQEWIQTEANEVRQMAELLFSEVGAA
FRKILEKASSAGLDDDSVKASSRLHSLELEQALLKEKAEFEYDPQHYLYFTGSQRYKPITGIGMPLPNFNTSFDGSPSIPKNLILNEYDPVYIPLLKEVE
QQSGARLFLPVGVNDTVVPIYDDEPTSIVAYALVSSAYQAQIYEPETTKNAGDSMPQHLNSVNPLSQSFDDSISDTLKSIGSMDESILSLTSSKSSLVSD
LVSHTKDLHARISFNAEGPLGKVKYTVTCYYAKRFEALKNICCPRDLDFVRSLSRCKKWGAQGGKSNVFFAKTLDDRFIIKQVTKTELESFIKFGPSYFK
YLTESIITRSPTCLAKILGIYQVSKQVKGGKETKMDILVMENLLFRRDVSRLYDLKGSSRSRYNADTSGSNKVLLDQNLIEAMPTSPIFVGNKAKRLLER
AVWNDTAFLASIDVMDYSLLVGVDEGKHELVLGIIDFMRQYTWDKHLETWVKASGILGGSKNSMPTVISPQQYKKRFRKAMTAYFVMVPDHWSPTTMIVP
SGTQSDAAFDENSLQAQSL
|
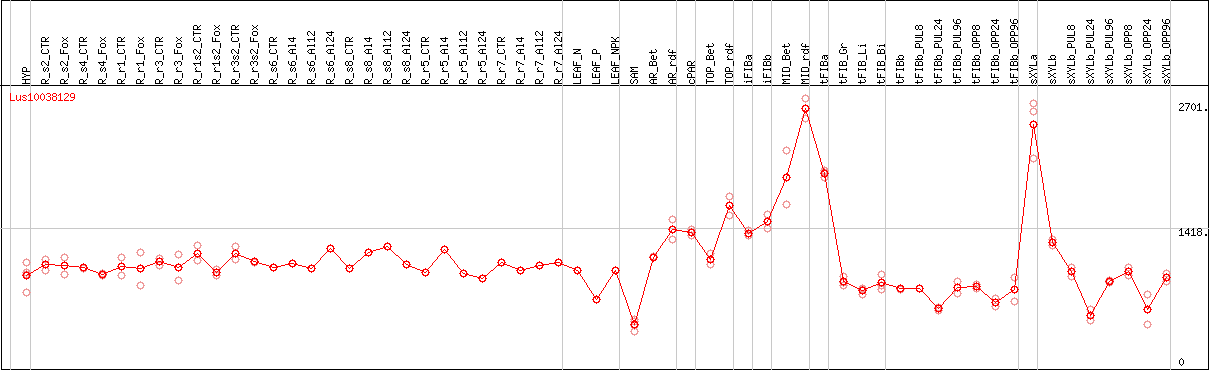
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038129 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
