External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G20750 61 / 3e-12
GATA
GATA29
GATA transcription factor 29 (.1)
AT4G36620 56 / 3e-10
GATA
GATA19, HANL2
hanaba taranu like 2, GATA transcription factor 19 (.1)
AT2G18380 55 / 6e-10
GATA
GATA20, HANL1
hanaba taranu like 1, GATA transcription factor 20 (.1)
AT3G50870 55 / 8e-10
GATA
GATA18, HAN, MNP
MONOPOLE, HANABA TANARU, GATA TRANSCRIPTION FACTOR 18, GATA type zinc finger transcription factor family protein (.1)
AT4G32890 53 / 5e-09
GATA
GATA9
GATA transcription factor 9 (.1)
AT5G25830 53 / 5e-09
GATA
GATA12
GATA transcription factor 12 (.1)
AT5G26930 50 / 8e-09
GATA
GATA23
GATA transcription factor 23 (.1)
AT3G54810 51 / 2e-08
GATA
GATA8, BME3, BME3-ZF
GATA TRANSCRIPTION FACTOR 8, BLUE MICROPYLAR END 3-ZINC FINGER, BLUE MICROPYLAR END 3, Plant-specific GATA-type zinc finger transcription factor family protein (.1.2)
AT5G49300 50 / 2e-08
GATA
GATA16
GATA transcription factor 16 (.1)
AT3G60530 50 / 5e-08
GATA
GATA4
GATA transcription factor 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042504
56 / 3e-10
AT3G20750 65 / 1e-12
GATA transcription factor 29 (.1)
Lus10020684
53 / 2e-09
AT4G16141 94 / 9e-24
GATA type zinc finger transcription factor family protein (.1)
Lus10011430
54 / 3e-09
AT4G32890 237 / 9e-77
GATA transcription factor 9 (.1)
Lus10029863
53 / 3e-09
AT4G16141 90 / 1e-22
GATA type zinc finger transcription factor family protein (.1)
Lus10037572
54 / 4e-09
AT4G32890 236 / 2e-76
GATA transcription factor 9 (.1)
Lus10028301
53 / 4e-09
AT3G50870 135 / 4e-39
MONOPOLE, HANABA TANARU, GATA TRANSCRIPTION FACTOR 18, GATA type zinc finger transcription factor family protein (.1)
Lus10025829
53 / 5e-09
AT4G32890 221 / 2e-70
GATA transcription factor 9 (.1)
Lus10038273
52 / 9e-09
AT4G32890 219 / 1e-69
GATA transcription factor 9 (.1)
Lus10016849
50 / 2e-08
AT3G06740 154 / 6e-49
GATA transcription factor 15 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0167
Zn_Beta_Ribbon
PF00320
GATA
GATA zinc finger
Representative CDS sequence
>Lus10038150 pacid=23158155 polypeptide=Lus10038150 locus=Lus10038150.g ID=Lus10038150.BGIv1.0 annot-version=v1.0
ATGCAGGGTGGGGAGACCGATCTGAGAATCGGTTCAAGTAGTGGAATAGTGAATGAGCAATATGGATATGGAACTCCAACATTTGGTATGGTTAACAAAG
ACCCAAACAACAACAACTACAACTACAACTACAACAACCCATCAGGTGTGGGAGTTCCGACATCAGCTGCGCTTGCTCCTCCTCTGGCCGCTCTTGCTCC
TACTGATCAGGCTACCTTCGTTAATGATTTGATGATTCCCCGCAATCTAAAAAGGCGTTTGTTTTGCAACACATTTGACACTCCTATGTGGCGTCGCGGA
CCTCTTGGCCCGAAGACGCTATGCAATGCGTGTGGACTTGAGTACCATAAGGATGAGAACAAAAACAAGGCAATGCAAGCTTCTACAAGCAATTGA
AA sequence
>Lus10038150 pacid=23158155 polypeptide=Lus10038150 locus=Lus10038150.g ID=Lus10038150.BGIv1.0 annot-version=v1.0
MQGGETDLRIGSSSGIVNEQYGYGTPTFGMVNKDPNNNNYNYNYNNPSGVGVPTSAALAPPLAALAPTDQATFVNDLMIPRNLKRRLFCNTFDTPMWRRG
PLGPKTLCNACGLEYHKDENKNKAMQASTSN
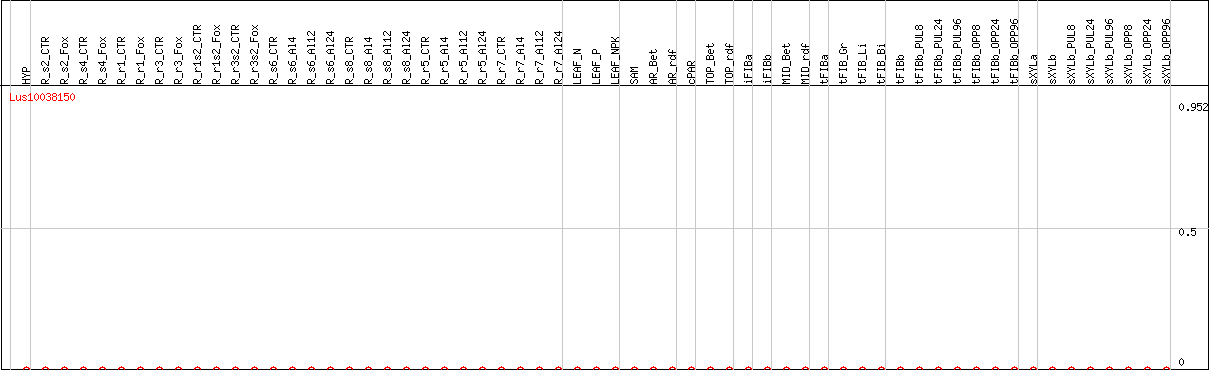
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038150 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.