External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G66870 172 / 1e-51
AS2
LBD36, ASL1
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
AT2G23660 154 / 7e-45
AS2
LBD10
LOB domain-containing protein 10 (.1.2)
AT3G27650 145 / 3e-43
AS2
LBD25
LOB domain-containing protein 25 (.1)
AT2G30130 145 / 1e-42
AS2
PCK1, LBD12, ASL5
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
AT5G63090 144 / 4e-42
AS2
LOBB, LOB
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
AT1G65620 141 / 5e-41
AS2
AS2
ASYMMETRIC LEAVES 2, Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
AT1G31320 136 / 3e-39
AS2
LBD4
LOB domain-containing protein 4 (.1)
AT2G40470 133 / 2e-37
AS2
ASL11, LBD15
ASYMMETRIC LEAVES2-LIKE 11, LOB domain-containing protein 15 (.1)
AT3G11090 129 / 7e-37
AS2
LBD21
LOB domain-containing protein 21 (.1)
AT2G30340 130 / 4e-36
AS2
LBD13
LOB domain-containing protein 13 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025840
381 / 1e-134
AT5G66870 184 / 4e-57
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10006679
317 / 1e-108
AT5G66870 169 / 1e-50
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10007024
296 / 1e-100
AT5G66870 173 / 2e-52
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10022463
185 / 6e-57
AT5G66870 221 / 8e-71
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10016769
180 / 6e-55
AT5G66870 222 / 3e-71
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10005313
177 / 2e-54
AT5G66870 209 / 4e-67
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10009337
159 / 2e-46
AT2G23660 183 / 7e-56
LOB domain-containing protein 10 (.1.2)
Lus10019633
158 / 3e-46
AT5G66870 181 / 7e-55
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Lus10007525
145 / 1e-42
AT2G30130 204 / 1e-67
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G039500
171 / 2e-51
AT5G66870 256 / 1e-84
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Potri.005G134900
164 / 7e-49
AT5G66870 254 / 8e-84
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Potri.001G345700
152 / 1e-45
AT3G27650 208 / 5e-70
LOB domain-containing protein 25 (.1)
Potri.012G083500
149 / 3e-44
AT5G63090 223 / 4e-75
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Potri.015G082200
147 / 2e-43
AT5G63090 223 / 4e-75
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Potri.010G184400
145 / 7e-43
AT2G30130 227 / 7e-77
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Potri.008G072800
144 / 2e-42
AT2G30130 229 / 8e-78
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Potri.005G097800
142 / 3e-42
AT1G31320 193 / 6e-64
LOB domain-containing protein 4 (.1)
Potri.013G081200
144 / 7e-42
AT2G40470 249 / 2e-84
ASYMMETRIC LEAVES2-LIKE 11, LOB domain-containing protein 15 (.1)
Potri.009G076900
142 / 9e-42
AT2G30130 228 / 2e-77
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03195
LOB
Lateral organ boundaries (LOB) domain
Representative CDS sequence
>Lus10038261 pacid=23158259 polypeptide=Lus10038261 locus=Lus10038261.g ID=Lus10038261.BGIv1.0 annot-version=v1.0
ATGTCACACTCGCAACAACCATGCGCGGCATGCAAGCTACAAAGGAGGAAGTGCACGCAGGAATGCATATTCGCACCCCATTTCCCACCGGACCAGCCTC
AGAAATTCGCCGACATCCACAAGGTTTTCGGAGCCAGCAACGTCGCAAAGTTGTTAAGTGAGCTCCCTACCATGAGCAGGGAAGAAGCAGTGATATCCCT
GTCTTACGAGGCGGGAGCCAGGATTCGTGACCCTGTATATGGTTGTGTAGGAGCCATCTCTGAATTACAACACAGACTAAAGCAAATACACATGGAAATT
GATGCGACCAGACAAGAGCTCCTCAAATACATGGGACCTAATAATCAGCAGCAGGATCAAGCTTTTATGATCCCCAATTTGGCTACTGCATCATTCATTG
CCCCGTCGACACACCATCAAAATCAACAGCAACAGCAACAGAACATGATGATGCAAATGATGGGCAGAGGTGGAGGAGGACATCAGATGTTGGAAGCTCA
ATTGATGGCTACTGAGCTAAATAGTACTACTGATAGAGGAGAACAATTACAACAACAACAACATCATGAGATGTTTAGGGTATATGAACAGAGAGAACAA
CAACAACAACAACAACATCATAATCAACATGAAGTGATTGGGTTTAGCAATGGAAATGGGATAATGGATGATCATGTTCCAAATTCTATTGGGTTTAATC
AGATTAGTATTGGGGAGGTTGAACCATGTTTGGCTCTTGGAAGCATGGATGATCAAAACAGTCATCATCATCAACAACAGGTAAATAATGAGCAATCACA
AGTGGAGATTCCGGTTCATCATGAGCAGCTTTTTTTCCATTCAGAGCATCATGAGCCAGACCTTAATCATCATCAGAGTTCAGGGAGTGATGAAGGTGGG
AATTCAGGTCCCAGTCTGAGTTAG
AA sequence
>Lus10038261 pacid=23158259 polypeptide=Lus10038261 locus=Lus10038261.g ID=Lus10038261.BGIv1.0 annot-version=v1.0
MSHSQQPCAACKLQRRKCTQECIFAPHFPPDQPQKFADIHKVFGASNVAKLLSELPTMSREEAVISLSYEAGARIRDPVYGCVGAISELQHRLKQIHMEI
DATRQELLKYMGPNNQQQDQAFMIPNLATASFIAPSTHHQNQQQQQQNMMMQMMGRGGGGHQMLEAQLMATELNSTTDRGEQLQQQQHHEMFRVYEQREQ
QQQQQHHNQHEVIGFSNGNGIMDDHVPNSIGFNQISIGEVEPCLALGSMDDQNSHHHQQQVNNEQSQVEIPVHHEQLFFHSEHHEPDLNHHQSSGSDEGG
NSGPSLS
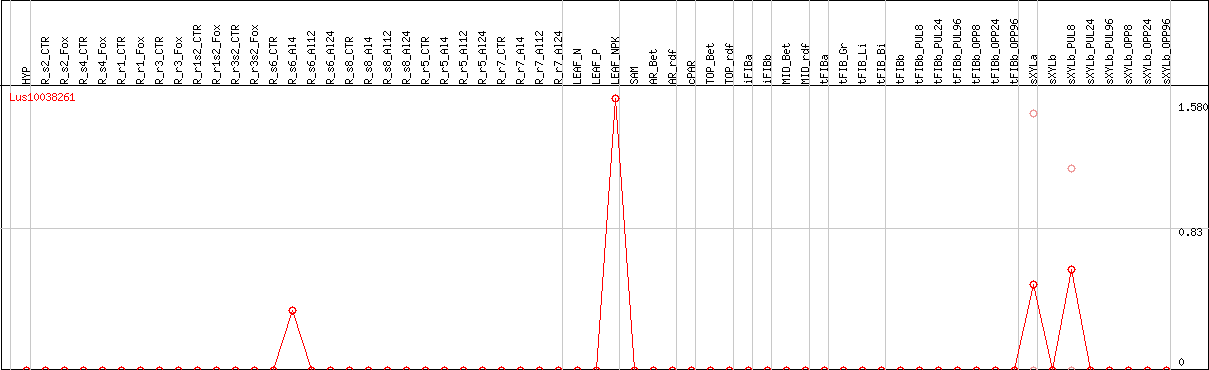
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038261 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.