External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G43000 231 / 3e-75
NAC
ANAC042, JUB1, JUNGBRUNNEN1
NAC domain containing protein 42 (.1)
AT3G12910 222 / 2e-71
NAC
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G39820 192 / 3e-59
NAC
ANAC094
NAC domain containing protein 94 (.1)
AT1G26870 188 / 5e-57
NAC
FEZ, ANAC009
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT2G02450 169 / 1e-49
NAC
LOV1, ANAC034, ANAC035
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
AT1G61110 160 / 4e-47
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT1G69490 157 / 1e-46
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT5G62380 158 / 4e-46
NAC
ANAC101, VND6
VASCULAR-RELATED NAC-DOMAIN 6, NAC-domain protein 101 (.1)
AT5G63790 155 / 2e-45
NAC
ANAC102
NAC domain containing protein 102 (.1)
AT3G04060 154 / 1e-44
NAC
ANAC046
NAC domain containing protein 46 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036194
429 / 2e-153
AT3G12910 224 / 1e-72
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10031189
236 / 4e-76
AT3G12910 264 / 9e-87
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10031767
210 / 5e-68
AT2G43000 236 / 3e-78
NAC domain containing protein 42 (.1)
Lus10015367
199 / 8e-62
AT1G26870 298 / 8e-99
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10030478
201 / 9e-62
AT5G39820 297 / 4e-98
NAC domain containing protein 94 (.1)
Lus10007263
198 / 1e-61
AT5G39820 300 / 3e-100
NAC domain containing protein 94 (.1)
Lus10036749
197 / 6e-60
AT1G26870 306 / 4e-100
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10037178
195 / 5e-59
AT5G39820 306 / 3e-101
NAC domain containing protein 94 (.1)
Lus10030446
168 / 1e-50
AT1G69490 302 / 2e-103
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G080900
261 / 4e-87
AT2G43000 257 / 3e-85
NAC domain containing protein 42 (.1)
Potri.003G149700
259 / 5e-86
AT2G43000 245 / 2e-80
NAC domain containing protein 42 (.1)
Potri.005G098000
245 / 2e-80
AT3G12910 271 / 2e-90
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.007G066300
243 / 1e-79
AT3G12910 269 / 2e-89
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.002G057200
235 / 8e-77
AT2G43000 282 / 4e-95
NAC domain containing protein 42 (.1)
Potri.005G205400
227 / 2e-73
AT2G43000 269 / 6e-90
NAC domain containing protein 42 (.1)
Potri.004G126901
210 / 1e-65
AT5G39820 305 / 1e-101
NAC domain containing protein 94 (.1)
Potri.017G082000
208 / 5e-65
AT1G26870 299 / 3e-98
FEZ, Arabidopsis NAC domain containing protein 9, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.003G046700
165 / 2e-48
AT2G02450 318 / 2e-106
LONG VEGETATIVE PHASE 1, Arabidopsis NAC domain containing protein 34, NAC domain containing protein 35 (.1.2)
Potri.008G089000
156 / 3e-46
AT1G69490 331 / 6e-115
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10038332 pacid=23158458 polypeptide=Lus10038332 locus=Lus10038332.g ID=Lus10038332.BGIv1.0 annot-version=v1.0
ATGGAGGAGGAGAAGCTGAAGCTGATGATTATGACGACCAAGAGGAGAAACGCAGCCGTTTCATGTTTCGTCGAGGAGGACAAGGACGACGACGTCGCTT
CACTTCCTGGGTTTCGATTTCACCCGACGGACGAGGAGCTTGTCGGATTTTATCTGAGGAGGAAAGTGGAGAAGAAGGCTTTGAAGATCGACATCATCAA
ACTTTGCGACGTCTACAAGTACCAGCCATGGGAGCTCCCAAGGGACGGAGGAGGAGGAGGAGAAAGGGAGATGTACTACTTTTGCAGGAGAGGGAGGAAG
TACAAGAACAGCGTCAGGCCCAACAGGGTGACAGCTGGCTGCGGATTCTGGAAAGCCACCGGAATCGACAAGTCCATTTATTCCGGCGGAGAGTGTGTCG
TCGGCCTTAAGAAATCACTAGTTTACTACCGAGGAAGTGCCGGAAAAGGCACCAAAACTGACTGGATGATGCACGAGTTCCGTCTCCCTCCCAACGATGG
CTATGAAGCAGAAGTTTGGACACTATGTCGTATATTCAAGCGAGAAGCTTCCTGCAGAAAGTACGCAACCACGGTGCAGCAACGGCAGGCGGTGAGTGAC
CGGAAGCGGTCGTCGCCGGTGGAGAATCATGGCTATTATTCAAGCAACGATATGGTGTCATCAGGACAGCTGATGGCGATGAATTACTTCGACCACGGAA
GCAGCACTAGTGCTGGTGATCAGCAGCAAGAAGCAGAAGAGATCAGTTTGGAAGAAGGGGAGTGGGATGAGCTTAGGCCAATGGTTGACTTTGCCCTCCT
CGATCATCATCATCGCCATTCCATCTTCTTCACCTAG
AA sequence
>Lus10038332 pacid=23158458 polypeptide=Lus10038332 locus=Lus10038332.g ID=Lus10038332.BGIv1.0 annot-version=v1.0
MEEEKLKLMIMTTKRRNAAVSCFVEEDKDDDVASLPGFRFHPTDEELVGFYLRRKVEKKALKIDIIKLCDVYKYQPWELPRDGGGGGEREMYYFCRRGRK
YKNSVRPNRVTAGCGFWKATGIDKSIYSGGECVVGLKKSLVYYRGSAGKGTKTDWMMHEFRLPPNDGYEAEVWTLCRIFKREASCRKYATTVQQRQAVSD
RKRSSPVENHGYYSSNDMVSSGQLMAMNYFDHGSSTSAGDQQQEAEEISLEEGEWDELRPMVDFALLDHHHRHSIFFT
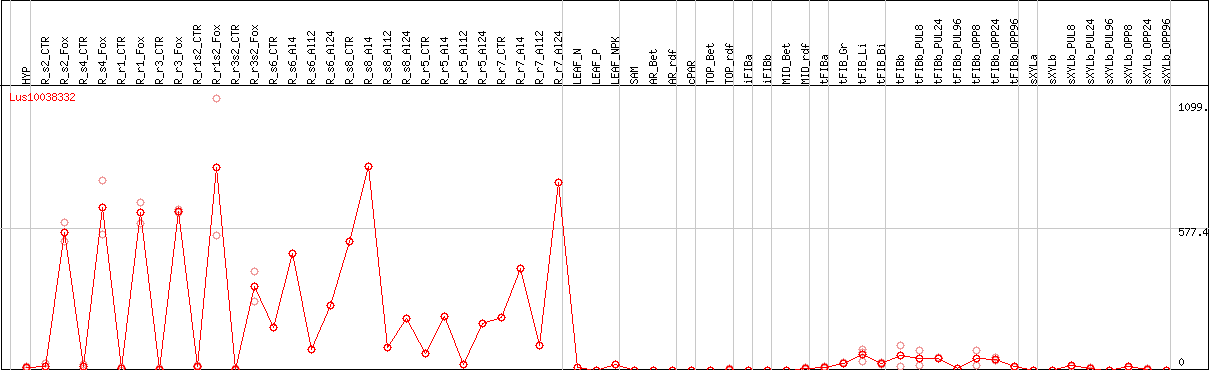
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038332 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.