External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G37990 336 / 4e-115
CAD-B2, ATCAD8, ELI3-2
CINNAMYL-ALCOHOL DEHYDROGENASE B2, ARABIDOPSIS THALIANA CINNAMYL-ALCOHOL DEHYDROGENASE 8, elicitor-activated gene 3-2 (.1)
AT4G37980 332 / 2e-113
ELI3-1, ATCAD7
CINNAMYL-ALCOHOL DEHYDROGENASE 7, elicitor-activated gene 3-1 (.1.2)
AT4G37970 328 / 7e-112
ATCAD6
cinnamyl alcohol dehydrogenase 6 (.1)
AT2G21730 323 / 1e-109
CAD2, ATCAD2
cinnamyl alcohol dehydrogenase homolog 2 (.1)
AT4G39330 322 / 3e-109
AtCAD1, ATCAD9
cinnamyl alcohol dehydrogenase 9 (.1.2)
AT2G21890 314 / 3e-106
CAD3, ATCAD3
cinnamyl alcohol dehydrogenase homolog 3 (.1)
AT3G19450 214 / 4e-67
CAD-C, ATCAD4
CINNAMYL ALCOHOL DEHYDROGENASE 4, GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
AT1G72680 205 / 1e-63
ATCAD1
CINNAMYL ALCOHOL DEHYDROGENASE 1, cinnamyl-alcohol dehydrogenase (.1)
AT4G34230 191 / 2e-58
CAD-5, ATCAD5, CAD-D, CAD6
cinnamyl alcohol dehydrogenase 5 (.1.2)
AT5G63620 67 / 2e-12
GroES-like zinc-binding alcohol dehydrogenase family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023268
508 / 0
AT4G37980 471 / 1e-167
CINNAMYL-ALCOHOL DEHYDROGENASE 7, elicitor-activated gene 3-1 (.1.2)
Lus10035956
380 / 2e-132
AT4G37990 509 / 0.0
CINNAMYL-ALCOHOL DEHYDROGENASE B2, ARABIDOPSIS THALIANA CINNAMYL-ALCOHOL DEHYDROGENASE 8, elicitor-activated gene 3-2 (.1)
Lus10025706
377 / 4e-131
AT4G37990 509 / 0.0
CINNAMYL-ALCOHOL DEHYDROGENASE B2, ARABIDOPSIS THALIANA CINNAMYL-ALCOHOL DEHYDROGENASE 8, elicitor-activated gene 3-2 (.1)
Lus10002089
304 / 3e-102
AT4G39330 577 / 0.0
cinnamyl alcohol dehydrogenase 9 (.1.2)
Lus10017285
268 / 2e-88
AT4G39330 413 / 1e-144
cinnamyl alcohol dehydrogenase 9 (.1.2)
Lus10003854
267 / 2e-88
AT4G39330 486 / 2e-174
cinnamyl alcohol dehydrogenase 9 (.1.2)
Lus10005611
263 / 2e-86
AT4G39330 409 / 9e-143
cinnamyl alcohol dehydrogenase 9 (.1.2)
Lus10027864
239 / 5e-77
AT3G19450 519 / 0.0
CINNAMYL ALCOHOL DEHYDROGENASE 4, GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
Lus10002812
238 / 1e-76
AT3G19450 523 / 0.0
CINNAMYL ALCOHOL DEHYDROGENASE 4, GroES-like zinc-binding alcohol dehydrogenase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.016G065300
372 / 3e-129
AT4G37990 492 / 1e-175
CINNAMYL-ALCOHOL DEHYDROGENASE B2, ARABIDOPSIS THALIANA CINNAMYL-ALCOHOL DEHYDROGENASE 8, elicitor-activated gene 3-2 (.1)
Potri.009G063400
367 / 6e-127
AT4G37980 490 / 1e-174
CINNAMYL-ALCOHOL DEHYDROGENASE 7, elicitor-activated gene 3-1 (.1.2)
Potri.006G199100
366 / 9e-127
AT4G37980 506 / 0.0
CINNAMYL-ALCOHOL DEHYDROGENASE 7, elicitor-activated gene 3-1 (.1.2)
Potri.001G268600
363 / 9e-126
AT4G37980 480 / 7e-171
CINNAMYL-ALCOHOL DEHYDROGENASE 7, elicitor-activated gene 3-1 (.1.2)
Potri.016G078300
360 / 2e-124
AT4G37970 482 / 2e-171
cinnamyl alcohol dehydrogenase 6 (.1)
Potri.001G307200
343 / 8e-118
AT4G39330 572 / 0.0
cinnamyl alcohol dehydrogenase 9 (.1.2)
Potri.001G372400
301 / 4e-101
AT4G39330 439 / 2e-154
cinnamyl alcohol dehydrogenase 9 (.1.2)
Potri.002G018300
259 / 8e-85
AT4G39330 409 / 5e-143
cinnamyl alcohol dehydrogenase 9 (.1.2)
Potri.009G063300
222 / 6e-72
AT4G37990 283 / 2e-95
CINNAMYL-ALCOHOL DEHYDROGENASE B2, ARABIDOPSIS THALIANA CINNAMYL-ALCOHOL DEHYDROGENASE 8, elicitor-activated gene 3-2 (.1)
Potri.006G024300
223 / 6e-71
AT1G72680 521 / 0.0
CINNAMYL ALCOHOL DEHYDROGENASE 1, cinnamyl-alcohol dehydrogenase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0296
GroES
PF08240
ADH_N
Alcohol dehydrogenase GroES-like domain
CL0063
NADP_Rossmann
PF00107
ADH_zinc_N
Zinc-binding dehydrogenase
Representative CDS sequence
>Lus10038536 pacid=23158395 polypeptide=Lus10038536 locus=Lus10038536.g ID=Lus10038536.BGIv1.0 annot-version=v1.0
ATGGCTGGAGAAGAAGAGCAAGAGCAAGTTCAAGCCTTTGGATGGGTTTCTCCCTTCCACTTCTCTCCGAGGGCTACGGGGGAGAAGGACGAGCAATTCA
AAGTGCTCTACTGTGGAATCTGTCACTCGGATCTCCACATGCTGAAAAACGAATGGGGGACGTCGATCTACCCGCTCGTTCCGGGCCATGAGATCGTCGG
CCTAGTAACTCGAGTGGGGAAGAATGTCGAGAAATTCAAAGTCGGGGACTGGGTGGGCGTCGGTTGCATGGTCGGATCATGCCGCAATTGCTACAACTGC
TCCAACGACCTCGCAAACTACTGCCCCGATTTCGTCCTAACATACTCCGCGCCCGCCAACCTCGTCCCCACGACGACCTACGGAGGCTGCTCGGACACCA
TGGTCTGCGACGAGCACTTTGTGGTCCAAATCCCGGACACCATCCCTATGGACACGGCCGCTCCTTTGCTGTGCGCCGGGATCACTGTTTACAGCCCGAT
CAGGGTTGGGCGGATTGAGATGACGGTCAAGTTTCTAAAAGCAATAGGGGTTAGAGTGATAGTGATCAGTACTTCCAATGGGAAGAAACCTGAAGCCCTG
GAGAGGCTTGGTGCTGACTCGTTCTTGGTTAGTAAAGATCCAGAGGAGTTGAATGCGACGGTCGGGAAATTGAATGGAATTATTAATATGGTTTTAGCAA
GTCAGTCGTTGTTGCCTTTGATCGGGTTGTTGAAGACTAACGACAAACTGATCTTGGTGGGTGTTCCGGAGAAGCCCCACGAGTTGCTCGCTTTTCTTTT
GTTGTTGGCAAAACACAACGTCATTGCCGACATTGAAGTCGTTACGATGGACTACGTGAATACCACCATGGAACGACTTACAAAGGCCGATGTCAAGTAC
AGATTCGTCATCAATGTTGCCAATACTATTTAA
AA sequence
>Lus10038536 pacid=23158395 polypeptide=Lus10038536 locus=Lus10038536.g ID=Lus10038536.BGIv1.0 annot-version=v1.0
MAGEEEQEQVQAFGWVSPFHFSPRATGEKDEQFKVLYCGICHSDLHMLKNEWGTSIYPLVPGHEIVGLVTRVGKNVEKFKVGDWVGVGCMVGSCRNCYNC
SNDLANYCPDFVLTYSAPANLVPTTTYGGCSDTMVCDEHFVVQIPDTIPMDTAAPLLCAGITVYSPIRVGRIEMTVKFLKAIGVRVIVISTSNGKKPEAL
ERLGADSFLVSKDPEELNATVGKLNGIINMVLASQSLLPLIGLLKTNDKLILVGVPEKPHELLAFLLLLAKHNVIADIEVVTMDYVNTTMERLTKADVKY
RFVINVANTI
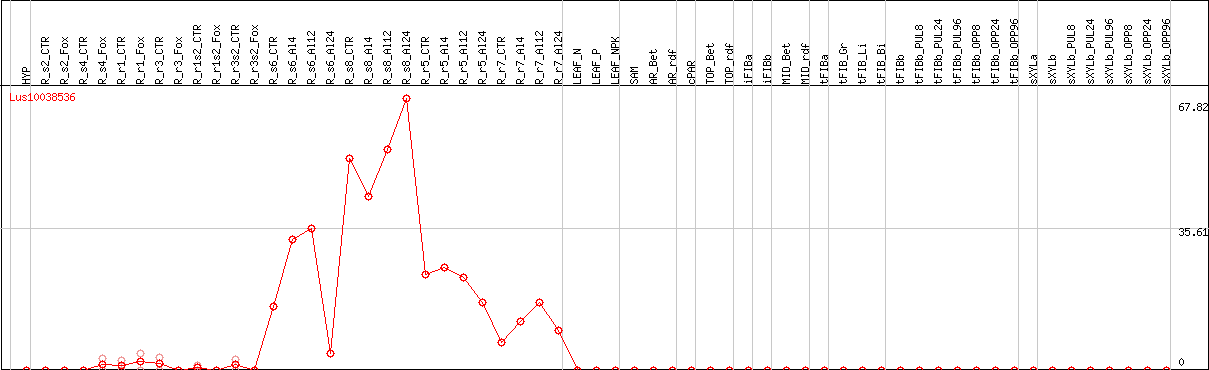
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038536 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.