External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G27950 65 / 3e-13
AP2_ERF
CRF4
cytokinin response factor 4 (.1)
AT4G11140 57 / 2e-10
AP2_ERF
CRF1
cytokinin response factor 1 (.1)
AT4G23750 54 / 4e-09
AP2_ERF
TMO3, CRF2
TARGET OF MONOPTEROS 3, cytokinin response factor 2 (.1.2)
AT1G71130 52 / 4e-09
AP2_ERF
CRF8
cytokinin response factor 8, Integrase-type DNA-binding superfamily protein (.1)
AT2G46310 51 / 2e-08
AP2_ERF
CRF5
cytokinin response factor 5 (.1)
AT5G53290 47 / 6e-07
AP2_ERF
CRF3
cytokinin response factor 3 (.1)
AT1G22985 40 / 0.0001
AP2_ERF
CRF7
cytokinin response factor 7, Integrase-type DNA-binding superfamily protein (.1)
AT1G43160 40 / 0.0001
AP2_ERF
RAP2.6, RAP2.06
related to AP2 6 (.1)
AT1G68550 40 / 0.0002
AP2_ERF
CRF10
cytokinin response factor 10, Integrase-type DNA-binding superfamily protein (.1.2)
AT5G61890 40 / 0.0002
AP2_ERF
Integrase-type DNA-binding superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027573
89 / 1e-21
AT4G27950 209 / 1e-64
cytokinin response factor 4 (.1)
Lus10039324
79 / 6e-18
AT4G27950 210 / 8e-65
cytokinin response factor 4 (.1)
Lus10017550
62 / 5e-12
AT4G23750 172 / 2e-51
TARGET OF MONOPTEROS 3, cytokinin response factor 2 (.1.2)
Lus10033938
61 / 2e-11
AT4G23750 201 / 6e-62
TARGET OF MONOPTEROS 3, cytokinin response factor 2 (.1.2)
Lus10032353
60 / 2e-11
AT4G23750 207 / 2e-64
TARGET OF MONOPTEROS 3, cytokinin response factor 2 (.1.2)
Lus10005149
50 / 5e-08
AT4G27950 106 / 6e-28
cytokinin response factor 4 (.1)
Lus10018756
45 / 7e-06
AT1G64280 506 / 2e-170
SALICYLIC ACID INSENSITIVE 1, NON-INDUCIBLE IMMUNITY 1, ARABIDOPSIS NONEXPRESSER OF PR GENES 1, regulatory protein (NPR1) (.1)
Lus10014054
42 / 5e-05
AT4G34410 139 / 4e-40
redox responsive transcription factor 1 (.1)
Lus10005805
40 / 0.0003
AT5G61890 120 / 4e-32
Integrase-type DNA-binding superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10038826 pacid=23150272 polypeptide=Lus10038826 locus=Lus10038826.g ID=Lus10038826.BGIv1.0 annot-version=v1.0
ATGGCTGATCCGAGTTTTCTCAGCCGGACGAAGTTCTCGAAGCACCGGAGTCAAGTCAGAAAGTACGCTCCCAAGCCGCCGGCGAAACGGAAGACGGAAA
AACACCCTTTGCCTCCGGCGGTGACGGCGGCGGAGGAGTTGGGTCGGAAAGTGTCCACCCGGGTGGTGAGGATATCGGTGACTGACCCGGATGCGACGGA
CTCCTCCAGCGACGAGGAAGCGGAGGTGTTCGGGATGATCAAGAGGTCCGAAAACGAGTACAACAGCGAGAGAAAAACGACGGAGGTGCGGCGTACGGCG
GCGGTGGGTGGTGCTCAGGGTGTGAAGAAGTTTAGGGGAGTTAGGCAGCGGCCATGGGGGAAGTGGGCGGCTGTCTCCTACTTCGGTTCTTCAGTTCAGG
AACCAATCGAGTGA
AA sequence
>Lus10038826 pacid=23150272 polypeptide=Lus10038826 locus=Lus10038826.g ID=Lus10038826.BGIv1.0 annot-version=v1.0
MADPSFLSRTKFSKHRSQVRKYAPKPPAKRKTEKHPLPPAVTAAEELGRKVSTRVVRISVTDPDATDSSSDEEAEVFGMIKRSENEYNSERKTTEVRRTA
AVGGAQGVKKFRGVRQRPWGKWAAVSYFGSSVQEPIE
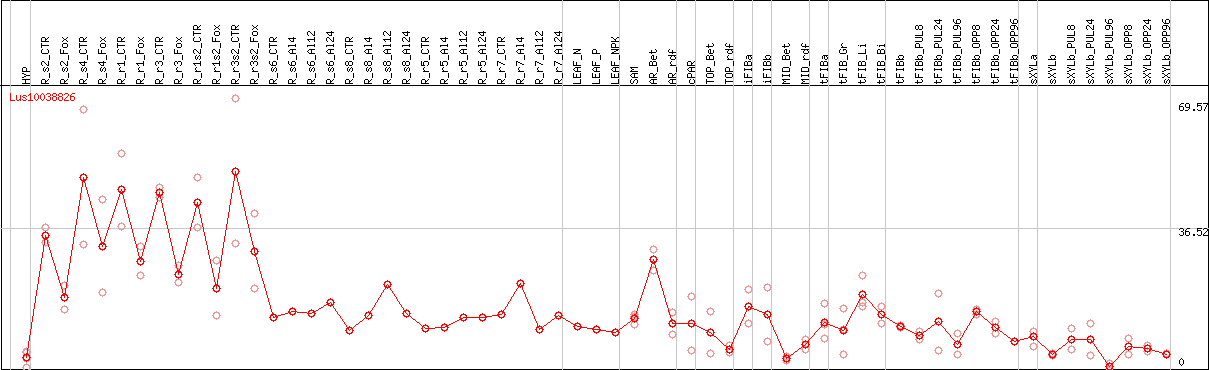
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038826 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.