External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G16380 62 / 2e-11
PAB6
poly(A) binding protein 6 (.1)
AT5G03580 55 / 3e-10
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G27330 55 / 5e-10
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT2G36660 56 / 2e-09
PAB7
poly(A) binding protein 7 (.1)
AT1G71770 56 / 2e-09
PAB5
poly(A)-binding protein 5 (.1), poly(A)-binding protein 5 (.2)
AT1G73530 52 / 2e-08
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT1G49760 52 / 3e-08
PABP8, PAB8
poly(A) binding protein 8 (.1), poly(A) binding protein 8 (.2)
AT4G34110 52 / 5e-08
PABP2, PAB2, ATPAB2
ARABIDOPSIS POLY\(A\) BINDING 2, poly(A) binding protein 2 (.1)
AT1G22760 51 / 1e-07
PAB3
poly(A) binding protein 3 (.1)
AT5G64200 50 / 1e-07
ATSC35, At-SC35
ARABIDOPSIS THALIANA ORTHOLOG OF HUMAN SPLICING FACTOR SC35, ortholog of human splicing factor SC35 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014958
293 / 2e-102
AT5G52545 55 / 2e-10
unknown protein
Lus10031534
54 / 2e-09
AT3G08000 146 / 2e-46
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10000826
54 / 8e-09
AT5G09880 589 / 0.0
Splicing factor, CC1-like (.1)
Lus10041728
54 / 1e-08
AT2G18510 436 / 1e-153
embryo defective 2444, RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10015146
51 / 1e-08
AT3G08000 145 / 5e-46
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10002093
54 / 2e-08
AT5G09880 600 / 0.0
Splicing factor, CC1-like (.1)
Lus10035556
53 / 2e-08
AT2G36670 623 / 0.0
Eukaryotic aspartyl protease family protein (.1.2)
Lus10027886
53 / 3e-08
AT4G34110 922 / 0.0
ARABIDOPSIS POLY\(A\) BINDING 2, poly(A) binding protein 2 (.1)
Lus10002835
53 / 3e-08
AT4G34110 918 / 0.0
ARABIDOPSIS POLY\(A\) BINDING 2, poly(A) binding protein 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Lus10038840 pacid=23150389 polypeptide=Lus10038840 locus=Lus10038840.g ID=Lus10038840.BGIv1.0 annot-version=v1.0
ATGGTATTTCTTTTTCCTTATCATGATTTCATACATTTCTCTGGCACTGATCATAACGGTTTCACCGATGAGAAAGCAGAAAGCAGAGTTTACGTTGGTA
ACCTTGACCTTAGAATAACAGAGGCGACTCTGATCAAGATGTTTTCCCCGTATGGTATGATCATCTCCGAGGACTTTCTATGGCACACCCGTGGCCCTAA
ACGCGGGGAGCCACGTGGCTTCGCGTTCGTCCAGTTCAGCTCCAAAGAGGAAGCTAGATTAGCCAAGGAGAAGATGCACGGAAGGTTAGCTTGTGGCCGG
CCATTGGTGGTACGTGTTTCCACCGAGCAAAACAACAACGATCAGTCATCGGCACCTCAAGGCTCTTCGAGAACAGCCAGGGAGACTAAGAAACCAGGAG
GCCTCCATCTCCTTACTTCTTCTTCACGTACCATGGGGCAGAAGAAGACGAGCAAGAGTGCGAAAATCGCAGCGATAAAGAACAAGTTGAGAGCCTTGGA
GGAAGAGACCACTGGTAGTAGTGTCGCCAAGAGACAGAAAACTGGCTGA
AA sequence
>Lus10038840 pacid=23150389 polypeptide=Lus10038840 locus=Lus10038840.g ID=Lus10038840.BGIv1.0 annot-version=v1.0
MVFLFPYHDFIHFSGTDHNGFTDEKAESRVYVGNLDLRITEATLIKMFSPYGMIISEDFLWHTRGPKRGEPRGFAFVQFSSKEEARLAKEKMHGRLACGR
PLVVRVSTEQNNNDQSSAPQGSSRTARETKKPGGLHLLTSSSRTMGQKKTSKSAKIAAIKNKLRALEEETTGSSVAKRQKTG
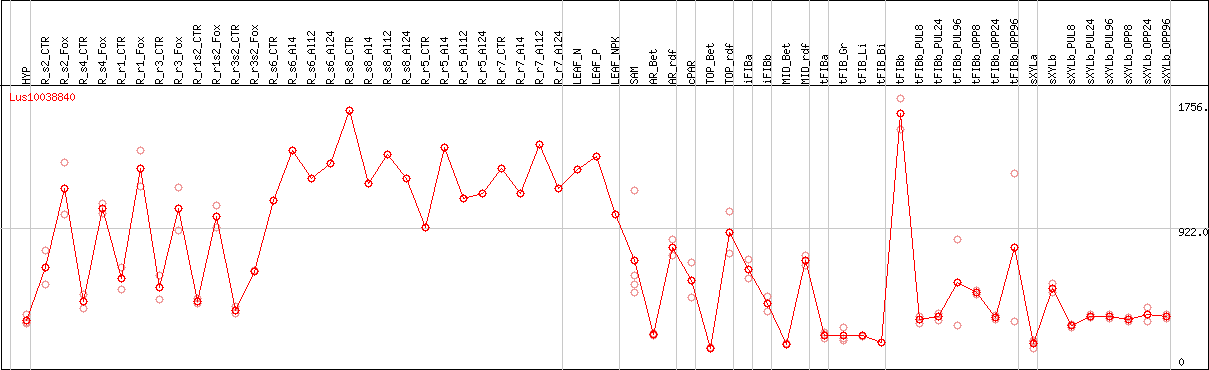
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10038840 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.