Lus10039014 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10039014 pacid=23150666 polypeptide=Lus10039014 locus=Lus10039014.g ID=Lus10039014.BGIv1.0 annot-version=v1.0
ATGGACCAACACAACCGGGCTCCGATGACCAATGAACAACACACCCGGGCTTCTCACACTCCGCCACGTGTACCCATCGCCACTTGTCCATCCTTCATTG
GCTCCAGTTCCTCATCACAACGAAACCAGCCACTGAGAAAGTCCTCCGCGTCAGCTTATATATATAATAACGAAAAACGTTTCACATCGTTTCCAGGAAA
AATAACGGCTCCCGCATCTCACAGCCTTTCTTCCTCTTCATTCGCCGATTTCCAGCTGAGCTCCTCAGCAGCCCAAGCTGAGAAAAATCCACCTCCCACT
GCTTCTGGCTCTTATTCAGTAGTTACTGATCCGTTAATGGCGGAATTACAGGGAAGGAGAGCGGCGGATTTGGGCTGTTTGTTTCTCATTTCTCGATCCA
TGCCGACGATGCTGATCAAATTGGGGTTGCTTCTCTGCTTCATTTCTGTAAACTTAATCCCGTCCCAATCGGCCGTCTCCAGCATAGATCTAGGGTCGGA
ATGGATGAAAGTAGCTGTAGTGAATCTCAAACCAGGTCAATCCCCGATTGCCATCGCGATCAATGAAATGTCAAAGCGCAAGTCTCCCGCCTTAGTGTCG
TTCCACTCTGGTACGCGTCTCCTCGCCGAGGAGGCTGCCGGGATCACAGCTCGTTACCCTAATAAAGTGTTCTCGCAGCTGCGGGATATGATAGGGAAAC
CGTACAAGGATGTGAAGGTAGTTTTGGATTCAATGTACCTGCCGTTTGAGGCTGTGGAGGATTCCAGGGGATCCGTTGCGTTTAAGGTTGATGACGGCCA
AGTTTATTCCGTTGAGGAATTGTTGGCTATGGTTTTGGGATACGCTTCGAATTTGGCTGAGGTACATGCCAAGCTACCTGTGAAGGATGCTGTCATTAGC
GTGCCGCCGTATTTTGGTCAGGCGGAGAGGAGAGGGTTGATTCAGGCAGCTCAGCTGGCTGGGATTAATGTTCTTACGTTGATGAATGAGCATTCTGGTG
CGGCATTGCAGTACGGGATTGATAAAGATTTCTCAAATGGATCGAGATATGTGATATTCTATGATATGGGTTCGAGTAGTACTTATGCTGCACTAGTCTA
TTATTCGGCCTACAACACCAAGGAGTATGGGAAGACTGTGTCTGTCAATCAGTTTCAGGTTAAGGATGTTAGATGGAATTCGCAGCTTGGAGGTCAGAGC
ATGGAAATGCGGTTGGTGGAATACTTTGCAGATGAATTTAACAAGCAAGTTGGAAATGGAGTTGATGTGCGGAAGTCTCCCAAGTCGATGGCTAAACTGA
AGAAGCAGGTTAAGCGGACAAAGGAAATATTGAGCGCAAACACAGCGGCTCCAATATCAGTTGAATCTCTATATGACGATCGGGATTTTAGGAGCACTAT
TACACGGGAGAAGTTTGAAGAGTTATGTGGTGATCTGTGGGACAGATCCCTTCTCCCCATCAAAGAAGTAATGAAGCATTCAGACCTGAAAGTGGATGAT
ATATCTGCTGTAGAGCTTATTGGAGGTGCCACAAGGGTGCCAAAGTTGCAGGCTAAGATCCAAGAATTTCTTGGAAGAAACGAGCTGGACAAACATCTGG
ATGCAGATGAAGCTATAGTTCTTGGTGGAGCATTGCATGCTGCAAATTTAAGTGATGGAATTAAATTGAACCGCAAGCTAGGAATGATTGATGGTTCTCC
ATATGGATTGTCCGTTGTACTGGATGGGCCAGACATGAAAGATGAGTACTCTAGGCAGTCACTGGTGCCTAGAATGAAAAAGGTTCCCAGTAAGATGTTC
AGATCCATCGTCCACAACAAAGATTTTGAAGTTTCACTTGAATATGAAACTGAGGAAATTTTACCACCCAGCATAAGCTCTCCAGTGTTCGCCCAGTATT
CTGTGTCTGGTCTTACTGAGGCGAGTGAGAAGTATTCATCACGGAATCTTTCCTCCCCTGTTAAGGCAAGTCTGCATTTCTCAATCAGTAGAAGTGGTGT
TCTCTCGTTGGACAGAGCAGATGCTGTTATTGAGATATCAGAGTGGGTAGAAGTCCCAATAAAAAAGAATCTTACCCTTGAAACTATCACCAACTCTTCA
AGTAGTTCTCCAGAAACTACCGAGAATACCACAACTAGCTCCACAACTGAGAGTTCTTCGGAGGAAAGTAATGAAAACCTACAATCTGATGGTGCAGCGG
GCAACGAATCTAACCCAGATGTCAAAGAACCAATTACCACTGAAGCGGCTACAGAAAAGAAGTTGAAGAAGCGGACTTTCAGGGTTCCATTGAAGATCAC
CGAAAAGACAGCAGGACCTGGAAAGCCTCTTTCAGAAGAATCTTTGGCTGAAGCTAAAAAGAAGCTTGAAGAATTGAACAAGAAGGACTCAGAGCGACGA
AGAACTGCTGAGTCGAAGAATAACTTGGAAGGATATATATACTCGACAAAAGAAAAGCTCGAAACAGACGAAGAGATCGAGAAGATATCTTCTAGTGAAG
AACGCCAATCATTCATTGAAAAGCTTGATGAAGTTCAAGACTGGCTATACATGGATGGTGAAGATGCAATTGCCAGTGAGTTTGAGAAACGTTTGGAGTC
CTTGAAAGTCATCGGTGACCCGATTTTTTTCAGACACAAGGAGCTCACAGATAGGCCAGCTGCAACGCAACTTGCTAGGAAATACCTCAGTGAGCTGAAA
CAGATTGTAGAAGGTTGGGAGGCAAACAAACCTTGGCTCCCTAAGGATAAAATAACCGAGGTTGTGAACGATGCGGAAAAGTTGAAGAGTTGGCTTGATG
AGAAGGAGGCTGAGCAGAAGAAGACTTCCGGATCCAGCAAGCCAGCCTTTACGTCTGAAGAAGTGTACACCAAAGTGTTCAAATTGCAAGACAAGGTTAC
TAGCATTAACAGAATTCCGAAGCCAAAGCCTAAAGTCGAGAAGCCCAAGAAGAACGAGACAGAGGAGCCTTCTAAGAAGGCTGAGACGGAAGAACCTAAG
AAGACAGATTCTGAGAAGAGCAGCGAGAGTTCGGAGAAAACCACTGGCTCTGATGGTGACAAAAATGGCGAAGATACTAAGCCACACGACGAGTTGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10039014 pacid=23150666 polypeptide=Lus10039014 locus=Lus10039014.g ID=Lus10039014.BGIv1.0 annot-version=v1.0
MDQHNRAPMTNEQHTRASHTPPRVPIATCPSFIGSSSSSQRNQPLRKSSASAYIYNNEKRFTSFPGKITAPASHSLSSSSFADFQLSSSAAQAEKNPPPT
ASGSYSVVTDPLMAELQGRRAADLGCLFLISRSMPTMLIKLGLLLCFISVNLIPSQSAVSSIDLGSEWMKVAVVNLKPGQSPIAIAINEMSKRKSPALVS
FHSGTRLLAEEAAGITARYPNKVFSQLRDMIGKPYKDVKVVLDSMYLPFEAVEDSRGSVAFKVDDGQVYSVEELLAMVLGYASNLAEVHAKLPVKDAVIS
VPPYFGQAERRGLIQAAQLAGINVLTLMNEHSGAALQYGIDKDFSNGSRYVIFYDMGSSSTYAALVYYSAYNTKEYGKTVSVNQFQVKDVRWNSQLGGQS
MEMRLVEYFADEFNKQVGNGVDVRKSPKSMAKLKKQVKRTKEILSANTAAPISVESLYDDRDFRSTITREKFEELCGDLWDRSLLPIKEVMKHSDLKVDD
ISAVELIGGATRVPKLQAKIQEFLGRNELDKHLDADEAIVLGGALHAANLSDGIKLNRKLGMIDGSPYGLSVVLDGPDMKDEYSRQSLVPRMKKVPSKMF
RSIVHNKDFEVSLEYETEEILPPSISSPVFAQYSVSGLTEASEKYSSRNLSSPVKASLHFSISRSGVLSLDRADAVIEISEWVEVPIKKNLTLETITNSS
SSSPETTENTTTSSTTESSSEESNENLQSDGAAGNESNPDVKEPITTEAATEKKLKKRTFRVPLKITEKTAGPGKPLSEESLAEAKKKLEELNKKDSERR
RTAESKNNLEGYIYSTKEKLETDEEIEKISSSEERQSFIEKLDEVQDWLYMDGEDAIASEFEKRLESLKVIGDPIFFRHKELTDRPAATQLARKYLSELK
QIVEGWEANKPWLPKDKITEVVNDAEKLKSWLDEKEAEQKKTSGSSKPAFTSEEVYTKVFKLQDKVTSINRIPKPKPKVEKPKKNETEEPSKKAETEEPK
KTDSEKSSESSEKTTGSDGDKNGEDTKPHDEL
|
DESeq2's median of ratios [FLAX]
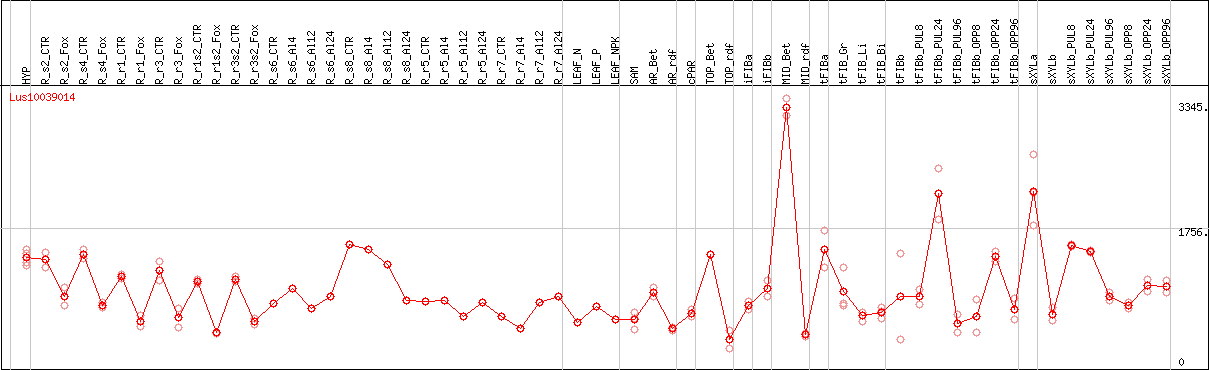
Coexpressed genes
Lus10039014 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
