Lus10039082 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10039082 pacid=23150606 polypeptide=Lus10039082 locus=Lus10039082.g ID=Lus10039082.BGIv1.0 annot-version=v1.0
ATGGCGGACAAGCTCGAGGGGCACCTTAAGGAAGTAGGATCCAAGCTAGACACTCCTCCTTCTTCCAAGGATGCTCTGCTTAAGCTTTTGAAGCAAGCTG
CTTCATGTCTCTCTGATATGGATCAATCGCCATCCCCGTCAATGTCAAAGTCCATGCAACCATTATTAACTGCAATTGTTAAGCCAGAATTGCTGCAGCA
TCAGGATAAGGACGTGAAGCTTCTAGTCGCAACCTGCATCTGTGAAATCACCCGGATTACTGCACCTGAAGCTCCTTACAGTGATGAAATTCTGAAGGTG
GATATATTTCAGTTGATTGTAAGCACCTTCAGTGGCTTAAGTGATACAACTGCGGGTCCATCTTTTGGTAGGAGAGCTCTCATTTTGGAGACTGTTGCCA
AATACCGGTCCTGTGTCGTGATGTTGGATCTTGAATGTGACGATCTCATTAAAAAGATGTTCAGTACATTTTTTTCTGTCACCAGGGATGACCATCAAGA
AAATGTACTTACGTCCATGCAAACCATCATGGTCGTTTTGATAGATGAAAGTGAAGAAGTTAATGATGATCTTTTACTTGTAATATTATCTGCTTTAGGC
CGTAATAAAAGTGATGTTTCTAAGGCTGCAAGGAAGCTTGCTATGAATATCATAGAACAGTGTGCAGTTCAGCTTGAAGCTGGCATCAAACGGATCCTTG
TTTCGTTAATGTCTGGAGATAACAGGTTGTCAGATTCTCAGATTGACTACCATGAAGTTATCTATGATATCTATTGTTGTGCTCCTCAGATATTGTCTGG
AGTGGTACCATACCTCACTGGAGAGTTGTTGACCGATCAGCTGGACACTCGTTTGAAAGCAGTAGGATTAGTTGGAGAACTTTTTTCTTTACCGGGATCC
ACAATCATCGAGACATTTCGGCCAATCTTTCTGGAATTCCTGAAAAGGTTGACGGACAGGGTAGTTGAAGTTAGAATGTCTGTTCTTGAGCATGTAAAGT
GCTGTTTATTGTCAAATCCTATTCGCACTGAGGCTTCTGAAATGATAGCTGCGCTATGTGAGCGATTGTTAGACTTTGATGAGAACATTAGGAGGCAGGT
CGTTGAGGTGCTCAGCAATATTGCTATTCATGCCCCTGGTTCTGTTCCTGCTGAAACTATAAAGCTGGTCTCCGATCGTCTAAGAGACAAATCCCTTACG
GTTAAAAAATGTACCATGGAGAGGATAGCTGAGATCTTCAGGGCTTATTGCAAGAAATCTTCAGATTCCTCCATCATTTCCCATGAATACGAATGGGTTC
CTGGGAAGATTTTGAGATGCTTTTACGATAAAGATTTCAGGTCAGATGCAATAGAGTATATACTCTCTGTTTCTTTATTCCCAAGCGATCTCTTGGTAAA
GGATAGAGTTACGCTTTGGGTCAGAATCTTCGCTACCTTTGACAAAATTGAACTGAAAGCTCTCGAGAAATTGCTGGAGCAAAAGCTAAGGTTACAACAA
GAGATGCAGAGGTACATGTCTCTTAGGTCAATGCATCAGGAGGGTGACGCACCAGAGATCCAAAAGAAGATTCTATTCTGCTTTCGGGTTATGTCTCGTT
CCTTCGCTGATCCTGCTAAGGCTGAGGAGAATTTTCTGATACTTGATCAACTGAAAGATACAAACGTCTGGAAACTTTTAGCTAGTCTGCTTGATGTAAA
TACTAGCTTGCAACAAGCTTGCACTAATCGGGACGAATTGCTCAAAATACTTGGTGAGAAACACGGGCTTTATGATTTTCTCAGCAGCCTGTCTGTCAAG
TGTTCGTATCTACTTTTCAGTAAGGAGTATGTTAAAGAATTTTTTTCGGAGGCTGTATCACATAAGTCTTCTGGGAATTCAGCATTGATTCAATCATCAA
TGAGTATATTGGTGACTCTTGCACGTTTTGGGCCATTGCTATTTGGGGGTGCTGAAGAAGAATTGATTAGCCTTTTAAAGGATGATAGTGAAATAATCAA
AGAAGGTGCATTACACGTTTTGGCTAAGGCCGGTGGCACTATCAGAGAGCAACTTGCAGTGTCCTCAAGTTCCATTGACCTCATATTGGAAAGGTTGTGC
CTAGAAGGTAGTCGAAGGCAGGCAAAATATGCTGTGCTTGCACTTGCAGCAATTACAAAGGATGATGGGCTCAAGTCTCTCTCTGTTCTATACAAGAAGC
TTGTTGGCATGCTGGAAGAGAAGAGAAATTTGCCTGCTGTTCTCCAGTCTCTGGGATGCATTGCACAAACTGCTATGTCAGTTTTTGAAACTAGAGAGAC
TGAAATTCAAGAATTTGTGAAAGACGGAATTCTGAATTGCAAGAGTCAGCCAGAAGATGATGCAAAAGTTTGTTGGGGCGATGTTAGTGAAACTAGTTTG
TTGAAGATATATGGGCTTAAGGTGTTTGTCAAAAGCCACTTACCTATTAAAGATGTGCACCTCCGTCCTGGTATTGGTGACCTTCTAGATATCCTGAAGA
ACCTACTGAAAGATGAAACATTACAGGGTACAGAATCAAGTTTGGTTGATAAAGCACATCTGAGGCTTGCTGCTGCGAAAGCAGTTCTTCGTCTGTCAAA
GCACTGGGATCATAAGATACCTGTTGATGTATTCCACTTGACTTTGAGAGTGGCTGAGAGTGCTTTCCCTGAAGCTAGGAAGATCTTTCTTGGCAAAGTT
CATCAGTATATAAAGGATCGCCTTCTTGACCCCAAGTATGCTTGTGCATTCTTGTTCTATCCGATAGAATCAGATCAGCTTGGTTTTGAGGAGGAAAAGC
AGAATCTCGTTGATATTATTCAAATGCATTGCCAGTCAAAGGCACGCCAAGTGTCTTTACAAAGTGATGCAAGTCCCTTAACAGCTTTTCCTGAATCGAT
TCTGCCATACGCGGTCCATTCACTGGCACATATCTCATGTCCTGACATAGACGAGTGTAAAGATGTTAAAGCATATGAGCCAATATATCGGCGACTATAC
TTGTTTCTTTCTTTGTTGATGCACAAAGAGGAAGAAGTAAAGTCAGAGGCTGACACCAATGCCAAAGATAGCATTTCTGTGATGACGTCTGTGTTTCAGG
CTATCAAGGACTCAGAAGATATCATGGATATGACAAAGTCTAAGAAGTCACGGGCTCTGTCTGAACTTGGATTGTCAATCGTCAAGCGCTTAGTTCCAAA
GGAGGATGACTTACAGATGTTGGCTTTACCAGTTACCCTGCCTTCCATGTTGTACAAGCTATATGAAAAGAAAGAAGAACAATCTCTGACAAATGAAGAG
AAGACATGGCTTGCGGAGGACAGTGTAATGACACATTTTCAGTCACTAAATTTTGAAACCGATGTAACACAAAAGGATCCTGCACCCGCTGCTGAGGAGG
AAGTTCGAAAAAATAGTGACAAAGAAGAGGATGATGCGCCTCTTGGTGAGATGGTAAAACAGTTGAAGGCCAAGGCAACAAAATCCGGAAGGAATAGAAG
GAATAAGAAAAAGAAATCTTCATCAGCTAAAGGCAAAAGCACGGAAAATGATTTTGATATTTTGAAGATGGTAAGGGAAATGGACGTGGATGATGTGGTA
GTTTCCGGTAAGCTTGAGTCAAGCAATGGTCATAAAGATGCTCCAAGTCAGAACCCGAAATCAGAACTGGAACACCAAAAGGCGGAGAAAAGAAAATCAA
TTGATGTAGCGTCTGTCCCGGTACCTAAGCGTCGGAGATCATCAACCATTCATACTGCTGCTAGATTGTCCCAGAGTCCAATGAAAGCTCTTTCAAGAGC
TTCAGCAGAAGATTCAAATCCTGATTTTAAAAACAAAGGCTCATTTCCAAAGAAGATGGGTAGAAGTGGTGACTCCGATTTTTTGGTATCATGTGTTAGA
AAGAGTAGAAATTCTTTACCAACGAAGGAATTCAAGATTTTTGATTTCGATGGTGATGATAGCGAGAATGACGCTAAACATTCCGATGAAGAAAATTGGA
CGAGCCCCAGTCTTCGCAGAAAGAACGACAAGAACAATGGAAAACATGTCGCAAAGCCTTCTACAGAATCTGCTTCGAAGAGGAAAAGAAAAAGTGTTTC
AGGAGGCCAAAAGCTCGCAAAGAACGATGGTGGAATAGATCTGGAAGCATTGATTGGTAATAGAATAAAGGTCTGGTGGCCAATGGACAAGGAGTTTTAT
GCTGGAACAATCAAGTCTTATGACCCTTTAAAGGGAAAGCATGTGATCTTATATGATGATGGAGATATAGAAGTTCTTCGTTTGGATAGAGAGCGTTGGG
AACTGGTTGACAAGAGTCAGAAGAATATGAAGAAGCTCAATTCGTCTAAGCGTGCTCCTTCTAAGGATGGGTCTGCCACACCGAAGAAAAGGAGTAAAGG
TGGTTCACAACAGAATAAGAGCTCCACCAAAAAAGCTAAAGCTAAAAGAACAAAGAAGAAAGCCTCAAAGCGAGTCCTGAAAGAGCCAGAAGATGATGAG
CCTTCTGACGGATCTGTTCCAGAACTTGCTGTTCCATCTTATGGAGAAGAAACAAAATCAGATGACTCTCAAGTTGAACGTGAGGAAGAAACCAAGACTC
TGAAGAAGGATGTAGAATCTGACAAGGAAGATACGTCAACTTCTGGAGGCAAACACTTGGAGGATGAAGCAGATAAGAAAAACTCCGATCATTCGGAGGA
TGATGATGGAGAAGGAGAACCAAGTTCTAAAGCAAAAGATTCTGACAACGAAAGTGAGAAACAAGAATCGGAGAAGAGTGGTGCTGCTGATGAGGGATCT
CATTCAGAAGAAAAGAAGGCAGAACAATCAGAAGATGAAGGCACATTAGAGGAAGCTGGTGATGGAGATGAATCAGGTTCAGAGGATAATGAAGATGCAG
ATGACGCCGAGGAGGATGAAAACGGTGCAAGTCCGGTTGCTATGGATGTGGATGATGAAATCCCAGACGATGAGCCTCTTAGCAAATGGAAGGGTAAATT
CGGGAAGAAGGCAGCATCAAAACGAGCAAAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10039082 pacid=23150606 polypeptide=Lus10039082 locus=Lus10039082.g ID=Lus10039082.BGIv1.0 annot-version=v1.0
MADKLEGHLKEVGSKLDTPPSSKDALLKLLKQAASCLSDMDQSPSPSMSKSMQPLLTAIVKPELLQHQDKDVKLLVATCICEITRITAPEAPYSDEILKV
DIFQLIVSTFSGLSDTTAGPSFGRRALILETVAKYRSCVVMLDLECDDLIKKMFSTFFSVTRDDHQENVLTSMQTIMVVLIDESEEVNDDLLLVILSALG
RNKSDVSKAARKLAMNIIEQCAVQLEAGIKRILVSLMSGDNRLSDSQIDYHEVIYDIYCCAPQILSGVVPYLTGELLTDQLDTRLKAVGLVGELFSLPGS
TIIETFRPIFLEFLKRLTDRVVEVRMSVLEHVKCCLLSNPIRTEASEMIAALCERLLDFDENIRRQVVEVLSNIAIHAPGSVPAETIKLVSDRLRDKSLT
VKKCTMERIAEIFRAYCKKSSDSSIISHEYEWVPGKILRCFYDKDFRSDAIEYILSVSLFPSDLLVKDRVTLWVRIFATFDKIELKALEKLLEQKLRLQQ
EMQRYMSLRSMHQEGDAPEIQKKILFCFRVMSRSFADPAKAEENFLILDQLKDTNVWKLLASLLDVNTSLQQACTNRDELLKILGEKHGLYDFLSSLSVK
CSYLLFSKEYVKEFFSEAVSHKSSGNSALIQSSMSILVTLARFGPLLFGGAEEELISLLKDDSEIIKEGALHVLAKAGGTIREQLAVSSSSIDLILERLC
LEGSRRQAKYAVLALAAITKDDGLKSLSVLYKKLVGMLEEKRNLPAVLQSLGCIAQTAMSVFETRETEIQEFVKDGILNCKSQPEDDAKVCWGDVSETSL
LKIYGLKVFVKSHLPIKDVHLRPGIGDLLDILKNLLKDETLQGTESSLVDKAHLRLAAAKAVLRLSKHWDHKIPVDVFHLTLRVAESAFPEARKIFLGKV
HQYIKDRLLDPKYACAFLFYPIESDQLGFEEEKQNLVDIIQMHCQSKARQVSLQSDASPLTAFPESILPYAVHSLAHISCPDIDECKDVKAYEPIYRRLY
LFLSLLMHKEEEVKSEADTNAKDSISVMTSVFQAIKDSEDIMDMTKSKKSRALSELGLSIVKRLVPKEDDLQMLALPVTLPSMLYKLYEKKEEQSLTNEE
KTWLAEDSVMTHFQSLNFETDVTQKDPAPAAEEEVRKNSDKEEDDAPLGEMVKQLKAKATKSGRNRRNKKKKSSSAKGKSTENDFDILKMVREMDVDDVV
VSGKLESSNGHKDAPSQNPKSELEHQKAEKRKSIDVASVPVPKRRRSSTIHTAARLSQSPMKALSRASAEDSNPDFKNKGSFPKKMGRSGDSDFLVSCVR
KSRNSLPTKEFKIFDFDGDDSENDAKHSDEENWTSPSLRRKNDKNNGKHVAKPSTESASKRKRKSVSGGQKLAKNDGGIDLEALIGNRIKVWWPMDKEFY
AGTIKSYDPLKGKHVILYDDGDIEVLRLDRERWELVDKSQKNMKKLNSSKRAPSKDGSATPKKRSKGGSQQNKSSTKKAKAKRTKKKASKRVLKEPEDDE
PSDGSVPELAVPSYGEETKSDDSQVEREEETKTLKKDVESDKEDTSTSGGKHLEDEADKKNSDHSEDDDGEGEPSSKAKDSDNESEKQESEKSGAADEGS
HSEEKKAEQSEDEGTLEEAGDGDESGSEDNEDADDAEEDENGASPVAMDVDDEIPDDEPLSKWKGKFGKKAASKRAK
|
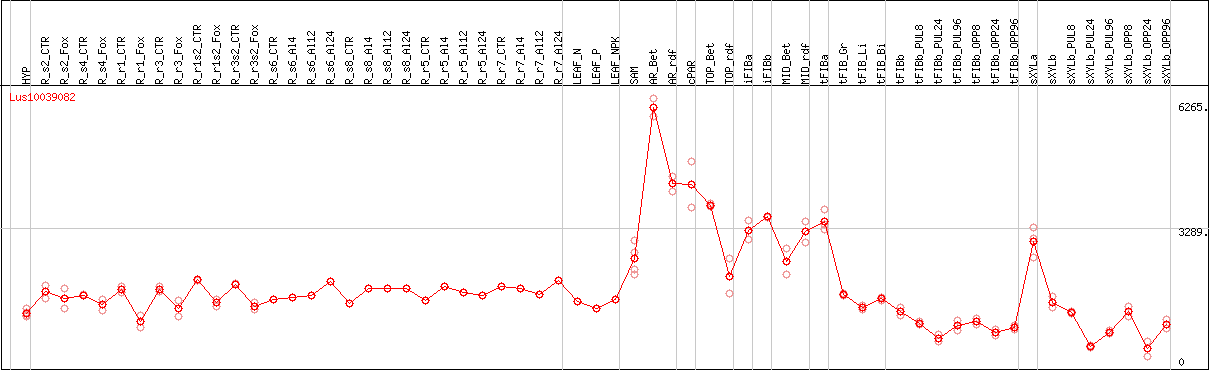
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039082 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
