Lus10039133 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10039133 pacid=23150595 polypeptide=Lus10039133 locus=Lus10039133.g ID=Lus10039133.BGIv1.0 annot-version=v1.0
ATGGCGAGTAGTATTAGTAGTAGTAGTAGCAGTATCATGGACCTTGACCATATCGGCTTCACAGTTTTGGTAGAGGAGGCCTCCCCGGAATCATCTTCAA
ATCCAACAGACGCCGTCATCTTCTTCGGCTTGTCTCTAGTGCTCGGGATCGCCAGCAGGCACGTCTTGCGCGGCACTCGTGTTCCTTACACCGTCGCTTT
ACTCATCATCGGCATTGTCCTCGGATCCTTAGAATATGGAACTAGTCATAAATTAGGGAAGATCGGAGATGGTATCCGACTTTGGGCGAGTATTGATCCT
GACCTTCTGTTGGCTATTTTCCTTCCTGCTCTGCTGTTTGAGAGTTCCTTCTCTATGGAACTCCATCAAATTAAGAGATGCATGGCGCAAATGCTTCTAC
TTGCTGGTCCAGGAGTTTTGCTCTCGACATTTTTCCTTGGATCGGCGTTGAAGCTCATCTTTCCCTATTCATGGAACTGGAAAACATCATTGTTACTTGG
GGGGCTGTTAAGTGCCACCGACCCTGTTGCTGTTGTTGCTCTTTTGAAAGAGCTCGGTGCAAGCAAAAAACTTAGTACGATTATTGAAGGAGAATCATTA
ATGAATGATGGTACAGCTATTGTGGTCTATCAGCTTTTCTATCGGATGGTTCTTGGACACAGTTATGGTGGGGGTGCGATAATTAAATTCTTGACACAAG
TGTCACTTGGAGCTGTAGGTATTGGTCTGGCTTTCGGGATCGCCTCTGTTTTGTGGCTTGGGTTTATTTTTAATGACACGGTGATTGAGATCTCACTCAC
GCTTGCCGTGAGCTATGTTGCTTACTTCACTGGACAAGAGGGCGCCGGTGTCTCGGGGGTTCTGACTGTGATGACTTTAGGAATGTTCTATGCTGCAGTT
GCTAAGACTGCTTTCAAGGGTGATGGTCAACAAAGCTTGCATCATTTTTGGGAAATGATTGCTTACATTGCGAATACTTTAATCTTCATCTTGAGTGGTG
TTGTTATAGCTGAAGGTGTTCTGAGCAGTGACAGCATTTTCCATAACCACGGACATTCTTGGGGATACCTTGTTCTTCTGTACGTTTTGGTCCAGCTTTC
TCGTTGTATAGTTGTTGGAGTTTTATTTCCTTTTCTGCGGTATTTTGGTTACGGGTTGGACTGGAAAGAGGCCACTATTCTCATATGGTCAGGTCTTCGA
GGAGCAGTTGCATTGTCGCTTTCTCTATCTGTTAAGGCAAGTTTCCTTTGTGAAATGTCTTACAATTTCATTTCAGCCTTTGCAAGGGAGGAGCAGAGGT
TGGGACGGCGAACTAGTGATAGCTCAGCTTTTATCAGCCCTGAGACGGGAAAACTGTTTGTTTTCTTCACTGGAGGAATTGTGTTTCTGACACTTATTGT
AAATGGATCAACTACCCAATTTGTTTTACGTCTTTTGGATATGGGCAAGCTGTCGGCTACCAAGAAACGTGTATTGGAGTATACAAGATATGAAATGTTG
AATAGAGCTCTAGAGGTTTTTGGTGATCTCGGAGATGACGAAGAACTTGGACCAGTGGACTGGCCCACAGTTAAAAGATATATTAAGACCTTGAGCAGTT
TGGAAGGAGAAAGTGTACATCCCCATGAACCATCTGAAACTGGTCTTGACCCTACAAATCTGAAAGATCTTCGTATACGCCTTCTTAATGGTGTTCAGGC
TGCCTACTGGGTGATGCTTGAAGAGGGAAGGATTACACAGGGCACCGCAAATATTTTGATGCAGTCTATAGATGAAGCGATGGACTCGGCAGCTCATGAG
CCTTTATCTGATTGGAAGGGCTTAAAATCAAATGTGAATTTTCCAAGCCACTACAAGTTCCTCCAGACAAGCTACTGCCCCCCAAAACTTGTTACTTATT
TCACTGTTGAGAGGCTGGAATCTGCATGTTATATCTGTGCTTCATTTCTGCGTGGCCACAGAATTGCACGGCAGCAGCTGCATGACTTTATAGGCGCCTG
GTTTCCTACTAGAAAAATGACAAAGGAGCATACTGTAGAAATCATATCAATTCTCCTTTTAATCTTAGTTCTTAGCTCAGAAGCTTTTTATTACATATTG
GCAGATCTAATTTGTCACTATTGTGTGATTAGCAGCTATGCTGAGATCTTATTCTATCATGCTCTTGTAGGCGACAGCAATATTGCTTCCATTGTCATTA
GTGAAAGCGAGGCAGAAGGAGAAGATGCAAGGAAGTTCTTGGAGGATGTTCGTGTTACATTTCCTCAGGTGTTACGTGTAGTGAAGACGAGACAAGTAAC
ATATTCAGTGCTTAACCACTTAATTGAGTATGTTCAAAATCTTGAGAAGACGGGGTTGTTAGAGCAGAAAGAGATGCTTCATCTTCATGATTCTGTGCAG
ACTGACTTGAAAAGGCTGCTGAGAAATCCTCCTCTTGTAAAGATACCCAAAATAAATGATTTGATCAATGTACACCCGCTGTTGGGTGCTCTCCCTCCTG
CCATCCGTGAACCACTAGCAGGTTCAAGCAAAGAAATAATGAAACTGCGTGGTGTTCCGATTTACAGAGAAGGCTCCAATCCCACTGGTGTGTGGTTCAT
TTCTAACGGCGTTGTCAAGTGGACAAGTAAGAGCATGAGGAGCAAGTACTCGCTTCATCCAACTTTCGCCCATGGTAGTACACTGGGACTGTACGAGGTG
CTAGTACGAAAGCCTTACATCTGTAGCATGGTAACAGATTCTGTGGTGCTTGGCTTTTTTGTTGAAAGTGAGAAAATACTATCAGCATTAAGCTCTGACC
CAGCAGTTGCAGATTTCTTCTGGCAGGAAAGCGCTATTGTGCTAGCAAAACTCTTACTTCCACAAATATTTGAGAAAATGTCAATGCATGATTTGAGAGC
TCTTATTGTTGAAAAGTCGACCATGACCACCTACATTAGGGGAGAAACCATAGAAATACCTACCCACTCCGTAGGGTTTCTTTTGGAAGGGTTTATAAAA
TCTCATGGTTTCCAAGAAGAATTGATTACTTCACCGGCAGCTTTATTTCCACAATATGGAGCAGCTTTATTTCCACAATATGGGGACCAAAGTAGCAAGA
CCTCTAGTTTTACTCATCAGGGGATGTGGTATCAAGTCGAGACGAGAGCAAGAGTGATCATATTCAAGACCGTATCATTTGAAGGGGAGGATGCCCTGAA
GAGAAGGTCATCGTCTTTGGTGTCAGTTGATAATCATATGCACAGGCCCCCAAGTAGAGAGCACGGTGGACTCATGAGTTGGCCTGAAAGCTTTTACAAG
CAGCAGAAGGAGAAAGACAACCACCAAACAACGGAAGAGCGGCCAGCCAACAGCTTATCGGCCAAAGCAATGCAGCTAAGCATGTTCGGCAGCATGGTTG
ACGTCCATGGTGGTCGGGCGCAAAGTTCTTCGAGCAGTGAAGGGAAAGGAGCTCGTAACATGGGGAAAGCAGCATCGTTCCAAGGCTACCCGAGTGGCGA
AGGGCCATCTTCGACTAGAGGAAGGAAGGTGGATGACGAAGGAGCCGCCATTGCTGCGGTGGAAAGTGGGGGCATCGGCACTGTGCAGAACTTGGAGGAG
AGGAGTGAGAGCGAAGATGATGATGACGATGAGGATGTTATCATAAGAATCGACTCACCAAGTACGCTGTCCTTCCGTCAGGTTTCTGGAGCAAAGCAGT
TAGGAGGAGGAGGAGAAGGCCGCGGCTTTGTATTCCAAGTTTGTCAGCCCACCACCGTCGCCTCGCCGGTCGCCGCCGCTAGTGTTGAGTGA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10039133 pacid=23150595 polypeptide=Lus10039133 locus=Lus10039133.g ID=Lus10039133.BGIv1.0 annot-version=v1.0
MASSISSSSSSIMDLDHIGFTVLVEEASPESSSNPTDAVIFFGLSLVLGIASRHVLRGTRVPYTVALLIIGIVLGSLEYGTSHKLGKIGDGIRLWASIDP
DLLLAIFLPALLFESSFSMELHQIKRCMAQMLLLAGPGVLLSTFFLGSALKLIFPYSWNWKTSLLLGGLLSATDPVAVVALLKELGASKKLSTIIEGESL
MNDGTAIVVYQLFYRMVLGHSYGGGAIIKFLTQVSLGAVGIGLAFGIASVLWLGFIFNDTVIEISLTLAVSYVAYFTGQEGAGVSGVLTVMTLGMFYAAV
AKTAFKGDGQQSLHHFWEMIAYIANTLIFILSGVVIAEGVLSSDSIFHNHGHSWGYLVLLYVLVQLSRCIVVGVLFPFLRYFGYGLDWKEATILIWSGLR
GAVALSLSLSVKASFLCEMSYNFISAFAREEQRLGRRTSDSSAFISPETGKLFVFFTGGIVFLTLIVNGSTTQFVLRLLDMGKLSATKKRVLEYTRYEML
NRALEVFGDLGDDEELGPVDWPTVKRYIKTLSSLEGESVHPHEPSETGLDPTNLKDLRIRLLNGVQAAYWVMLEEGRITQGTANILMQSIDEAMDSAAHE
PLSDWKGLKSNVNFPSHYKFLQTSYCPPKLVTYFTVERLESACYICASFLRGHRIARQQLHDFIGAWFPTRKMTKEHTVEIISILLLILVLSSEAFYYIL
ADLICHYCVISSYAEILFYHALVGDSNIASIVISESEAEGEDARKFLEDVRVTFPQVLRVVKTRQVTYSVLNHLIEYVQNLEKTGLLEQKEMLHLHDSVQ
TDLKRLLRNPPLVKIPKINDLINVHPLLGALPPAIREPLAGSSKEIMKLRGVPIYREGSNPTGVWFISNGVVKWTSKSMRSKYSLHPTFAHGSTLGLYEV
LVRKPYICSMVTDSVVLGFFVESEKILSALSSDPAVADFFWQESAIVLAKLLLPQIFEKMSMHDLRALIVEKSTMTTYIRGETIEIPTHSVGFLLEGFIK
SHGFQEELITSPAALFPQYGAALFPQYGDQSSKTSSFTHQGMWYQVETRARVIIFKTVSFEGEDALKRRSSSLVSVDNHMHRPPSREHGGLMSWPESFYK
QQKEKDNHQTTEERPANSLSAKAMQLSMFGSMVDVHGGRAQSSSSSEGKGARNMGKAASFQGYPSGEGPSSTRGRKVDDEGAAIAAVESGGIGTVQNLEE
RSESEDDDDDEDVIIRIDSPSTLSFRQVSGAKQLGGGGEGRGFVFQVCQPTTVASPVAAASVE
|
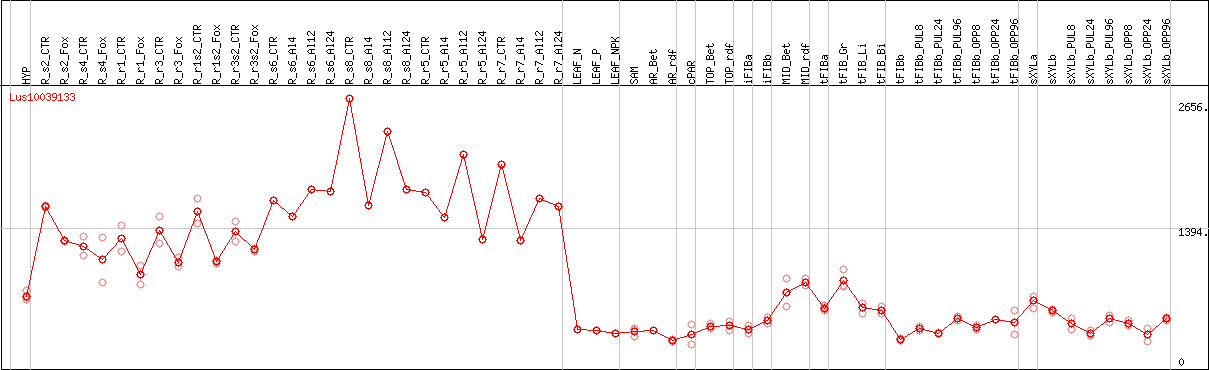
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039133 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
