External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G23940 206 / 2e-65
PEL3, DCR, EMB3009
PERMEABLE LEAVES3, EMBRYO DEFECTIVE 3009, DEFECTIVE IN CUTICULAR RIDGES, HXXXD-type acyl-transferase family protein (.1)
AT5G01210 104 / 6e-27
HXXXD-type acyl-transferase family protein (.1)
AT2G39980 102 / 4e-26
HXXXD-type acyl-transferase family protein (.1)
AT5G42830 93 / 9e-23
HXXXD-type acyl-transferase family protein (.1)
AT5G07850 87 / 9e-21
HXXXD-type acyl-transferase family protein (.1)
AT5G07860 82 / 6e-19
HXXXD-type acyl-transferase family protein (.1)
AT5G07870 80 / 3e-18
HXXXD-type acyl-transferase family protein (.1)
AT5G67150 76 / 1e-16
HXXXD-type acyl-transferase family protein (.1)
AT3G50270 64 / 1e-12
HXXXD-type acyl-transferase family protein (.1)
AT3G50280 60 / 4e-11
HXXXD-type acyl-transferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039256
293 / 4e-99
AT5G23940 576 / 0.0
PERMEABLE LEAVES3, EMBRYO DEFECTIVE 3009, DEFECTIVE IN CUTICULAR RIDGES, HXXXD-type acyl-transferase family protein (.1)
Lus10027500
288 / 2e-97
AT5G23940 571 / 0.0
PERMEABLE LEAVES3, EMBRYO DEFECTIVE 3009, DEFECTIVE IN CUTICULAR RIDGES, HXXXD-type acyl-transferase family protein (.1)
Lus10035519
108 / 2e-28
AT5G01210 520 / 0.0
HXXXD-type acyl-transferase family protein (.1)
Lus10040222
96 / 8e-24
AT2G39980 633 / 0.0
HXXXD-type acyl-transferase family protein (.1)
Lus10028267
96 / 8e-24
AT2G39980 631 / 0.0
HXXXD-type acyl-transferase family protein (.1)
Lus10031149
85 / 8e-20
AT3G50280 305 / 3e-99
HXXXD-type acyl-transferase family protein (.1)
Lus10024809
80 / 5e-18
AT5G42830 442 / 8e-153
HXXXD-type acyl-transferase family protein (.1)
Lus10027778
66 / 4e-15
AT5G01210 141 / 6e-42
HXXXD-type acyl-transferase family protein (.1)
Lus10031148
67 / 2e-13
AT5G67150 284 / 5e-91
HXXXD-type acyl-transferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0149
CoA-acyltrans
PF02458
Transferase
Transferase family
Representative CDS sequence
>Lus10039255 pacid=23175545 polypeptide=Lus10039255 locus=Lus10039255.g ID=Lus10039255.BGIv1.0 annot-version=v1.0
ATGGCGGACAGCTACTTCGGAAACCTGATCCAGGCGATATTCACCGTCCTACCGGCCGGAATGCTCATCAGCAACCCGCCGGGGTTCGGGGCGTCGATGA
TACAGGGGGTGATCGAGAAGCACGACGCGAAGGCGATCGAAGGGAGGAACGCGGAGTGGGAGGCGGCGCCGAAGATATTCCAGTTCAAGGACGCCGGAGT
GAACTGCGTGGCGGTGGGGAGCTCGCCGAGGTTCAAGGTCTACGACGTGGATTTCGGATTCGGGAAGCCCGAGAGTGTCCGGAGCGGGTTGAATAACCGG
TTCGACGGGATGGTGTATTTGTACCAAGGGAAGAACGGAGGGATCGATGTGGAGATTAGCTTGGAGAGTGCGGCGATGGAGAAGCTGGAGAAGGATAAGG
AGTTTCTTATGGAGGGAGTTGACTGTAGTTTGGCCGCTTGA
AA sequence
>Lus10039255 pacid=23175545 polypeptide=Lus10039255 locus=Lus10039255.g ID=Lus10039255.BGIv1.0 annot-version=v1.0
MADSYFGNLIQAIFTVLPAGMLISNPPGFGASMIQGVIEKHDAKAIEGRNAEWEAAPKIFQFKDAGVNCVAVGSSPRFKVYDVDFGFGKPESVRSGLNNR
FDGMVYLYQGKNGGIDVEISLESAAMEKLEKDKEFLMEGVDCSLAA
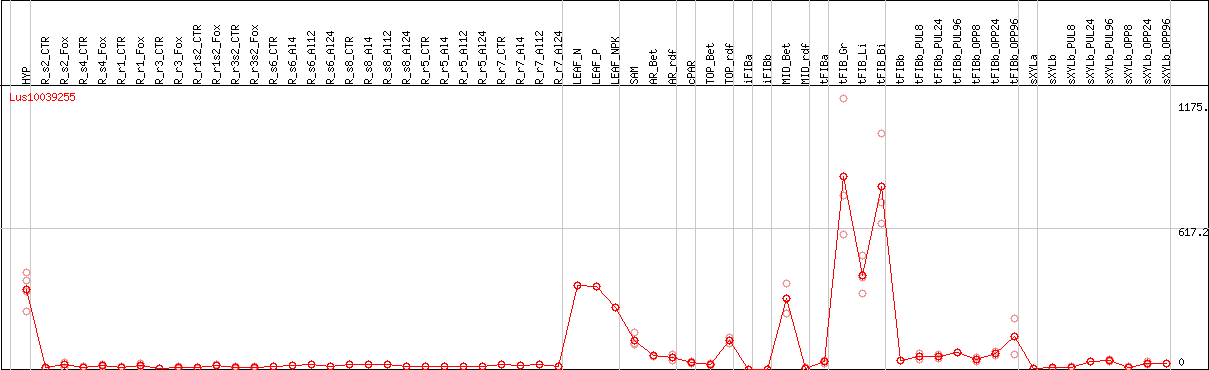
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039255 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.