Lus10039491 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10039491 pacid=23164995 polypeptide=Lus10039491 locus=Lus10039491.g ID=Lus10039491.BGIv1.0 annot-version=v1.0
ATGGCCGGCAAGTGTAGGCATCGGGTAGCTGTGAAGAAAGTGGAGGTTGGGGAGGGGGTGGAAGTGGACTGGGTGCAAGCTCAGCTGGAAAACTTGCGGC
GTGCTTCAATGTGGTGTAGAAACGTGTGTACTTTCCATGGTGTTGTGAAAGTGGATGGGTGTTTAGGGCTTGTAATGGATAAGTATTGTGGCTCTGTTCA
GTCTGAGATGCAGCAGAATGAAGGAAGGCTCACCCTGGAGCAAATTCTAAGATATGGGGCTGACATAGCACGTGGGGTTGCTGAACTTCATGCAGCAGGT
GTTGTCTGCATGAACTTAAAACCTTCCAATCTTCTCCTGGATTCAAGTGGCCGTGCTGTTGTTTCTGATTATGGACTTGCTTCCATTTTAAAAAAACCAA
CTTGCCGCAAAGCTAAAACAGAATGTGCTTCTGCCAAAATTCATTCATGCATGGACTGCATAATGCTCAGTCCACACTACACGGCACCAGAGGCTTGGGA
GCCGGTCAAGAAGTCTCTGAATCTTTTCTGGGATGATGCAATCGGCATATCTGTGGAATCAGATGCCTGGAGCTTTGGATGTACATTGGTAGAGATGTGC
ACCGGTGCTACTCCGTGGGCTGGTCTGAGTGCAGAAGAAGTTTACTGTGCTGTTGTTAAAGCTCGAAAATTGCCTCCACAGTATGCCAGTGTGGTTGGTG
TTGGAATTCCTAGAGAATTATGGAAAATGGTCGGTGAATGCTTACAGTTCAAACCATCAAGGAGGCCAACGTTTAATGCAATGTTAGCGATATTCATCCG
TCATCTGCAGGAGCTGCCACGTACCCCTCCTGCAAGTCCTGATAATGACTATGTACAAAAACCTGGTTCAAATGTAGCTGAACCATTACCACCATCTGAT
TTGGAGGATTTGCCAGACAACTCCAGCCATCTACATAGACTCGTCTCGGAGGGAAATGTAATTGGTGTTAGAGATCTACTAGCAAAGGCTGCATCAGGAA
GTAGTGGTGATTCTGTATACAGACTGCTAGAAGCACAGAATGCTGATGGCCAAACAGCACTTCACCTGGCTTGTAGACGTGGGTGTGCAGAACTCGTTCA
GACCATACTGGAGTATAGAGAAGCAAATGTTGATGTCCTGGACAAGGATGGTGATCCACCACTTGTTTTTGCTTTAGCTGCTGGATCACCAGAATGTGTT
CATGCTCTAATTAGGAGCGGGGCTAATGTCAGATCCCGATTAAGGGAAGGGTTTGGTCCATCGGTTGCTCATGTTTGTGCCTACCATGGCCAGCCAGATT
GTATGCGTGAGCTTTTACTTGCTGGAGCGGATCCAAATGCTGTGGATGATGAAGGGGAATCTGTACTACATAGAGCTCTATCAAAGAAGTATACTGATTG
TGCTATTGCTATACTGGAAAATGGAGGCTGCAGATCAATGGCTTTTGTGAATTCAAAAAATTTGACACCATTGCACTTGAGTGTAGCAACATGGAGTGTG
GCTGTTGTCAGAAGGTGGGTAGAAGTCGCCTCCCAAGATGACATTGCTGAGGCAATTAATATTCCAAGCCCAGTTGGAACTGCCTTATGTATGGCTGCTA
GTTCCAAGAAAGATCATGAAATTGAAGGGAGACAATTAGTGCAAATTTTGCTCACTGCTGGAGCAGATCCAACTTCTCAGGATGCACCTCATGGGAGAAC
AGCCTTGCACACAGCTTCTATGACTAATGATGTTGAGTTAGTGAAGATAATTCTGGATGCTGGAGTTGATGTGAACATCCGGAACATGCATAACACCATA
CCTCTCCATGTAGCTCTGGCCAGAGGAGCAAAATCATGTGTTGGCATACTCTTAAATGCTGGGGCAAATTGCAACATCCAGGATGATGAAGGTGATACTG
CCTTTCACATAGCAGCTGATGCTGCAAAAATGATACGGGAAAATCTTGAATGGCTGATCGTTATGCTTGGGAATCCGAATGCAGCCGTTGAAGCAAGAAA
TCACAGGCTAGTGGCTTTAGATGATTCAGAAAATGAATGGCTGAACTCTACTGTGCTTTATGTACTTGTTGGTAACAGTGGTAAGACATTACGCGATTAT
TTGGAAACTCTGCCACGGGAATGGATTTCTGAAGACCTAATGGAGGCTCTCTTGAAGAGGGGAGTGCATTTATCTCCTACGGTATATGAAGTTGGAGATT
GGGTGAAGTTCAAGAGAAGTTTGACCACTCCTGCAAAGGGATGGCAAGCTGCAAAACACAAGAGTGTAGGGTTTGTTCAAACAGTTGTTGACAAGGACAA
TCTAGTCGTCTCATTCTGTACTGGGGAGGCTCACGTTTTAGCCAGTGAAGTATGGAAGGTGATTCCCTTGGATCGTGGTCAGCATGTCAGGCTCAAAGCA
GAGATAAAAGAGCCAAGGTTTGGATGGCGTGGCCAATTACGAGACAGCATTGGAACTGTCCTGTGTGTTGATGATGATGGCATTTTACGTGTTGGGTTTC
CTGGAGCGTCTCGAGGGTGGAAAGCTGACCCCGCAGAAATGGAGAGAGTTGAAGAATTCAAGGTTGGTGACTGGGTTCGTATTCGCCCCACACTAACAAC
AGCCAAGCATGGATTAGGATCTGTAACACCAGGAAGCATTGGCATCGTGTATTGCATTAGACCAGATGGTAGTCTTTATTTAGAATTGAGTTATCTTCCA
CAGCCATGGCATTGTGAACCAGAGGAGGGTGAAGCTGTTACTCCTTTCAGGATTGGTGACCGTGTATGTGTAAAGCGTTCTGTTGCTGAACCTAGATATG
CTTGGGGCGGTGAGACCCATCACAGTGTAGGAAGGGTTAGCGAGATAGAAAACGATGGCTTGTTAATAATTGAAATACCAAATCGACCTATTCCATGGCA
AGCTGATCCATCTGACATGGAAAAAGTAGAGGATTTCAAGGTTGGAGATTGGGTCAAGGTAAAAGCTTCAGTTTCTTCTCCTATCTATGGGTGGGAGGAT
GTTTCACGGAACAGCATCGGTGTGATTCACAGTTTAGAAGATGACGGCAATATTGGTGTTGCCTTCTGTTTCAGAAGCAAGCCCTTCTGCTGTTCTGTGA
CAGATGTGGAAAAGGTTCTTCCTTTTGAGCTAGGACAGGAAGTGCGTGTGATGGCTTCCGTTACTCAGCCACGACTAGGGTGGTCAAATGAAAGTCCTGC
AACTGTTGGGAAAATTGCTAAAATTGACATGGATGGAACTTTGAACGTGTTGGTTGCTGGCAGACGTGGTTTATGGAAAATGTCTCCTGGAGATGCAGAA
CGGCTTTCCGGATTTGAAGTTGGTGATTGGGTACGCTCCAAACCGAGCCTAGGGACCAGGCCTAGTTATGATTGGAATACTATAGGAAAGGAAAGCTTAG
CTGTTGTGCATAGTGTGCAAGAGACCGGTTACCTGGATCTAGCTTGCTGTTTCCGAAAGGGAAGGTGGAGTACTCATTACACGGATGTCGAAAAGGTTCC
AAGCATGAAAGTAGGGCAGCATGTACGGTTTCGGACTGGAATTGTTGAGCCAAGGTGGGGCTGGAGGGGTGCTCAGCTTGATTCACGTGGCATAATAACT
AGTGTTCATGCTGATGGAGAAGTCAGAGTTGCTTTCTTTGGACTGTCAGGGTTATGGAGAGGCGATCCTGCAGATCTGGAGATAGAGCGAATGTTTGAAG
TGGGTGAATGGGTGAGGTTAGCGAAAGATGCAAGCAACTGGAAGTCAATTGCTCCAGGATGCGTTGGTGTTGTACAGGGTATAGGATATGATGGGGATAA
ATGGGACGGAAGTACTTACGTGGGGTTTTGTGGCGAACAAGAAAGATGGGTGGGCCCTACTTCTCATCTCGAAAGAGTGGAGAGGCTTGTCGTCGGCCAA
AAAGTTAGGGTTAAACTCTCTGTAAAACAACCTAGGTTTGGATGGTCAGGTCATAATCATGGAAGTCTTGGAACAATAGCAGCTATTGATGCTGATGGGA
AGTTGAGAATGTACACTCCGGCAGGGTCGAAAACTTGGATGCTTGACCCATCCGAAGTCGAACTAGTGGAGGAAGATGAACTCGGAATAGGGGACTGGGT
GAGAGTTAGGAAATCAGTTGCAACGCCTACTCACCAATGGGGAGAGGTAAATCATTCAAGCATCGGAGTAGTGCATCGGATGCAAGGTGGGGAGCTATGG
GTTGCATTTTGTTTCATGGAGAGGCTATGGCTTTGTAAGGCATGGGAAATGGAAAGGCTAAGACCATTTGAAATAGGGGACAAGGTAAGGATCCGAGACG
GGCTGATAACACCGCGGTGGGGATGGGGAATGGAGACACACACGAGTAAAGGTGAGGTAGTTGGGGTGGATGCAAATGGAAAGTTGAGGATAAAGTTCCA
GTGGAGAGATGGGAGGCCATGGATTGGTGATCCTGCTGATATTGTTGTAGACGAAAGCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10039491 pacid=23164995 polypeptide=Lus10039491 locus=Lus10039491.g ID=Lus10039491.BGIv1.0 annot-version=v1.0
MAGKCRHRVAVKKVEVGEGVEVDWVQAQLENLRRASMWCRNVCTFHGVVKVDGCLGLVMDKYCGSVQSEMQQNEGRLTLEQILRYGADIARGVAELHAAG
VVCMNLKPSNLLLDSSGRAVVSDYGLASILKKPTCRKAKTECASAKIHSCMDCIMLSPHYTAPEAWEPVKKSLNLFWDDAIGISVESDAWSFGCTLVEMC
TGATPWAGLSAEEVYCAVVKARKLPPQYASVVGVGIPRELWKMVGECLQFKPSRRPTFNAMLAIFIRHLQELPRTPPASPDNDYVQKPGSNVAEPLPPSD
LEDLPDNSSHLHRLVSEGNVIGVRDLLAKAASGSSGDSVYRLLEAQNADGQTALHLACRRGCAELVQTILEYREANVDVLDKDGDPPLVFALAAGSPECV
HALIRSGANVRSRLREGFGPSVAHVCAYHGQPDCMRELLLAGADPNAVDDEGESVLHRALSKKYTDCAIAILENGGCRSMAFVNSKNLTPLHLSVATWSV
AVVRRWVEVASQDDIAEAINIPSPVGTALCMAASSKKDHEIEGRQLVQILLTAGADPTSQDAPHGRTALHTASMTNDVELVKIILDAGVDVNIRNMHNTI
PLHVALARGAKSCVGILLNAGANCNIQDDEGDTAFHIAADAAKMIRENLEWLIVMLGNPNAAVEARNHRLVALDDSENEWLNSTVLYVLVGNSGKTLRDY
LETLPREWISEDLMEALLKRGVHLSPTVYEVGDWVKFKRSLTTPAKGWQAAKHKSVGFVQTVVDKDNLVVSFCTGEAHVLASEVWKVIPLDRGQHVRLKA
EIKEPRFGWRGQLRDSIGTVLCVDDDGILRVGFPGASRGWKADPAEMERVEEFKVGDWVRIRPTLTTAKHGLGSVTPGSIGIVYCIRPDGSLYLELSYLP
QPWHCEPEEGEAVTPFRIGDRVCVKRSVAEPRYAWGGETHHSVGRVSEIENDGLLIIEIPNRPIPWQADPSDMEKVEDFKVGDWVKVKASVSSPIYGWED
VSRNSIGVIHSLEDDGNIGVAFCFRSKPFCCSVTDVEKVLPFELGQEVRVMASVTQPRLGWSNESPATVGKIAKIDMDGTLNVLVAGRRGLWKMSPGDAE
RLSGFEVGDWVRSKPSLGTRPSYDWNTIGKESLAVVHSVQETGYLDLACCFRKGRWSTHYTDVEKVPSMKVGQHVRFRTGIVEPRWGWRGAQLDSRGIIT
SVHADGEVRVAFFGLSGLWRGDPADLEIERMFEVGEWVRLAKDASNWKSIAPGCVGVVQGIGYDGDKWDGSTYVGFCGEQERWVGPTSHLERVERLVVGQ
KVRVKLSVKQPRFGWSGHNHGSLGTIAAIDADGKLRMYTPAGSKTWMLDPSEVELVEEDELGIGDWVRVRKSVATPTHQWGEVNHSSIGVVHRMQGGELW
VAFCFMERLWLCKAWEMERLRPFEIGDKVRIRDGLITPRWGWGMETHTSKGEVVGVDANGKLRIKFQWRDGRPWIGDPADIVVDES
|
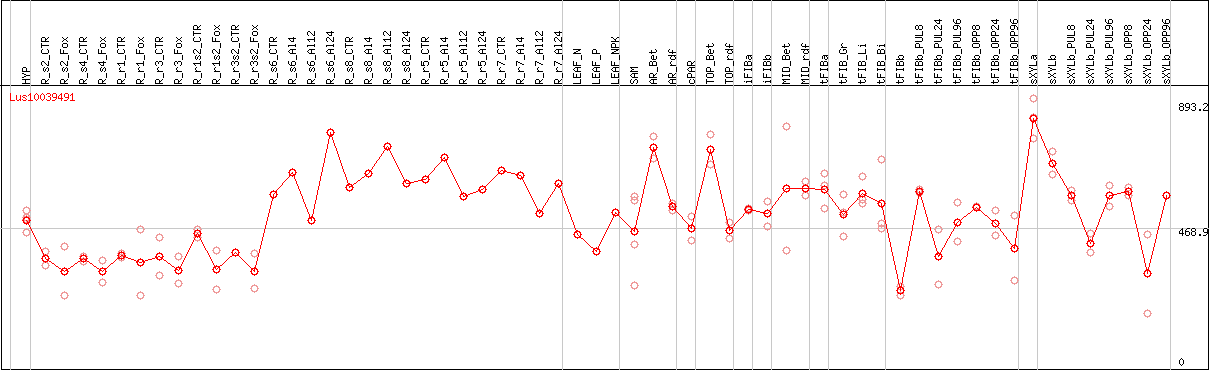
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039491 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
