Lus10039514 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10039514 pacid=23165133 polypeptide=Lus10039514 locus=Lus10039514.g ID=Lus10039514.BGIv1.0 annot-version=v1.0
ATGAATGCATTGACTACTGTTCTTAAAGTTCTCTTGGTCTCATTGGTCTGTATGGGTCTCTGGGACGAGACATTCGGTGCAGGCTGTAGTAAATATGAAA
GAGAAGCACTTTTGATGTTCAGGGCTGATCTCGAAGACCCTTTCAGGTGGCTATCATCTTGGGTTGGCCAAGGAGATTGTTGCACATGGTACGGTGTTGT
CTGCGACAATTTTACTGGCCATGTCACTGAGCTCCATCTCGGGTGGCCATCTTCCTCGGAGAAATTGGGTTTTACTGGTAAGATAAATCCTTCCTTGCTT
GAGTTGAAGTATCTCACTTATTTGTGCTTGTCAAACAACGACTTTCAAGGGACTCCCATTCCGAGTTTTCTTGGGTCTATGAGCAATTTAACTTCCCTTT
ACCTCAACGAAGCTGGATTTGGAGGAGTTATTCCCCATCAATTTGGGAACCTCACAAATCTGAAACAGCTTGGTTTGGAAGCTGCTGACTATGTGGTTTA
TGCTGATAGTTTGCAGTGGCTTTCTGGTCTGCATTCATTGAATTACCTTGATTTGAGTAGTGTTGATCTTAGCAAGGCCTCTAACTGGTTGGAGATGATC
AACACTCTGCCATCGCTTTCGACATTGTATCTTTCAAACTGCCAAATCCAGCATAGCCTTCCACTCGTAAATGTCAACCTGTCATCATCACTCTCTGTTT
TGGATTTGTATCACAATAAATTTGGCCAAACTTCAATTATTACTAGTTGGGTTTCTCATCTCAAGGGCCTTACTTATTTGAATCTAGGAAACAACAATTT
TGGCAGTCCGATTCCTTGCGAACTTCGAAACCTTACCTCTCTCAGGAGCCTTTACTTCGATGGCAACCATTTCAATCATTCAATACCCAACTGGTTGTAT
GGTTTACACTATCTCGAGTACCTCAATCTACGTGGAAATAAGTTAGAGGGTACAGTCTCACATTCCATTGGGAACTTGTCTTCTCTTGTGAAACTTGACC
TGTCATTGAATCCTGGCCTGGAATTCGAAATGGGGATTCCGGATTCTTTCAAACAGTTCTGTCATTTGAGAGCACTTTTTCTGGAATATGTTAAGGTAAA
CCAAAGTTTGTCAGAAGTTCTTGATATCCTGTCAGAATGTGCTTCGGATTCTCTAGAGGTATTATCTATGTATAGTAGCCAACTGTTTGGTCAGTTGACG
AACCGGATCGGAAAGTTCAACAGATTGGAACGTTTGGTACTCGATGGTAATTTCATTTCTGGTCCACTTCCAGTTTCTATAGGGGAATTGAAATCCTTGA
ATTATGTGAACCTCGCAAACAACCAGATAAATGGTACTATTCCAGCGAGTTTCGGGTTGCTGGCAGAGTTGGAATCTGTTGATATTTCTCAGAACTCGAT
GCAGGGCGATGTCTTCGCTGAAATCCAGTTTGCCAGCCTCACAAAGTTGGGTATGTTTCTTGCATCCGGAAACCAAATGGTGTTGAAAGATGGACCAAAT
TGGTTTCCTTCGAAAGGACTTACAATACTGGACCTGGATTCCTGGTACGTTGGACCCGGCTTTCCTAATTGGCTCAAGAATATGGGGAGTCTGAAGTCTC
TATCTCTATCAAACTCTGGAATTTCAGAACCAATTCCGGATTGGTTCTGGAGCATGTCGTCTGGATTTTACTATACGAATCTGTCTCACAACCAAATCCC
AGGAAGGCTTCCACATGTCATCTCTTCTCTTGTTGAATATGACACATTGTTTGATTTAAGCTTCAACAGCTTGGAAGGTCCGTTGGCTTCGATTTCGTCC
AACTTGACTTTGTTCGACATGTCCAACAATTTACTCTCTGGAGACTTGGTCAAGTTCCTATGTTTCAATCCGACTCAGATGAGGAATACACAATATCTCA
GCCTTGGAGGTAATCTGTTGTCTGGAAATTTACCTGATTGCTTTAGTAATTGGAGGAGTTTGAAGGTGCTGAGACTAGGAAACAATAAGTTTACAGGAAG
TATTCCAAAGTCAATTGGAACTTTGAGCTCACTTAAGTCTTTGCACATTCAGAACAATAGCTTGACTCGTGAAATTCCCCTTTCCATTGGAAAAAAATGC
TCTGCTTTGGTCACAATTAACTTGAGTTATAATCAATTGGAAGGAGAGATTCCACAGTGGGGTGGGGTGAGCCTTTCTAGACTGTGTATACTAAATCTGC
GTGGAAATCGGTTTCGTGGAGTCTTACCGAATGAGCTTTGCAACCTGCAGTCTTTACAAATCCTAGACCTTTCTCACAACATGCTTTCTGGCAGTTTGCC
TCAATGTCTGGCCAATTTAAGTGCAATGGCTGTACCTACGAATGACCGGCTTTCCCAAATTTATCTGTTCTTCGGTGGGGGAGTTGTGTACAGCGAGAAT
CAAGTCATTGTGACGAAAGGGAGCTTTGCTGAATACAGCACCATACTTAACTATGTCAGAAGTTTAGACCTTTCCTGCAATAACTTATCCGGAGTAGTTC
CCTGGCAGATTTCAGACCTGATGGGATTGCAGTACCTAAATTTGTCGCATAACTCACTGTCGGGAATGATTCCTGAAGGTTTAGGAAAAATGAGAGCACT
GGAATCCATGGACTTGTCTGTAAACCGGCTATCGGGGGTAATACCGGGTGGAGTTGCGAGCCTGACATTTTTGAGTCATTTGAACCTGTCTTTCAACAGG
TTGAGTGGGAGGATTCCTTCGAGTACTCAGCTGCAGAGTTTCGATACATCCAGCTTTGTCGGTAACAAAGAACTATGTGGTCCTCCATTGAATGAGGGTT
GTAGAAACACAGTGGGTGATGGGTTTGCTGGTAATGGTGGTGTTAAAGGAGATGAAGAATATTATTATGCATTCCATTGCTTCTGTGCTCTTGGGTTTGG
AGCGGGGTTCGGATTTAGTGTTTTCATTTTTCAATATATCGCTGATGAAGAACATACTTTACAAAGGGTGGATCACGTGGGACACAAGTTATGCAACTTT
CTCATGTTTGTACGTAGTCCTCAAGGAACCGGATCGGATCTTCCAGGTACAGGAATTAGGATGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10039514 pacid=23165133 polypeptide=Lus10039514 locus=Lus10039514.g ID=Lus10039514.BGIv1.0 annot-version=v1.0
MNALTTVLKVLLVSLVCMGLWDETFGAGCSKYEREALLMFRADLEDPFRWLSSWVGQGDCCTWYGVVCDNFTGHVTELHLGWPSSSEKLGFTGKINPSLL
ELKYLTYLCLSNNDFQGTPIPSFLGSMSNLTSLYLNEAGFGGVIPHQFGNLTNLKQLGLEAADYVVYADSLQWLSGLHSLNYLDLSSVDLSKASNWLEMI
NTLPSLSTLYLSNCQIQHSLPLVNVNLSSSLSVLDLYHNKFGQTSIITSWVSHLKGLTYLNLGNNNFGSPIPCELRNLTSLRSLYFDGNHFNHSIPNWLY
GLHYLEYLNLRGNKLEGTVSHSIGNLSSLVKLDLSLNPGLEFEMGIPDSFKQFCHLRALFLEYVKVNQSLSEVLDILSECASDSLEVLSMYSSQLFGQLT
NRIGKFNRLERLVLDGNFISGPLPVSIGELKSLNYVNLANNQINGTIPASFGLLAELESVDISQNSMQGDVFAEIQFASLTKLGMFLASGNQMVLKDGPN
WFPSKGLTILDLDSWYVGPGFPNWLKNMGSLKSLSLSNSGISEPIPDWFWSMSSGFYYTNLSHNQIPGRLPHVISSLVEYDTLFDLSFNSLEGPLASISS
NLTLFDMSNNLLSGDLVKFLCFNPTQMRNTQYLSLGGNLLSGNLPDCFSNWRSLKVLRLGNNKFTGSIPKSIGTLSSLKSLHIQNNSLTREIPLSIGKKC
SALVTINLSYNQLEGEIPQWGGVSLSRLCILNLRGNRFRGVLPNELCNLQSLQILDLSHNMLSGSLPQCLANLSAMAVPTNDRLSQIYLFFGGGVVYSEN
QVIVTKGSFAEYSTILNYVRSLDLSCNNLSGVVPWQISDLMGLQYLNLSHNSLSGMIPEGLGKMRALESMDLSVNRLSGVIPGGVASLTFLSHLNLSFNR
LSGRIPSSTQLQSFDTSSFVGNKELCGPPLNEGCRNTVGDGFAGNGGVKGDEEYYYAFHCFCALGFGAGFGFSVFIFQYIADEEHTLQRVDHVGHKLCNF
LMFVRSPQGTGSDLPGTGIRM
|
DESeq2's median of ratios [FLAX]
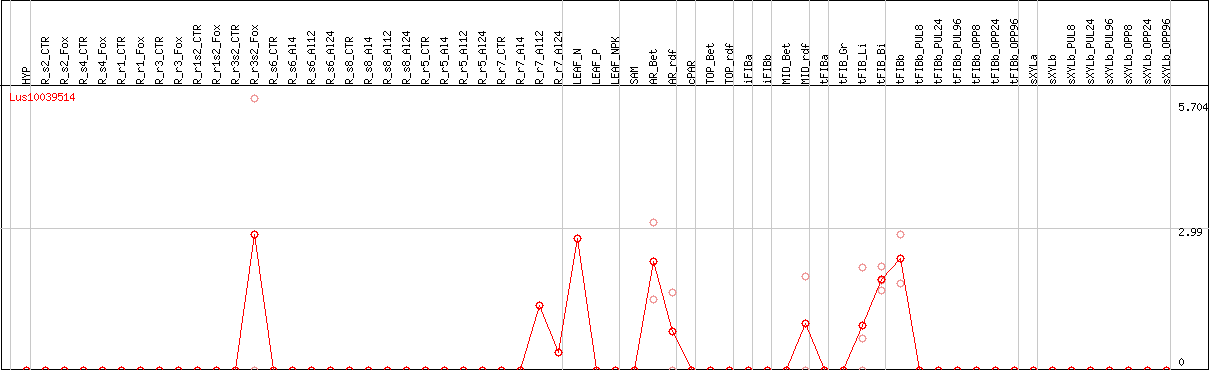
Coexpressed genes
Lus10039514 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
