External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G55790 169 / 3e-49
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
AT5G56060 156 / 5e-47
Domain of unknown function (DUF2431) (.1)
AT5G56075 141 / 7e-41
Domain of unknown function (DUF2431) (.1)
AT5G25030 129 / 4e-37
Domain of unknown function (DUF2431) (.1)
AT4G26485 120 / 1e-33
Domain of unknown function (DUF2431) (.1)
AT1G55800 118 / 1e-31
Domain of unknown function (DUF2431) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000235
269 / 9e-90
AT1G55790 176 / 7e-51
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Lus10002759
269 / 9e-90
AT1G55790 176 / 7e-51
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Lus10016345
257 / 6e-85
AT1G55790 183 / 2e-53
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Lus10032900
170 / 3e-51
AT1G55790 256 / 1e-81
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Lus10032898
165 / 2e-49
AT1G55790 232 / 2e-72
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Lus10032901
108 / 8e-28
AT1G55790 196 / 1e-58
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Lus10016346
76 / 2e-16
AT1G55800 72 / 8e-15
Domain of unknown function (DUF2431) (.1)
Lus10032899
44 / 6e-05
AT1G55790 62 / 2e-21
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G002700
180 / 3e-56
AT1G55790 185 / 1e-55
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Potri.001G438100
172 / 2e-53
AT1G55790 207 / 4e-64
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Potri.003G101200
176 / 2e-52
AT1G55790 276 / 8e-88
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Potri.011G141700
169 / 3e-52
AT1G55790 198 / 1e-60
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Potri.011G141900
164 / 6e-50
AT1G55790 219 / 7e-69
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Potri.011G165600
160 / 5e-47
AT1G55790 275 / 2e-88
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Potri.001G438200
140 / 4e-40
AT1G55790 169 / 1e-48
Domain of unknown function (DUF2431) (.1), Domain of unknown function (DUF2431) (.2)
Potri.011G141501
90 / 2e-22
AT4G26485 102 / 2e-28
Domain of unknown function (DUF2431) (.1)
Potri.016G003133
49 / 8e-08
AT1G55800 47 / 1e-07
Domain of unknown function (DUF2431) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF10354
DUF2431
Domain of unknown function (DUF2431)
Representative CDS sequence
>Lus10039534 pacid=23165253 polypeptide=Lus10039534 locus=Lus10039534.g ID=Lus10039534.BGIv1.0 annot-version=v1.0
ATGAAGAAGAAAAAGGGTTCATTTGGGAAGAAACTCATATCCTACCTACTTGGATGCATATGCTCCAACCCTGAAAAGGAAGAAAGATATTTCCACCAGG
AGTTTCAGCTTGCTTCTTCAGCTTCCGCTTCCTCCTGTGATGACTCGGAACTCATTCTCCATGATCTTGATGATGATGATGATCAATCTGAGATATGGAA
ATCCCACTACTGCAATACTCATAGGATTCTACTGGTTGGTGAAGGCGACTTCTCTTTCTCTGCTTCTCTCGCCAGAGCTTTTGGATCTGCTTCCAACATT
ATCGCCTCTTCTCTTGATTCTCGGGAGTTCTTGAGGAAAAATTACAGGAGATCCATGGCCAACATCGAAGAGCTCATCTCCAGAGGGTGCCTAGTGTTGC
ATGGTGTGGACGCAACTAAAATGGCAAGGCTCAAATCTCTCAAGACCATGAAATTCGACCGTATCATTTTCAACTTCCCACATTCAGGCTTCTTCCCTGA
TGAATCCTCCCATTCACAGATTTGCCGACACCAAAAGCTGATACAAGAGTTCATGATGAATGCGAAGAAGCTGGTTTCAGAAGACGGGGAAATTCACATA
TCCCACAAGACAAATGGGTTCCATCTGAAGTGGGATTTGGAAGGTCTTGGTGAATATGTTGGGCTTCAGCTCATAGAAGAAGTTCCCTTTGATTTCGCTG
CTTTTACAGGATATCGAACCAAGTATGGTTTTGGAGGCAACAAGAACTTCAACTCAAAGCCCAGCAGCACTTACAAATTTGGACTTCCGTGA
AA sequence
>Lus10039534 pacid=23165253 polypeptide=Lus10039534 locus=Lus10039534.g ID=Lus10039534.BGIv1.0 annot-version=v1.0
MKKKKGSFGKKLISYLLGCICSNPEKEERYFHQEFQLASSASASSCDDSELILHDLDDDDDQSEIWKSHYCNTHRILLVGEGDFSFSASLARAFGSASNI
IASSLDSREFLRKNYRRSMANIEELISRGCLVLHGVDATKMARLKSLKTMKFDRIIFNFPHSGFFPDESSHSQICRHQKLIQEFMMNAKKLVSEDGEIHI
SHKTNGFHLKWDLEGLGEYVGLQLIEEVPFDFAAFTGYRTKYGFGGNKNFNSKPSSTYKFGLP
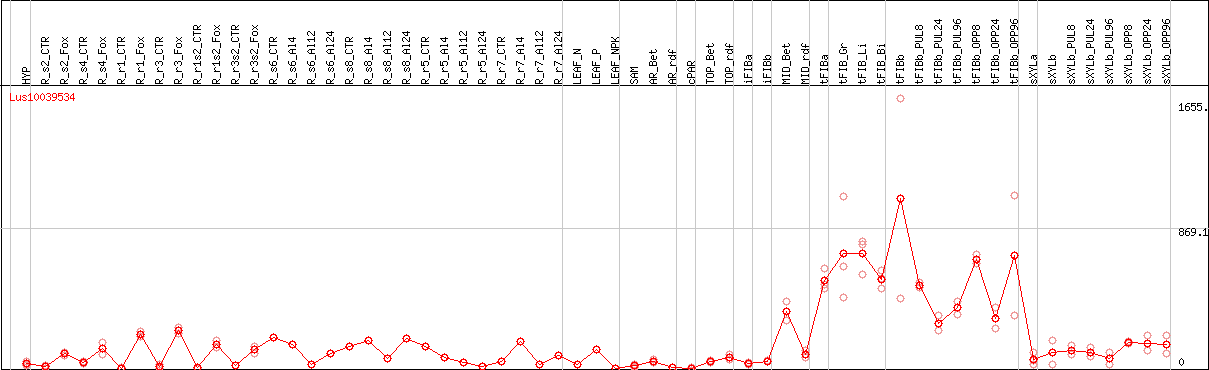
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039534 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.