External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G12410 87 / 8e-21
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT5G48350 78 / 6e-18
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT4G13870 79 / 2e-17
WRNEXO, ATWRNEXO, WEX, ATWEX
Werner syndrome-like exonuclease (.1.2)
AT2G36110 73 / 1e-15
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G12470 65 / 7e-13
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G56310 65 / 2e-12
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G12460 63 / 4e-12
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G12430 57 / 8e-10
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G12440 56 / 2e-09
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G62320 52 / 2e-08
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029478
338 / 4e-120
AT5G48350 82 / 2e-19
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10029479
188 / 6e-61
AT2G36110 87 / 4e-21
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10039164
141 / 4e-42
AT4G13870 105 / 2e-27
Werner syndrome-like exonuclease (.1.2)
Lus10013769
137 / 3e-40
AT4G13870 105 / 4e-27
Werner syndrome-like exonuclease (.1.2)
Lus10043049
79 / 7e-18
AT2G36110 121 / 4e-34
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10022556
62 / 1e-11
AT4G13870 246 / 1e-80
Werner syndrome-like exonuclease (.1.2)
Lus10010930
59 / 2e-10
AT1G56310 640 / 0.0
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10010357
55 / 4e-09
AT3G11770 66 / 7e-13
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10036491
54 / 9e-09
AT3G11770 64 / 4e-12
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0219
RNase_H
PF01612
DNA_pol_A_exo1
3'-5' exonuclease
Representative CDS sequence
>Lus10039591 pacid=23165119 polypeptide=Lus10039591 locus=Lus10039591.g ID=Lus10039591.BGIv1.0 annot-version=v1.0
ATGGAGGTTAACTACGTGAGATCGGTCTACTTCGGCGGCTACGAAATCGAAACCACGGTTACCGACCAGGCCTACGCTGTCGAAAACTGGGTTCGATCCT
TCCCGCCTGGTTGGAAGCGGATAGTGGGACTGGACTGCGAGTGGATGCCCTTCTTTTATAACAACCAGGACAACATAACGGCTGTGCTCCAACTCTGCGT
CGAAACCAAGTGCATCATCATACAGATGTCCTACCTCGAATACATCCCCCAGGCCCTGGCCGATTTCCTCGAGAATCCCAATAACTATTTCGCAGGGGTC
GGGGTGGGCGCTGACGTGAGTAGGTTAACAGCAGATTACGGGCTCAGCTGCTCCTCAACTGCTGACATTCAGACCATCGCTAGGGAAGAGTGGAACTGGA
GTAATCTTCCTGGGCTGAAGAATATTTTGCGCCATGTGGAAGGGCTTGTCATGGAGAAACCGCAGCATATTAGGCTGAGCAATTGGCAGAACAGAGATCT
AGACCCCGAGCAGGTTGAGTATGCTTGCATTGATGCTTTTGCTTGTTATAAGATCGGCTACAATCTGCGACGCCGCATCGATGCTTTTATTTACTAG
AA sequence
>Lus10039591 pacid=23165119 polypeptide=Lus10039591 locus=Lus10039591.g ID=Lus10039591.BGIv1.0 annot-version=v1.0
MEVNYVRSVYFGGYEIETTVTDQAYAVENWVRSFPPGWKRIVGLDCEWMPFFYNNQDNITAVLQLCVETKCIIIQMSYLEYIPQALADFLENPNNYFAGV
GVGADVSRLTADYGLSCSSTADIQTIAREEWNWSNLPGLKNILRHVEGLVMEKPQHIRLSNWQNRDLDPEQVEYACIDAFACYKIGYNLRRRIDAFIY
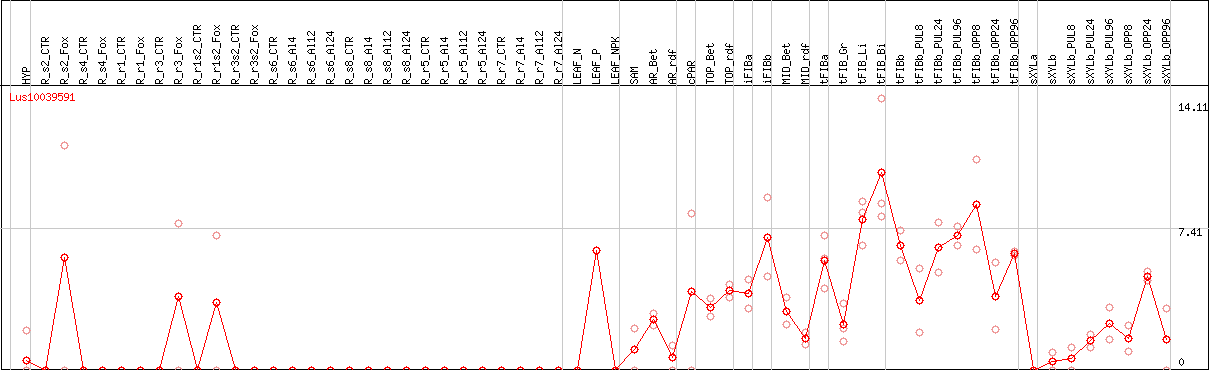
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039591 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.