External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G02120 65 / 5e-16
PDF2.1, LCR70
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 70, Scorpion toxin-like knottin superfamily protein (.1)
AT2G02100 65 / 5e-16
LCR69, PDF2.2
low-molecular-weight cysteine-rich 69 (.1)
AT2G02130 62 / 1e-14
PDF2.3, LCR68
low-molecular-weight cysteine-rich 68 (.1)
AT1G61070 59 / 9e-14
LCR66, PDF2.4
PLANT DEFENSIN 2.4, low-molecular-weight cysteine-rich 66 (.1)
AT5G63660 54 / 9e-12
LCR74, PDF2.5
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 74, Scorpion toxin-like knottin superfamily protein (.1)
AT2G31957 46 / 1e-08
LCR75, LCR79
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 79, low-molecular-weight cysteine-rich 75 (.1)
AT2G02140 46 / 1e-08
LCR72, PDF2.6
low-molecular-weight cysteine-rich 72 (.1)
AT2G31953 40 / 3e-06
LCR76
low-molecular-weight cysteine-rich 76 (.1)
AT2G02147 34 / 0.0006
LCR73
low-molecular-weight cysteine-rich 73 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029546
105 / 7e-32
AT2G02120 65 / 1e-15
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 70, Scorpion toxin-like knottin superfamily protein (.1)
Lus10000290
103 / 2e-31
AT2G02120 67 / 2e-16
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 70, Scorpion toxin-like knottin superfamily protein (.1)
Lus10019226
61 / 3e-14
AT2G02120 81 / 6e-22
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 70, Scorpion toxin-like knottin superfamily protein (.1)
Lus10004290
60 / 6e-14
AT2G02130 78 / 1e-20
low-molecular-weight cysteine-rich 68 (.1)
Lus10004289
59 / 1e-13
AT2G02130 78 / 1e-20
low-molecular-weight cysteine-rich 68 (.1)
Lus10029544
56 / 4e-12
AT2G02130 87 / 4e-24
low-molecular-weight cysteine-rich 68 (.1)
Lus10019227
47 / 9e-09
AT2G02130 66 / 1e-15
low-molecular-weight cysteine-rich 68 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.T011201
61 / 2e-14
AT2G02130 76 / 6e-20
low-molecular-weight cysteine-rich 68 (.1)
Potri.T011200
61 / 2e-14
AT2G02130 76 / 6e-20
low-molecular-weight cysteine-rich 68 (.1)
Potri.T011301
57 / 4e-13
AT5G63660 62 / 2e-14
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 74, Scorpion toxin-like knottin superfamily protein (.1)
Potri.T011300
57 / 4e-13
AT5G63660 62 / 2e-14
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 74, Scorpion toxin-like knottin superfamily protein (.1)
Potri.004G138100
52 / 9e-11
AT5G63660 71 / 4e-18
LOW-MOLECULAR-WEIGHT CYSTEINE-RICH 74, Scorpion toxin-like knottin superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0054
Knottin_1
PF00304
Gamma-thionin
Gamma-thionin family
Representative CDS sequence
>Lus10039622 pacid=23165031 polypeptide=Lus10039622 locus=Lus10039622.g ID=Lus10039622.BGIv1.0 annot-version=v1.0
ATGGGTGATGCTACCGGAGCAAGAAGATGTGAGTATGAAAGCAGGAGATTCAAAGGTACATGTGAGGTAGACAAAAATTGTGCCGATGTTTGCAAGCTCG
AAGGCTTCCCCGGAGGTGATTGCCAGGGCTTCCGCACCGGTTGCTACTGCACCAGGCCTTGCTAA
AA sequence
>Lus10039622 pacid=23165031 polypeptide=Lus10039622 locus=Lus10039622.g ID=Lus10039622.BGIv1.0 annot-version=v1.0
MGDATGARRCEYESRRFKGTCEVDKNCADVCKLEGFPGGDCQGFRTGCYCTRPC
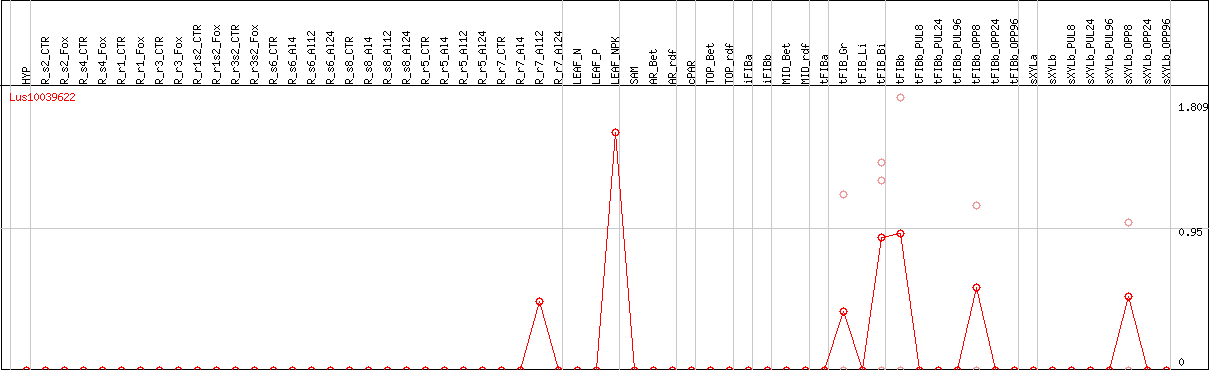
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039622 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.