External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G37890 246 / 1e-75
EDA40
embryo sac development arrest 40, Zinc finger (C3HC4-type RING finger) family protein (.1), Zinc finger (C3HC4-type RING finger) family protein (.2)
AT5G49665 226 / 5e-68
Zinc finger (C3HC4-type RING finger) family protein (.1)
AT2G22680 220 / 3e-66
WAVH1
WAV3 homolog 1, Zinc finger (C3HC4-type RING finger) family protein (.1)
AT5G65683 209 / 5e-62
WAVH2
WAV3 homolog 2, Zinc finger (C3HC4-type RING finger) family protein (.1)
AT2G38970 72 / 1e-13
Zinc finger (C3HC4-type RING finger) family protein (.1)
AT5G60710 68 / 2e-12
Zinc finger (C3HC4-type RING finger) family protein (.1)
AT1G08050 48 / 8e-06
Zinc finger (C3HC4-type RING finger) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003485
495 / 1e-171
AT2G22680 522 / 9e-177
WAV3 homolog 1, Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10028225
84 / 4e-20
AT5G49665 100 / 7e-26
Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10041321
60 / 1e-09
AT5G60710 730 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10028224
57 / 8e-09
AT5G49665 286 / 5e-91
Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10023527
56 / 3e-08
AT2G38970 887 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10016102
56 / 3e-08
AT5G60710 764 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10040413
52 / 3e-07
AT2G38970 846 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10021458
52 / 4e-07
AT5G60710 738 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10042930
50 / 0.0001
AT5G49665 397 / 7e-129
Zinc finger (C3HC4-type RING finger) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G107902
256 / 4e-79
AT5G49665 675 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Potri.008G038450
73 / 5e-14
AT2G38970 889 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Potri.010G223766
71 / 3e-13
AT2G38970 892 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Potri.003G148400
59 / 1e-09
AT2G38970 475 / 3e-159
Zinc finger (C3HC4-type RING finger) family protein (.1)
Potri.001G082150
58 / 5e-09
AT2G38970 466 / 4e-156
Zinc finger (C3HC4-type RING finger) family protein (.1)
Potri.009G010350
56 / 2e-08
AT5G60710 763 / 0.0
Zinc finger (C3HC4-type RING finger) family protein (.1)
Potri.006G000900
46 / 2e-05
AT3G54780 147 / 1e-37
Zinc finger (C3HC4-type RING finger) family protein (.1), Zinc finger (C3HC4-type RING finger) family protein (.2), Zinc finger (C3HC4-type RING finger) family protein (.3)
Potri.016G001300
46 / 3e-05
AT5G60710 156 / 1e-40
Zinc finger (C3HC4-type RING finger) family protein (.1)
PFAM info
Representative CDS sequence
>Lus10039635 pacid=23165218 polypeptide=Lus10039635 locus=Lus10039635.g ID=Lus10039635.BGIv1.0 annot-version=v1.0
ATGAGCGTCAACGACGCTCTGAAAAAGGCAGTGAAAGTGATCGAAGACCGACGTGACAAGAACCCGGTCGCCACCGTCTTAATCCTATCGGAAGATCACC
AGGATCGGTCCCGAAACACTTCGCTCTCTTGTTCGCCGCTGGCGTTGGTTTCCACCACGCGCTACTCGCAATTCGGGACTCCCGTTCATTCGATTAGCTT
CGGAGATTTGGGCGCGTACAAGCACGCTGCGGTAGAAGATGCCTTCTCCAAGTGTTTGGGTAGGTTGTTACGGGTCGTGGTTCAAGATCTGAAGATTCAA
CTCGGGTTCGTTACGGGTTCCGCCCCGGCAGAAATAGTAGCAGCATATCCGCTAACGGGTCCCCCGACTGTGTTCGTACCCGGGTCGATCGGACTAGGCG
ATCTCCACGTCGAGGAAGAGAGGGAGGTTCTCCTGGAAGTGAGAGTACCATCATCATCATCTTCGTCCGGTGGGTCCCGACGCCTGTTGTCCGTACGATC
GTCCTTTAAGGACACCTCATTGCAGGAGCAAATGAACGGTAGAGATCAGGTGATGATCGTTCCAAGGCCAGAAACCGTCCGATCCGGAAGGCTGAAGAAC
GTACACGTGTCGGTTCGAGCAGTAGCGGAATCTCGGAGATTAGTTGAGCACGGCGATTTATCAGGGGCGTATCACTTGTTGTCATCGGCTCGAGCTTTGT
TGATGCAGAGGAGCGACAGCTCGGCTGGGGAGTGTCTACAGTGCGTAGAGGCAGAGCTAGCAGAGCTGCAGAAGAAGAAGAAGCCGCTGATGACGACGGG
GAGGCAGAGCCAGCAGCAACAGAGGAGCGGTAACCGGCAGTCTGAAGCGCAAGTGGAGGAAAAGACGGAGGCGTTGACGCCGACGTCGGCTTGGCGAGCC
GCGGAGAAGTTGGCTAAGTTGGCGATCATGAGGAAGCACATGAACAGAGTCAGCGATTTGCACGGCTTTGAGAACGCAAGATTTTGA
AA sequence
>Lus10039635 pacid=23165218 polypeptide=Lus10039635 locus=Lus10039635.g ID=Lus10039635.BGIv1.0 annot-version=v1.0
MSVNDALKKAVKVIEDRRDKNPVATVLILSEDHQDRSRNTSLSCSPLALVSTTRYSQFGTPVHSISFGDLGAYKHAAVEDAFSKCLGRLLRVVVQDLKIQ
LGFVTGSAPAEIVAAYPLTGPPTVFVPGSIGLGDLHVEEEREVLLEVRVPSSSSSSGGSRRLLSVRSSFKDTSLQEQMNGRDQVMIVPRPETVRSGRLKN
VHVSVRAVAESRRLVEHGDLSGAYHLLSSARALLMQRSDSSAGECLQCVEAELAELQKKKKPLMTTGRQSQQQQRSGNRQSEAQVEEKTEALTPTSAWRA
AEKLAKLAIMRKHMNRVSDLHGFENARF
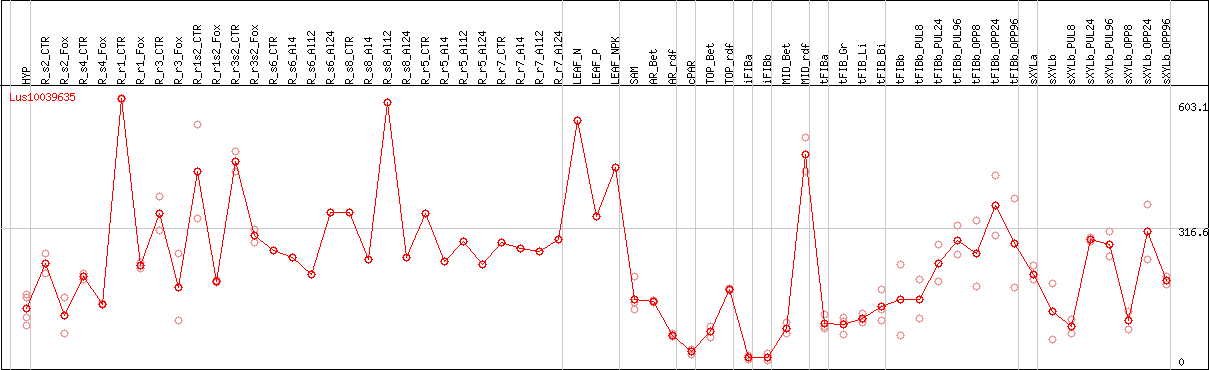
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039635 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.