External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G22870 414 / 1e-146
EMB2001
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G11480 268 / 4e-89
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G58370 105 / 3e-25
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
AT3G12080 47 / 1e-05
EMB2738
embryo defective 2738, GTP-binding family protein (.1.2)
AT1G08410 44 / 9e-05
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT2G27200 44 / 0.0001
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G52980 41 / 0.0007
AtNug2
nuclear/nucleolar GTPase 2, GTP-binding family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011703
531 / 0
AT2G22870 427 / 5e-152
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10022126
278 / 5e-93
AT5G11480 418 / 8e-148
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10011673
277 / 1e-92
AT5G11480 418 / 1e-147
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10009919
102 / 3e-24
AT5G58370 473 / 9e-165
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
Lus10018881
44 / 9e-05
AT3G07050 706 / 0.0
nucleostemin-like 1, GTP-binding family protein (.1)
Lus10028574
44 / 0.0001
AT3G07050 743 / 0.0
nucleostemin-like 1, GTP-binding family protein (.1)
Lus10004103
44 / 0.0002
AT1G08410 756 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10023835
43 / 0.0002
AT1G52980 708 / 0.0
nuclear/nucleolar GTPase 2, GTP-binding family protein (.1)
Lus10013359
43 / 0.0002
AT1G08410 745 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G006300
425 / 6e-151
AT2G22870 434 / 2e-154
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.006G243200
294 / 2e-99
AT5G11480 422 / 2e-149
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.018G037000
283 / 5e-95
AT5G11480 421 / 7e-149
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.014G006501
193 / 2e-62
AT2G22870 199 / 2e-65
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.014G007450
135 / 3e-40
AT2G22870 131 / 3e-39
embryo defective 2001, P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.019G128000
92 / 8e-21
AT5G58370 461 / 1e-158
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
Potri.007G021900
43 / 0.0001
AT5G66470 535 / 0.0
RNA binding;GTP binding (.1)
Potri.009G017000
42 / 0.0003
AT1G08410 755 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Potri.001G402500
41 / 0.0008
AT1G52980 811 / 0.0
nuclear/nucleolar GTPase 2, GTP-binding family protein (.1)
Potri.002G241100
41 / 0.001
AT3G07050 667 / 0.0
nucleostemin-like 1, GTP-binding family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF01926
MMR_HSR1
50S ribosome-binding GTPase
Representative CDS sequence
>Lus10039653 pacid=23165138 polypeptide=Lus10039653 locus=Lus10039653.g ID=Lus10039653.BGIv1.0 annot-version=v1.0
ATGCTTCTCCGGTACTCCCGTCTGGCAACTTTCCCCCTCATTACACCTCCAGCGACTCGGAAGTTTCTCCCTCTGAGCACCCATTTCAAATTCTCGATTG
AAAGATTTGCTTCCTCTATCTCACCGAGTCATGTCTCCGCTTCAAAGGCGGCAAAGCCATCGGCGTCGGACCCAGCGAGGTCTGTTCGGACGGTTCTCTT
CGTTCCCCCGGGAGTTGAGCTGGAGGAGGTGAGGGAGGACATGATTTTATCCGGTTCGAACATCGTAGTCGGACCATATGCGGGTCATTCGCAGATAAAG
GAAGTGGAGTTCGTGAAGAGCAGTGCGCGGGCAAAGGATTGCCCCAAAGATGATAAACCCGAGTTCGCCATTCTGGGGAGGTCCAATGTTGGAAAGTCAT
CGCTAATCAATTCTTTGGTTCGGAAGAAAGAAATTGCTCTTACATCTAAAAAGCCAGGGAAGACTCAGTTGATCAATCATTTCTTGGTTAACAAAAGTTG
GTACCTTGTTGATTTGCCTGGTTACGGGTTCGCCAAAGCATCAGATGCAGCTAGAATGGATTGGTCTGGATTCACAAAAGGATACTTTCTGAACAGAGAG
AGTCTGGTTGGTGTGCTACTCCTAATTGATGCAAGCGTTCCTCCTCAAAAGATAGACCTGGACTGCGCAAATTGGCTTGGACGAAACAAAATACCATTAA
CTTTTGTTTTAACCAAGTGTGATAAGGTGAAAGGCGGGAAAGGAACGAGGCCAGAAGAGAACATCAGTAATTTCCAGGAGTGGATAGATGAGAGTTACAG
ACAGCAACAACTACCTTGGATCATGACTAGTAGTGTCACCGGGTTAGGAAGGGATGAGCTTCTCTTGCATATGTCACAACTCAGAAACTACTGGGATCAG
TAA
AA sequence
>Lus10039653 pacid=23165138 polypeptide=Lus10039653 locus=Lus10039653.g ID=Lus10039653.BGIv1.0 annot-version=v1.0
MLLRYSRLATFPLITPPATRKFLPLSTHFKFSIERFASSISPSHVSASKAAKPSASDPARSVRTVLFVPPGVELEEVREDMILSGSNIVVGPYAGHSQIK
EVEFVKSSARAKDCPKDDKPEFAILGRSNVGKSSLINSLVRKKEIALTSKKPGKTQLINHFLVNKSWYLVDLPGYGFAKASDAARMDWSGFTKGYFLNRE
SLVGVLLLIDASVPPQKIDLDCANWLGRNKIPLTFVLTKCDKVKGGKGTRPEENISNFQEWIDESYRQQQLPWIMTSSVTGLGRDELLLHMSQLRNYWDQ
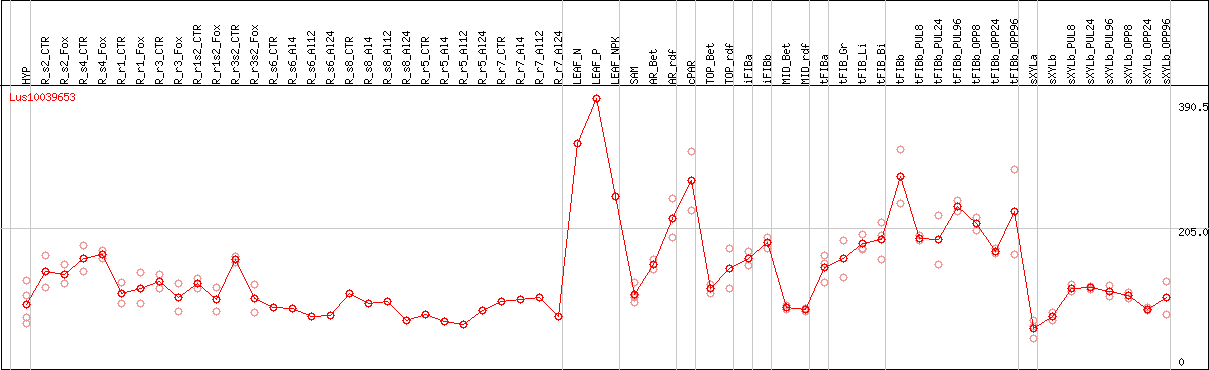
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039653 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.