Lus10039684 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10039684 pacid=23165109 polypeptide=Lus10039684 locus=Lus10039684.g ID=Lus10039684.BGIv1.0 annot-version=v1.0
ATGGAGAAATCGTGTTCTCTACTGGTTCACTTCGACAAGGGATCGCCAGCGGTTGCGAATGAGATTAAGGAAGCTCTGGAAGGGAATGATGTTTCGGCCA
AGATTGATGGCATGAAGAAGGCAATTAGCCTTCTGTTGAATGGTGAAAACTTGCCTCAGCTTTTCATCACTATTCTTAGATATGTACTGCCCTCTGAGGA
TCACACCATACAGAAGCTGTTGCTGTTCTACTTGGAGATCATTGATAAGAAAGATGCTTCGGGGAAGCTTCTTCCGGAGATGATTTTGATATGCCAGAAT
CTGATGAACAATCTTCAACACCCTAACGAGTACATCCGTGGGGTGACTTTGCGGTTTCTTTGCCGGATTCACGAGACAGAGATTATAGAGCCGTTAATCC
CTTCGGTGCTGCAGAATCTGGAGCATAGGCACCCCTTTATTAGGAGGAATGCCATCTTTGCAGTGATGTCCATCTATAAGCTTCCCCAGGGCGACCAGCT
GCTCATCGATGCTCCGGAAATGATCGAGAAGGCTCTTTCCTCAGAGCACGATCCTTCAGCCAAGCGCAATGCGTTCCTGATGCTTTTCACTTGTGCACAA
GATCGAGCTATTAATTATCTTTTCAGCCACGTTGATAAGATCCCTGAATGGGATGAGAAGCTTCAGATGGTTGTTTTGGAGTTGATTCGTAAAGTTTGTA
AAACAAGTAGGGGAGAGAAGGGAAGATACATCAAAATCATCATATCATTGTTGACTGCACCTTCGAGTGCTGTGGTTTACGAGTGTGCTGGAACTCTTGT
TTCTCTTTCATCTGCTCCTACTGCCATTAAAGCTGCTGCTAACACATACGGGCAACTCCTTTTGTCTCAGAGTGATAACAATGTCAAGCTTATTGTGCTT
GATCGCTTGCAAGAGCTAAAGTCTTCTCACAGGGAAATCATGGTCGAGATGGTTATGGACGTGCTTAGGGCACTCTCTAGCCCCAATCTTGATATACGCC
GCAAGACCCTTGATATTGTTCTTGACTTGATCACTGCTAGGAATGTGAATGAAATTGTTCAGATGCTCAAGAAGGAAGTTGTTAAGACCCAGAGTGGAGA
GCTTGAAAAGAATGGCGAGTACAGGCAGATGCTTATACAAGCCATCCATTTTTGTGCTATTAAGTTCCCGGAGGTGGCAAGCATAGTGGTTCACTTGTTG
ATGGATTTCCTTGGAGATAGCAACATTGCTTCAGCTATTGATGTGGTTGTTTTTGTCCGGGAAATAATCGAAACCAACCCGAAACTCAGAGTTTCGATAG
TAACAAGGCTGTTGGATACTTTCTATCAGATCCGAGCAGCCAGGGTATGCTCTGGTGCCCTTTGGATCATTGGGGAGTACTGCGTTTCCCTTTCTGAAGT
AGAGAGTGGCATAGCAACAATCAAGCAGTGTCTTGGAGAATTGCCGTTTTTCTCTATTTCAGAAGAAGGTGAAGTTACTGATTCTTCCAAAAAGCCACCA
CAACCGAGTTCCATCACAGTCTCTTCAAGAAAACCCGCTATCCTTTCTGATGGGACATATGCAACACAAAACGCTGCCTCTGAAACAGCATTTTCTCCGC
CTACTGTAGTTCAAGGATCACTGTCCACTGGAAATTTGAGATCCCTTCTTCTCACTGGCGATTTTTTCCTTGGAGCAGTTGTTGCATGCACTCTAACAAA
ACTGGTTCTGAGACTCGAAGAGGTGCAATCTTCTAAACTGGAGGTCAACAAGGCAGCCACACAGACTTTGTTGATAATGGTCTCCATGCTTCAATTGGGA
CAGTCTTCTGTTCTGCCTCATCCAATTGACAGCGATTCATATGATAGGATCCTTCTGTGCATTAGGTTGCTATGTAATACTGGTGATGACATCCGGAAGA
TATGGTTGCAATCATGCCGTCAAAGCTTTGTGAAAATGCTCTCAGAAAAGCAATTGAGGGAAACTGAGGAATTGAAGGCAAAGGCTCAGGTTTCTCATGC
ACAACCAGATGATCTTATCGACTTCTATCATTTAAAGAGCAGGAAGGGTATGAGTCAGCTGGAGTTGGAAGATGAAGTACAAGATGATTTGAAGCGTGCA
ACGGGAGAGTTCATCAACGATGGCAATGACGCTAATAAGCTCAACCGAATCCTTCAGCTCACTGGATTTAGTGACCCTGTGTATGCCGAAGCACATGTCA
AGGTCAACCACTACGACATCGTACTTGATGTCACTGTCATCAACCGGACAAAGGAAACTCTCCAAAACTTGTGCCTGGAGTTGGCTACAATGGGCGATCT
CAAGCTAGTTGAGCGTCCTCAGAATTATGCCTTGGCACCAGAATCAAGCAAACAGATAAAGGCCAACATCAAGGTCTCCTCAACTGAGACTGGCGTCATT
TTCGGTAACATTGTTTTCGAGACATCCAATGTGATGGAACGTTCAGTTGTCGTCCTCAACGATATCCACATCGATATCATGGATTACATCTCTCCTGCTT
CTTGCTCGGATGCTGCGTTCCGGACTATGTGGGCAGAATTTGAATGGGAAAATAAGGTTGCTGTTAACACAACAATTCAGAACGAGAAGGAATTCCTGGA
CCATATCATCAAGTCTACAAATATGAAGTGTCTGACGGCACCGTCTGCACTTGATGGAGAGTGTGGATTCCTGGCTGCGAACTTGTACGCCAAGAGTGTG
TTCGGCGAGGACGCATTGGTGAACATAAGCCTGGAGAAGGAAGGAGATGGAAAGCTCAGTGGATACATCAGGATTCGGAGCAAGACGCAAGGAATCGCGC
TCAGCCTTGGTGATAAAATCACCTTGAAGCAGAAAGGTGGCAGTAGTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10039684 pacid=23165109 polypeptide=Lus10039684 locus=Lus10039684.g ID=Lus10039684.BGIv1.0 annot-version=v1.0
MEKSCSLLVHFDKGSPAVANEIKEALEGNDVSAKIDGMKKAISLLLNGENLPQLFITILRYVLPSEDHTIQKLLLFYLEIIDKKDASGKLLPEMILICQN
LMNNLQHPNEYIRGVTLRFLCRIHETEIIEPLIPSVLQNLEHRHPFIRRNAIFAVMSIYKLPQGDQLLIDAPEMIEKALSSEHDPSAKRNAFLMLFTCAQ
DRAINYLFSHVDKIPEWDEKLQMVVLELIRKVCKTSRGEKGRYIKIIISLLTAPSSAVVYECAGTLVSLSSAPTAIKAAANTYGQLLLSQSDNNVKLIVL
DRLQELKSSHREIMVEMVMDVLRALSSPNLDIRRKTLDIVLDLITARNVNEIVQMLKKEVVKTQSGELEKNGEYRQMLIQAIHFCAIKFPEVASIVVHLL
MDFLGDSNIASAIDVVVFVREIIETNPKLRVSIVTRLLDTFYQIRAARVCSGALWIIGEYCVSLSEVESGIATIKQCLGELPFFSISEEGEVTDSSKKPP
QPSSITVSSRKPAILSDGTYATQNAASETAFSPPTVVQGSLSTGNLRSLLLTGDFFLGAVVACTLTKLVLRLEEVQSSKLEVNKAATQTLLIMVSMLQLG
QSSVLPHPIDSDSYDRILLCIRLLCNTGDDIRKIWLQSCRQSFVKMLSEKQLRETEELKAKAQVSHAQPDDLIDFYHLKSRKGMSQLELEDEVQDDLKRA
TGEFINDGNDANKLNRILQLTGFSDPVYAEAHVKVNHYDIVLDVTVINRTKETLQNLCLELATMGDLKLVERPQNYALAPESSKQIKANIKVSSTETGVI
FGNIVFETSNVMERSVVVLNDIHIDIMDYISPASCSDAAFRTMWAEFEWENKVAVNTTIQNEKEFLDHIIKSTNMKCLTAPSALDGECGFLAANLYAKSV
FGEDALVNISLEKEGDGKLSGYIRIRSKTQGIALSLGDKITLKQKGGSS
|
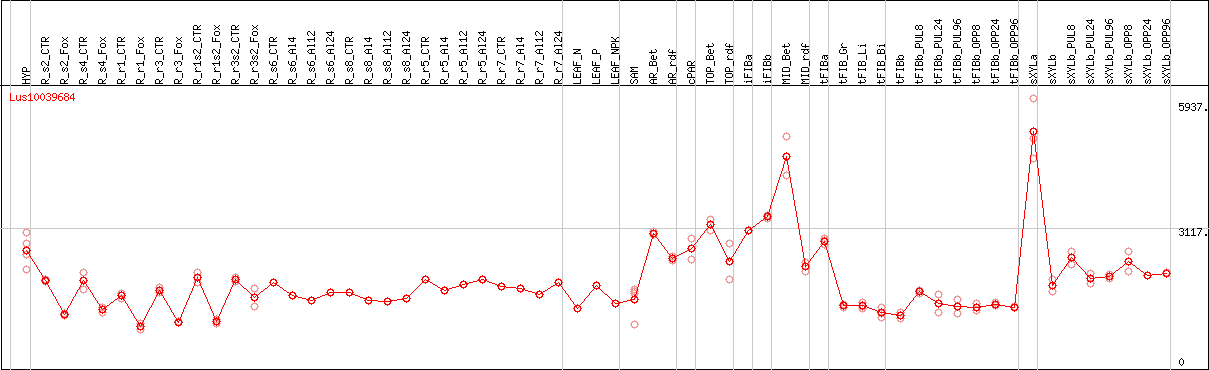
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10039684 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
