External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13840 100 / 2e-26
FZR3
FIZZY-related 3 (.1.2)
AT4G11920 92 / 7e-24
FZR1, CCS52A2
FIZZY-RELATED 1, cell cycle switch protein 52 A2 (.1)
AT4G22910 91 / 1e-23
CCS52A1, FZR2
cell cycle switch protein 52 A1, FIZZY-related 2 (.1)
AT4G33270 70 / 6e-16
AtCDC20.1, CDC20.1
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
AT4G33260 69 / 1e-15
AtCDC20.2, CDC20.2
cell division cycle 20.2, Transducin family protein / WD-40 repeat family protein (.1.2)
AT5G27570 65 / 4e-14
AtCDC20.5
cell division cycle 20.5, Transducin/WD40 repeat-like superfamily protein (.1)
AT5G27945 64 / 4e-14
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G27080 64 / 8e-14
AtCDC20.3
cell division cycle 20.3, Transducin family protein / WD-40 repeat family protein (.1)
AT5G26900 63 / 1e-13
AtCDC20.4
cell division cycle 20.4, Transducin family protein / WD-40 repeat family protein (.1)
AT5G52820 41 / 1e-05
WD-40 repeat family protein / notchless protein, putative (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023401
110 / 1e-30
AT5G13840 546 / 0.0
FIZZY-related 3 (.1.2)
Lus10040282
111 / 2e-30
AT5G13840 685 / 0.0
FIZZY-related 3 (.1.2)
Lus10001192
91 / 3e-23
AT4G22910 734 / 0.0
cell cycle switch protein 52 A1, FIZZY-related 2 (.1)
Lus10024482
91 / 3e-23
AT4G22910 733 / 0.0
cell cycle switch protein 52 A1, FIZZY-related 2 (.1)
Lus10037595
69 / 3e-16
AT4G33270 402 / 2e-141
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10037593
69 / 1e-15
AT4G33270 728 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10006853
67 / 7e-15
AT4G33270 729 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10006851
67 / 9e-15
AT4G33270 728 / 0.0
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
Lus10038263
62 / 2e-14
AT4G33270 108 / 1e-29
cell division cycle 20.1, Transducin family protein / WD-40 repeat family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Lus10040004 pacid=23173562 polypeptide=Lus10040004 locus=Lus10040004.g ID=Lus10040004.BGIv1.0 annot-version=v1.0
ATGGTGTGGAAGTATCCGTCTTTATCAAAGGTTGCAACTCTAACTGGACACAGTTTGCTGGTTCTCTCCCTTGCCATGTTGCCTGACGGCCAGACCATAG
TGACTGGAGCGGGAGATGAAACTCTGCGGTTCTGGAATGTCTTCCCATCCATGAAAACACCTGCGCCGGTTCAAGGGCACTTTTTGTGA
AA sequence
>Lus10040004 pacid=23173562 polypeptide=Lus10040004 locus=Lus10040004.g ID=Lus10040004.BGIv1.0 annot-version=v1.0
MVWKYPSLSKVATLTGHSLLVLSLAMLPDGQTIVTGAGDETLRFWNVFPSMKTPAPVQGHFL
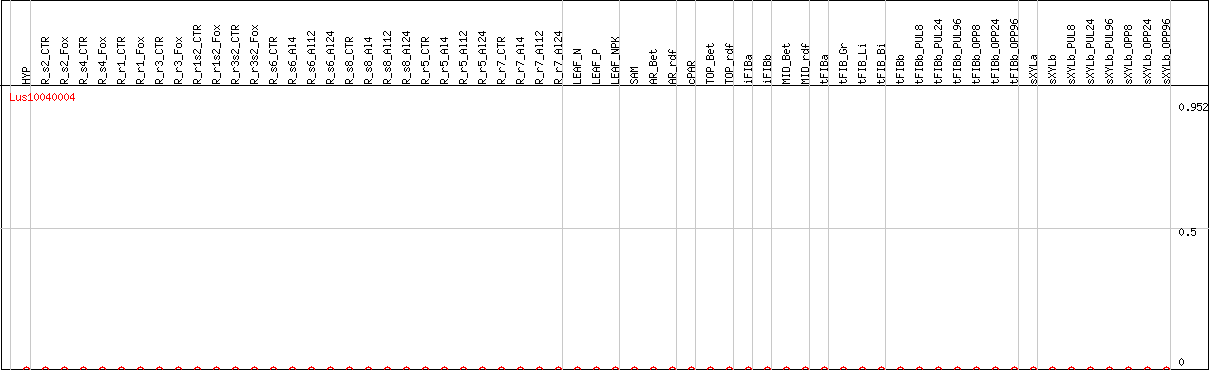
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040004 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.