Lus10040006 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10040006 pacid=23173514 polypeptide=Lus10040006 locus=Lus10040006.g ID=Lus10040006.BGIv1.0 annot-version=v1.0
ATGAGCCCTACCGATTTTTACTCTGCCCCTGATGGGACTATTCCGGTTTTCTACCCGGATACCGGCGGCGGATTTGTTGGTGTAGGGTATAGGAATGTGG
CGCCGGCTGCGATTCCCCAATGGGTGCCGGTGGCACCTGGTAATGTGAATATTGGCGTGGCAGCTCCTCAAGCTGTTGGTGGGTTTGGTTATAATCCCAA
TTTGGGGAATAGGGTAGTTGGTAATGCTGTTGATCAATCTAGTAGTGAAATGTTGGTTGGACTTAGTTCCAATCCGCCGCTTAATGGCAATGGGCACACT
GAATTTTTGAATGCAGGGTATATTTATAATCCCAGTTTAGGTAATAGGGGGAGTGGTAGTGGTGGTTGTGGTGGTGGTAGGACTGATTACGCTAGCGAAG
ATGGCGGGGATGATTCGGTTTCTGGGAAGAAAGTTAAGCTTTTGTGTAGTTTTGGAGGGAAGATTCTTCCTAGACCTATTGATGGAGCATTGAGATACGT
CGGAGGTCAGACAAGGATCATTAGCTTGAGAAGGGAAGTGAATTTCAGTGAGCTAATGCAGAAGATGGTCGATACTTATGGACAACCTGTTGTTTTGAAG
TACCAGCTTCCCGAGGAGGATTTGGATGCATTGGTGTCAGTTTCATGTCCCGAAGATGTCGATAACATGTTGGATGAGTATGACAAGTTAGTGGAGAGAT
CTTCTGACGGTTCAGCTAAGTTAAGGGTCTTTTTGTTTTCAGCCACAGAGCTTGATGCTTCTGGTATGGTACATTTAGGAGATATGCAGGATAATGAACA
GCGATATGTCGACGCGGTTAATGGGATTGGTGGGATCTCGAGGAAGGAAAGTATGGCGAGTGCAACTTCCACACAGAATTCTGATATTAGTGGTGCTGAA
GCTGCTGACAACATCTCAGGCAGCGGACAAGGGGAAATGAGCAGACCTCCATCAGGCGGCGGGCTTACCCCAAGATTGGTGCAAGTGGATCCTAACTCCC
CACCACTTTATGCAGCAGATACATCATCATCTGTTTCATTAGGTATTCCACTTATCAACTCCGCTTCTCCTACAGCTTTAATCTCCCCACATGAAGTCGA
CTTTGAAAGGTCTGCACCTGTTAATGTCTCACAGCAGCACTTCGGTTACGAGTTACCACAAACCGGAATGCCCATTCAACCAACTGCACCTCATTTCCAG
GCATATCGGAATCCTCGTCAGGATTATTTACATCCCCCTCCTCAAATAGGATATCCAAACCCTCAAATGGCGGGGACTGCTGGTGGGTCAGTATTTGCTC
AGCAGCCAATGCATGATGTTAATGCTTCTATGGCTCCTCATCAGTATATTCCTACAGAGCACATGACTATGGCTCCTGTGAGACCTTTAATTGTGCAGCC
ATTGATGCAAGCTCAGCAAACTCGGGTGGACCATTATCCAGATGACAGGACATATGGAACACGTCTTCGTTCAGAAGATAGTGCAAAGCTGCAACCTGTG
AACCGAATAATGGTGACTGGAGCAGCAGGAGAAAGCACCCTGGTTCAGGGTGATGGGGCTCGACATATAGTCACGGGTAATCTGGAACAGACAATTGGGG
CAGCTCAATCAGAAGCAATCGATTTGCCCCGGAGTGCTGAAGCAATTCACGAGTTGGAGAGTATTTCTCTCAGGAAAAAGGAAGCTCCTGAGCAACCTAA
GATGCCAGTTCCTGTTGATATGACAAGTCTACCATGTGATGCACAGACTAATTATGGACAGTTGATGGGTAAAACACCTCAGTCTCAGGTAGGAGATTCT
ATTCAGCAGCAGCAGGCAGCGACAACTATGTATCCAATTAACCAAAATGCACTATTAAAAATGCATGTTCGTGACGTGACTCATGTTGCTGGTGGTCCTA
TTCAAACTTCTGAAGGCATGATCTATGAGTCTCCAAAGGCACCTGCAGGTAACCTTCCTCTTGTTGTTCCAAAAGAAAATACGTTCAGCTCGTTTTTCTC
CCGCGACCCGCTGAAACCAATTGAAGGCATGAAAGGTGCTTCTGAAATTAATTTATGTAATGAGCTCAACAAGCTGCCCAGTGATCCGAAAACCCAGCTA
GTAGCTGTGAGGGAGCTATTTGATGATGCACTTAACAAACCTCAAGATTCGAATCTCATCAAATCTGAAGTTCAGCCTACTGCTGAATTCTCATATCTAC
ACAGCTCTCAGCCCATGGAACCATATAATGTTGCTCAACCATCATATCCGGGTGATCCAGGAGCACATCTGCATTCCATGGTTGGTACCCTCTTGGATTC
TGGTGATGTCAGTTGTGTCAATCAAAAGCATGTCGATGAGCCAGTATATCAAGTTGATAGACTCCCACCTGTTAGTGAGAGAAATGATGACATCTCTAAG
TTATTACCGGATACGCTTTCTGAGGATGTGGAGTCTTTGATACTGAGTGAGAATGCACCATCTTCAATATCGCCATCTAGTAGGGTTGAGGGTATTCAGG
AATCGTCAGATTCTGTTTTCAGCAACCAGGACCCTTGGAACTTGAGGCACGGCGATACACATTATCCTCCTAAACCTCTCAAGATTGGGACGAGAAAAGA
TACTGTTGGTGCTGGTGATCCTTTCAGGGAAAATCATCCGGGTAATTCTGGAGAATCAATTCCAGACGTGCTGTTGGACGGTGGAACTTCCCAAACCCTA
TGCAGTTCGGAGGATCCTCTGTCATCTAAAGGGTCCGAGGATAAGCTTATTAAACAAGAACTTCAGGCTGTTGCTGAGGGAGTAGCTGCATCTGTACTTA
AATCGAGTATACCCGATGCTGACTTAACAGGTCATGAAAGAAATGAATCTCTTTCTAAAGTAATGGATTCCGAGGAAGTCCCAAATAAAGATGTGGAGAC
ACAGTATAAAGTAAAAGCTGAGGATCTGAAGAACAAGCAGCCAGAGAGAGTGAAGTTGGGATTTCCAATGTCAGAAGGTGCAGGACGGCTACAGATCATT
AAGAGCAGTGACCTTGAAGAGTTGCAAGAATTGGGATCTGGTACCTTTGGCACTGTGTATCATGGAAAGTGGAGGGGTACGGATGTTGCTATTAAGCGAA
TCAACGACAGGTGTTTTGTGGGGAAGCCTTCTGAGCAAGAACGCATGATAGATGATTTCTGGAATGAAGCTATCAAGCTAGCTGACTTGCACCATCCAAA
TGTGGTTGCTTTTTATGGTGTCGTTCTTGATGGCCCAGGAGGTTCTGTTGCAACAGTGACAGAGTATATGGTAAATGGCTCTTTGCGGACTGCTTTGCTT
AAGAATGAGAGGAGCCTTGACAAGCGGAAGCGCTTGTTGATTGCAATGGACGTGGCTTTTGGGATGGAATATTTACATGCTAAAAACATTGTGCATTTCG
ACCTCAAGAGTGACAATCTGCTTGTCAATCTTAGAGATCCACACCGTCCGATATGCAAGGTTGGCGATCTGGGGCTGTCCAAGGTAAAATGTCAAACGCT
CATCTCAGGTGGTGTGAGAGGAACGCTTCCATGGATGGCACCGGAGCTTTTGAATGGCAGCAGCAGCCTCGTCTCTGAAAAGGTTGATGTTTTCTCATTT
GGCATCGTGTTGTGGGAGCTCCTGACTGGGGAAGAGCCATATGCGGACTTGCACTACGGCGCGATTATAGGTGGTATTGTGAGCAATACATTAAGGCCAG
CAGTGCCAGAGCATTGTGATCCGGTGTGGAAATCATTGATGGAAAGATGCTGGGCTTCTGAACCACCCCAGAGACCAAGCTTCACTGCGGTTGCAAACGA
GCTGCGAGCTATGGCTGCCAAGATACAAACGCAGGCTCAAAAGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10040006 pacid=23173514 polypeptide=Lus10040006 locus=Lus10040006.g ID=Lus10040006.BGIv1.0 annot-version=v1.0
MSPTDFYSAPDGTIPVFYPDTGGGFVGVGYRNVAPAAIPQWVPVAPGNVNIGVAAPQAVGGFGYNPNLGNRVVGNAVDQSSSEMLVGLSSNPPLNGNGHT
EFLNAGYIYNPSLGNRGSGSGGCGGGRTDYASEDGGDDSVSGKKVKLLCSFGGKILPRPIDGALRYVGGQTRIISLRREVNFSELMQKMVDTYGQPVVLK
YQLPEEDLDALVSVSCPEDVDNMLDEYDKLVERSSDGSAKLRVFLFSATELDASGMVHLGDMQDNEQRYVDAVNGIGGISRKESMASATSTQNSDISGAE
AADNISGSGQGEMSRPPSGGGLTPRLVQVDPNSPPLYAADTSSSVSLGIPLINSASPTALISPHEVDFERSAPVNVSQQHFGYELPQTGMPIQPTAPHFQ
AYRNPRQDYLHPPPQIGYPNPQMAGTAGGSVFAQQPMHDVNASMAPHQYIPTEHMTMAPVRPLIVQPLMQAQQTRVDHYPDDRTYGTRLRSEDSAKLQPV
NRIMVTGAAGESTLVQGDGARHIVTGNLEQTIGAAQSEAIDLPRSAEAIHELESISLRKKEAPEQPKMPVPVDMTSLPCDAQTNYGQLMGKTPQSQVGDS
IQQQQAATTMYPINQNALLKMHVRDVTHVAGGPIQTSEGMIYESPKAPAGNLPLVVPKENTFSSFFSRDPLKPIEGMKGASEINLCNELNKLPSDPKTQL
VAVRELFDDALNKPQDSNLIKSEVQPTAEFSYLHSSQPMEPYNVAQPSYPGDPGAHLHSMVGTLLDSGDVSCVNQKHVDEPVYQVDRLPPVSERNDDISK
LLPDTLSEDVESLILSENAPSSISPSSRVEGIQESSDSVFSNQDPWNLRHGDTHYPPKPLKIGTRKDTVGAGDPFRENHPGNSGESIPDVLLDGGTSQTL
CSSEDPLSSKGSEDKLIKQELQAVAEGVAASVLKSSIPDADLTGHERNESLSKVMDSEEVPNKDVETQYKVKAEDLKNKQPERVKLGFPMSEGAGRLQII
KSSDLEELQELGSGTFGTVYHGKWRGTDVAIKRINDRCFVGKPSEQERMIDDFWNEAIKLADLHHPNVVAFYGVVLDGPGGSVATVTEYMVNGSLRTALL
KNERSLDKRKRLLIAMDVAFGMEYLHAKNIVHFDLKSDNLLVNLRDPHRPICKVGDLGLSKVKCQTLISGGVRGTLPWMAPELLNGSSSLVSEKVDVFSF
GIVLWELLTGEEPYADLHYGAIIGGIVSNTLRPAVPEHCDPVWKSLMERCWASEPPQRPSFTAVANELRAMAAKIQTQAQK
|
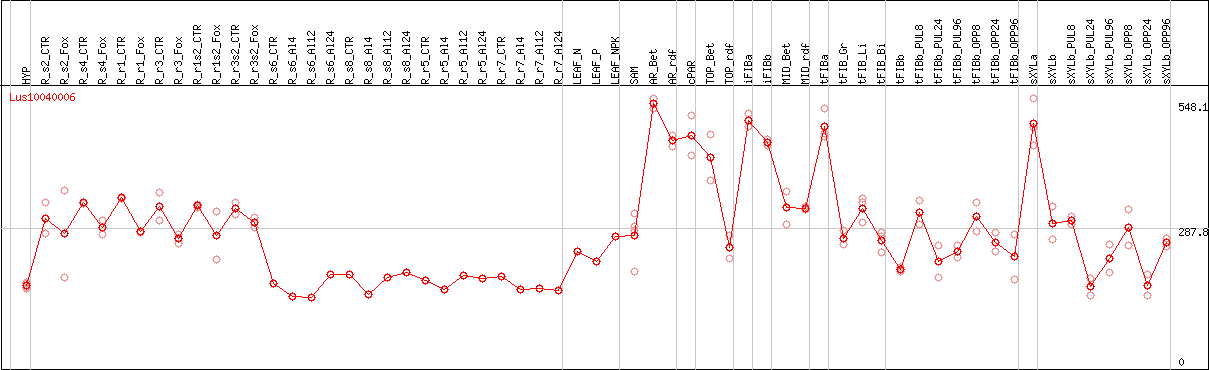
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040006 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
