External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G23180 245 / 5e-77
RLK4, CRK10
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
AT4G23290 242 / 2e-76
CRK21
cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.2)
AT4G23150 243 / 3e-76
CRK7
cysteine-rich RLK (RECEPTOR-like protein kinase) 7 (.1)
AT4G23240 234 / 7e-76
CRK16
cysteine-rich RLK (RECEPTOR-like protein kinase) 16 (.1)
AT4G23230 234 / 3e-74
CRK15
cysteine-rich RLK (RECEPTOR-like protein kinase) 15 (.1)
AT4G11530 238 / 4e-74
CRK34
cysteine-rich RLK (RECEPTOR-like protein kinase) 34 (.1)
AT4G23140 237 / 6e-74
RLK5, CRK6
cysteine-rich RLK (RECEPTOR-like protein kinase) 6 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 6 (.2)
AT4G23270 235 / 2e-73
CRK19
cysteine-rich RLK (RECEPTOR-like protein kinase) 19 (.1)
AT4G21380 238 / 4e-73
ARK3
receptor kinase 3 (.1)
AT4G23220 236 / 6e-73
CRK14
cysteine-rich RLK (RECEPTOR-like protein kinase) 14 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019599
389 / 2e-138
AT4G23290 180 / 6e-53
cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.2)
Lus10019061
403 / 5e-137
AT4G21380 338 / 5e-104
receptor kinase 3 (.1)
Lus10036314
268 / 3e-89
AT4G23180 315 / 6e-102
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Lus10024081
260 / 1e-81
AT1G61500 333 / 5e-102
S-locus lectin protein kinase family protein (.1)
Lus10041639
259 / 3e-81
AT4G11900 338 / 7e-104
S-locus lectin protein kinase family protein (.1)
Lus10033752
233 / 3e-75
AT4G03230 497 / 6e-169
S-locus lectin protein kinase family protein (.1)
Lus10036313
233 / 1e-74
AT4G03230 241 / 9e-71
S-locus lectin protein kinase family protein (.1)
Lus10007602
241 / 3e-74
AT1G65800 788 / 0.0
receptor kinase 2 (.1)
Lus10023391
244 / 4e-74
AT1G11300 520 / 5e-168
protein serine/threonine kinases;protein kinases;ATP binding;sugar binding;kinases;carbohydrate binding (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G111900
377 / 2e-126
AT3G16030 367 / 2e-114
CALLUS EXPRESSION OF RBCS 101, lectin protein kinase family protein (.1)
Potri.018G111700
367 / 9e-123
AT4G23200 363 / 1e-115
cysteine-rich RLK (RECEPTOR-like protein kinase) 12 (.1)
Potri.018G111800
365 / 3e-122
AT4G23200 362 / 4e-115
cysteine-rich RLK (RECEPTOR-like protein kinase) 12 (.1)
Potri.018G111751
307 / 3e-106
AT4G23290 255 / 2e-81
cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.1), cysteine-rich RLK (RECEPTOR-like protein kinase) 21 (.2)
Potri.018G111600
323 / 1e-105
AT4G23180 377 / 2e-120
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
Potri.005G014802
255 / 2e-82
AT1G11330 454 / 5e-152
S-locus lectin protein kinase family protein (.1.2)
Potri.011G125201
244 / 2e-81
AT4G27300 328 / 1e-107
S-locus lectin protein kinase family protein (.1)
Potri.004G023550
250 / 7e-79
AT4G05200 541 / 0.0
cysteine-rich RLK (RECEPTOR-like protein kinase) 25 (.1)
Potri.001G441400
251 / 2e-78
AT3G16030 523 / 1e-174
CALLUS EXPRESSION OF RBCS 101, lectin protein kinase family protein (.1)
Potri.011G027900
246 / 7e-78
AT4G23180 542 / 0.0
cysteine-rich RLK (RECEPTOR-like protein kinase) 10 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Lus10040020 pacid=23173490 polypeptide=Lus10040020 locus=Lus10040020.g ID=Lus10040020.BGIv1.0 annot-version=v1.0
ATGCTGGTGTACGAATTCATGCCCAACAAAAGCCTCGACCTCTATCTTCTGGATCCAGTGAGGAAGCTAAGGTTAGACTGGAACAAGAGGGTTAGGATCA
TTGAAGGAGTGACTCAGGGGCTATTGTACCTTCAGGAGTACTCAAACTTCACCATAATTCATAGGGATCTCAAGGCGAGCAACATCCTTCTGGATGATAC
CATGAACCCTAAGATCTCCGATTTTGGGATGGCCAAGCTCTTTAGGAAAGATATCGATGAAGCCAGCACTAGCAATATTGTGGGAACATATGGATACGTT
CCACCTGAATATGTCAAGAAAGGGATATACTCTATGAAATACGATGTCTACAGCTTCGGAGTTTTGCTCCTGCAAATTGTCAGTGGTAAAAGAACCAGAC
TTTACTATGGTCCCAATCAAGACATGCATCTCTTGGAACATGCTTACGAATGTTGGAAGGGTGGTAAAGGAACCGACTTCTTTGACCCGTCATTAGACGA
TTCGGCATCACAATGTAAGCTGGTGACATGTCTACAAGTGGCGTTGCTGTGCGTTCAGGAGAATCCTGACGAGAGGCCGACAATGTTGGAAGTGTCATCT
GCACTGAGAAATGGTGCAGCAGCTTTGGAAATGCCTGTTCCAAAAAGACCAGCCTTTTCCGTGTCTAAGGTAAGCGAAAAGGGATCTGCAACGAGTTCCA
GGCCAGAAACTACTTCGTTCAACGACGCTGAAATATCTCAACTAGAACCTCGTTGA
AA sequence
>Lus10040020 pacid=23173490 polypeptide=Lus10040020 locus=Lus10040020.g ID=Lus10040020.BGIv1.0 annot-version=v1.0
MLVYEFMPNKSLDLYLLDPVRKLRLDWNKRVRIIEGVTQGLLYLQEYSNFTIIHRDLKASNILLDDTMNPKISDFGMAKLFRKDIDEASTSNIVGTYGYV
PPEYVKKGIYSMKYDVYSFGVLLLQIVSGKRTRLYYGPNQDMHLLEHAYECWKGGKGTDFFDPSLDDSASQCKLVTCLQVALLCVQENPDERPTMLEVSS
ALRNGAAALEMPVPKRPAFSVSKVSEKGSATSSRPETTSFNDAEISQLEPR
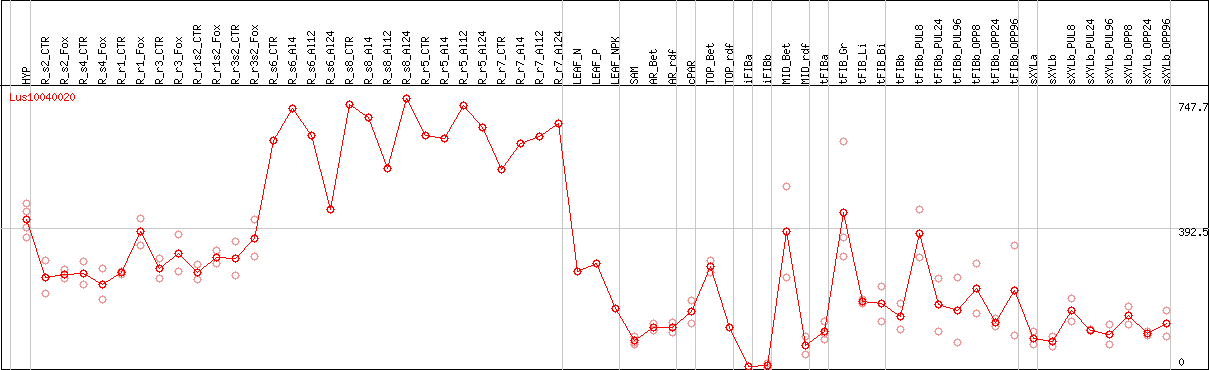
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040020 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.