External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G02090 243 / 3e-81
ATCSN7, COP15, CSN7II, FUS5
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004367
400 / 9e-143
AT1G02090 265 / 1e-89
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
Lus10028205
235 / 9e-78
AT1G02090 355 / 2e-125
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
Lus10042908
221 / 1e-71
AT1G02090 385 / 7e-136
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G137400
251 / 5e-84
AT1G02090 381 / 6e-135
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
Potri.014G047200
241 / 5e-80
AT1G02090 394 / 5e-140
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
Potri.003G081066
167 / 2e-52
AT1G02090 179 / 1e-57
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
Potri.001G153900
81 / 4e-19
AT1G02090 87 / 1e-21
FUSCA 5, CONSTITUTIVE PHOTOMORPHOGENIC 15, ARABIDOPSIS THALIANA COP9 SIGNALOSOME SUBUNIT 7, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2), Proteasome component (PCI) domain protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF01399
PCI
PCI domain
Representative CDS sequence
>Lus10040162 pacid=23173990 polypeptide=Lus10040162 locus=Lus10040162.g ID=Lus10040162.BGIv1.0 annot-version=v1.0
ATGGAAATTGAGCAGCGACAAGCTCCATTCATCGAGAAGTTCGTGGAGCAAGCCTCCGCCGTCGTCGACGGTCCTGCTCTCACGCGAATCGTCGTGGAAT
CCACCTCTCACCCTTCGGTTTTTGCCTTCTCCGATATTTTGTCCGTCCCCAGTTTCGACAAGCTTCAAGGAACCGCGGATTCGGCATTCGTCTATCTTCT
TAAACTATTCGCGTATGGCACATGGCGGGACTACAAAGCTAAATCTAGCGAACAGCCACAGCTGACGCCGAATCAAATCATCAAACTGAAGCAACTTACA
GTGCTTACTCTTGCTGAGGGAACCAAGGTTATGTCCTATGATACGCTTTTGGAGGAACTTGAATTCTCTAACATCCGTGAACTTGAAGATTTCCTTATTG
ATGAGTGTGTGTATGCGGGCATAATCAAAGGGAAGGTGAATCAATTAAGGAGATGCTTTGAGGTTAAATTTGCAGCCGCAAGGGACCTGATGCCTTGTGA
ATTTGATTCTGTGACAGAGACAACAAGAAATTGGCTGGAAACTTCAGGTAATATGCTAGCCCTTCTTGAAAAGCAGATTGACAAGGCTAATGCGATGGAG
GAATTGGAAACGAAGCATGGAATGCCAACTCGAAGCGGGGACGCAATGTTTCCAGAAGGTGAAGGGTGTTGTTACAGAGGCTCTCGTCCTCCTGCAATTC
GTTTCTTGGATGAGTTCTTTTGA
AA sequence
>Lus10040162 pacid=23173990 polypeptide=Lus10040162 locus=Lus10040162.g ID=Lus10040162.BGIv1.0 annot-version=v1.0
MEIEQRQAPFIEKFVEQASAVVDGPALTRIVVESTSHPSVFAFSDILSVPSFDKLQGTADSAFVYLLKLFAYGTWRDYKAKSSEQPQLTPNQIIKLKQLT
VLTLAEGTKVMSYDTLLEELEFSNIRELEDFLIDECVYAGIIKGKVNQLRRCFEVKFAAARDLMPCEFDSVTETTRNWLETSGNMLALLEKQIDKANAME
ELETKHGMPTRSGDAMFPEGEGCCYRGSRPPAIRFLDEFF
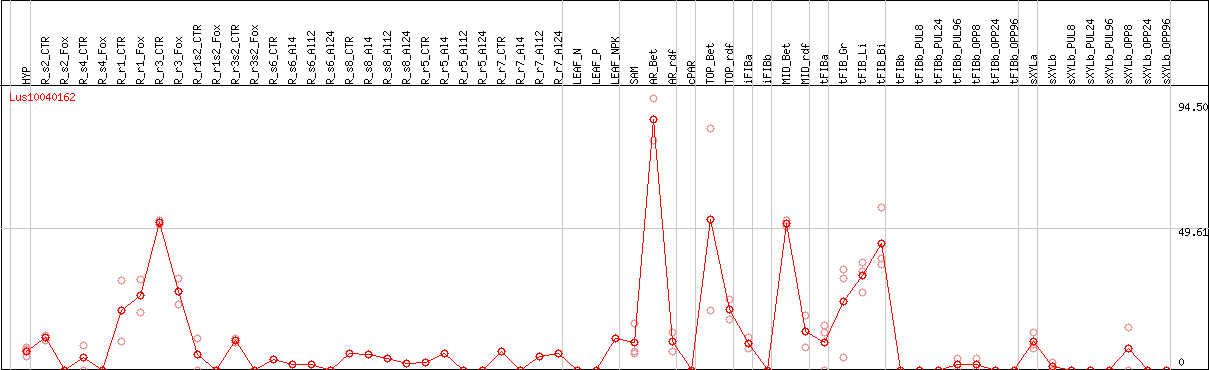
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040162 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.