External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G06080 126 / 3e-37
AS2
LBD33
LOB domain-containing protein 33 (.1)
AT3G58190 121 / 6e-35
AS2
LBD29, ASL16
ASYMMETRIC LEAVES 2-LIKE 16, lateral organ boundaries-domain 29 (.1)
AT2G45420 119 / 2e-33
AS2
LBD18, ASL20
LOB domain-containing protein 18 (.1)
AT3G03760 118 / 4e-33
AS2
LBD20
LOB domain-containing protein 20 (.1)
AT2G42430 116 / 2e-32
AS2
ASL18, LBD16
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
AT4G00210 112 / 2e-31
AS2
LBD31
LOB domain-containing protein 31 (.1)
AT2G42440 111 / 8e-31
AS2
Lateral organ boundaries (LOB) domain family protein (.1)
AT4G00220 108 / 7e-30
AS2
JLO, LBD30, ASL19
LOB DOMAIN-CONTAINING PROTEIN 30, JAGGED LATERAL ORGANS, Lateral organ boundaries (LOB) domain family protein (.1)
AT2G45410 105 / 6e-29
AS2
LBD19
LOB domain-containing protein 19 (.1)
AT2G31310 102 / 9e-28
AS2
LBD14
LOB domain-containing protein 14 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004707
241 / 2e-82
AT5G06080 154 / 1e-47
LOB domain-containing protein 33 (.1)
Lus10000613
118 / 1e-33
AT3G58190 215 / 1e-70
ASYMMETRIC LEAVES 2-LIKE 16, lateral organ boundaries-domain 29 (.1)
Lus10033873
117 / 2e-33
AT3G58190 220 / 1e-72
ASYMMETRIC LEAVES 2-LIKE 16, lateral organ boundaries-domain 29 (.1)
Lus10000707
117 / 7e-33
AT2G42430 222 / 2e-72
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10024877
114 / 1e-31
AT2G42430 211 / 6e-68
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10009298
114 / 1e-31
AT2G45410 193 / 2e-62
LOB domain-containing protein 19 (.1)
Lus10033872
111 / 1e-30
AT2G42430 212 / 6e-69
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10014757
109 / 6e-30
AT2G42430 208 / 2e-67
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10009930
98 / 9e-26
AT2G30340 230 / 5e-76
LOB domain-containing protein 13 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03195
LOB
Lateral organ boundaries (LOB) domain
Representative CDS sequence
>Lus10040274 pacid=23173852 polypeptide=Lus10040274 locus=Lus10040274.g ID=Lus10040274.BGIv1.0 annot-version=v1.0
ATGCAGGAGTGTGTTTTTGCACCTTACTTCAGTTATAACCAGGCTCCATCCCATTTCGGGGCAGTGCATAAGGTGTTTGGTGCAAGCAATGTTTCCAAGC
TACTACTTCGCTTACCGGTCAACTTTAGAAGCGATGCTGCCATCACTATTGCTTACCAAGCATTGGCACGTATCCAGGATCCAGCATATGGATGTGTGGC
TTACATCATGGCCCTCCAGCAACAGGTCGCCAGCATACAGCAAGAGATCGAAACTCTTAGCCACCAATTGGCTTTGAACTCGGTCACCCAGCTTGCCAGC
TGTGGTAGTTTACCTGCAAGTTCTGCATCGAACTACGAGACACAAATTTTCCAGCAGAATCAACAAAGTTTACAGCTTGACAATTCAGAACATATTGCTG
GAAGCATAGAGTTCGATAGCAAAATCACATCAGAGCTTTTGTCTCTGTACGGATGGGAAGACCAGAACCCTTTCTGGAACTCTGCACATCACACTTAG
AA sequence
>Lus10040274 pacid=23173852 polypeptide=Lus10040274 locus=Lus10040274.g ID=Lus10040274.BGIv1.0 annot-version=v1.0
MQECVFAPYFSYNQAPSHFGAVHKVFGASNVSKLLLRLPVNFRSDAAITIAYQALARIQDPAYGCVAYIMALQQQVASIQQEIETLSHQLALNSVTQLAS
CGSLPASSASNYETQIFQQNQQSLQLDNSEHIAGSIEFDSKITSELLSLYGWEDQNPFWNSAHHT
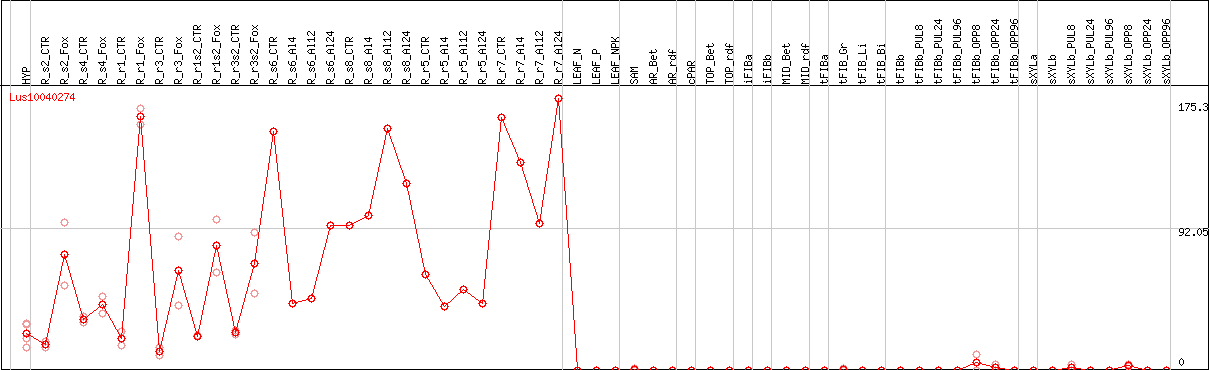
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040274 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.