Lus10040524 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10040524 pacid=23157794 polypeptide=Lus10040524 locus=Lus10040524.g ID=Lus10040524.BGIv1.0 annot-version=v1.0
ATGTTGCTGCAAATACCGCGTTTGACGAACTCTCTGAGGGAGCCATTCGATGTAGACCAGGCCTATCTCCATCGGAAAATCATCCTCCACAACCACTTCA
AGCCTAGCAAACATGGAAACTCCCTGAATGAGTCGGATCTTGCTAGGAAGATAGTTATTGGATGGGATGAAGCATCAACAGAAGTGCGACAGGTGTACAA
ACAATTTATTGGTGCTGTGGTGGAACTAATAGATGCCGAAGTTCCTTCCGAGGAGTTCAGAGAGGTGGCATTGACAGCTTACCGTCTCTTTGCCTTTCCA
GGGGCTGAAAAGGAGGAGGTTTCAGCCGGCATTATCAAGAGGGATAAGTCTGAATTGGAGCAAATTGTTGGTCATGCAACTAATGGTAGCCTGCAGAAAG
TTGCAGCCCTAGCAGAGAGACTCTCTGGTTTGCAATCTAGCAATTATGGTGCTGCATTCGCTCTTGAAAGTCGTCGAGATGGGTCGGAAAATGATTTCGA
GTTTGGAACCGACCTTCTCTTTCAAGCCCCGGCTAGGTTTTTGGTGGATGTCACTTTGGACGATGGAGAATTATTGGGCGAGGAAAGCACTGCACCATTT
TCAACGTTACGTGAAAGATTGCATGACCATAGAGACCATGTACATAATCATTCAACTCTTGGTGGGAACTTCAATTTAAGTTGGTTAAGAAATGCATGTG
ACCGCATAGCTAGTGAAAGTACCTCACAGCTTTCTAGGGATGATCTAGCAATGGCAATATGTCGTGTATTAGATTCAGGAAGACCGGGAGAGGAGATTGC
TGATGTTTTGCTGGATCTTGTTGGGGATGGTGCATTTGAAATTGTTCAAGATCTTATCTCACACCGGAAAGAACTTGTCGATGCAATTCAGCATGGATTT
TCAGTATTGAAATCAGATAAAATGGCTTCTAACAACCAATCACGGATGCCTAGTTATGGCACCCAGGTGACTGTACAAACAGAGTCAGAAAAGCAAATTG
ATAAACTCCGTCGCAAGGAAGAAAAAAGACATAGACGCGGAACAGAATATGGGGAGAATGATACATCAGCTACAACCTTTTCTTCATTACTTGAAGCAAG
TGAAAGAAAGACCCTGCTTGATGATCTGGTCAATTCAGGTCCGGGGACCCTTCAATCGGCAGTAACTGCTCTGCCTCAAGGTACTGTCAGGAAGCATTTC
AAAGGATATGAGGAAGTCATTATCCCACCAACACCAACTGCAGCCATGAAACCTGGAGAAAAGCTGGTTTGTGCACCAACAGGTGCTGGCAAAACAAATA
TAGCCATGATATCAATTCTACATGAGATTGGACAGCATTTCAAGGATGGCTACTTGCATAAGAATGAATTCAAAATAGTTTATGTTGCCCCTATGAAGGC
TTTAGCTGCAGAGGTTACGTCAACGTTTGGCCGTCGTTTAGCTCCACTGAATATGATTGTGAGAGAACTCACTGGGGACATGCAGCTCTCTAAGAATGAA
CTCGAAGAAACACAGATGATAGTGACAACTCCAGAGAAATGGGATGTCATTACTCGTAAGAGCAGTGACATGTCACTTTCAATGCTTGTAAAGCTCTTGA
TCATTGATGAAGTTCATCTTCTGAATGATGATAGAGGGCCTGTGATAGAGGCACTGGTAGCCAGGACTCTGCGTCAGGTGGAGTCAACACAGACAATGAT
TCGCATCGTGGGACTTTCAGCCACTCTACCTAATTACATGGAGGTCGCTCAGTTTCTCAGAGTGAATCCAGAAGCAGGTCTCTTCTTCTTTGATTCTAGT
TACCGTCCAGTGCCACTTGCACAACAATATATTGGAATTAGTGAACAGAACTTTGCAGCTCGCAATGAGTTACTAAATGAAGTTTGTTATAAGAAGGTTG
TTGATTCCCTAAAGCAAGGCCATCAAGCAATGGTATTCGTTCACTCTCGTAAGGACACAGCAAAAACAGCTGACAAATTGGTTGAACTTGCAAGGAAATA
TGAGGAAATGGAACTCTTTTGCAATGATAACCATCCGCAGTTTGGATTGATTAAGAAAGAAGTCTCGAAGTCCAGAAACAAAGATCTTGTTCAACTTTTT
GAATCTGCAGTTGGTGTTCATCATGCGGGGATGTTACGTTCTGATAGAGGCTTGACTGAACGGCTCTTCTCTGATGGACTATTGAAGGTACTTGTTTGCA
CAGCAACTTTGGCCTGGGGAGTGAATCTACCTGCTCACACTGTCGTTATCAAGGGGACTCAATTATATGACCCAAAAGCTGGTGGCTGGCGAGACCTTGG
CATGCTTGATGTCATGCAGATTTTTGGAAGGGCTGGAAGACCTCAGTTTGACAAGAGTGGTGAAGGTATTATAATCACATCTCATGAGAAACTGGCGTAT
TATCTGAGATTGCTGACAAGTCAGCTTCCAATTGAAAGTCAGTTTATTAGTTCCTTGAAGGACAACTTAAATGCTGAGGTGGCTTTGGGCACAGTAACTA
ATGTTAAGGAAGCTAGTGCGTGGCTTGGATATACGTATCTTTTCATCCGAATGCGGAAAAATCCTCTGGTTTATGGTATTGGGTGGGATGAGGTCATTGC
TGATCCTTCTTTGGGCTTAAAGAAAAGAGCTCTTATAACGGATGCTGCCCGTGCACTTGACAAGGCTAAGATGATGAGATTTGACGAAAAAAGTGGCAAC
TTTTATTGTACAGAACTCGGGCGTATTGCTAGCCACTTCTATATCCAGTATTCCAGTGTCGAAACATACAATGAAATGTTGAGACGCCACATGAGCGATA
GTGAGGTAATTGAAATGGTTGCACGGTCTTCCGAGTTTGAGAATATTGTTGTTCGAGAGGAGGAGCAGAATGAGTTAGAGTCGATTCTGCGTACGTCATG
CCCTTTAGAAGTTAAGGGGGGTCCTTCTAACAAATATGGAAAGATTTCAATACTCATCCAGTTGTATATATCCCGGGGTTCAATCGATACCTTTTCTTTG
GTCTCGGATGCTGCATATATTAGTGCAAGCTTGGCTCGAATTATGCGAGCATTATTTGAGATCTGTTTACGCAAGGGCTGGTGCGAGATGACGTTGTTCA
TGTTGGAGTATTGCAAAGCTGTGGATCGCCAGATGTGGCCATACCATCATCCTCTGAGACAATTTGACAGAGATCTTTCAGCAGAAATACTCAGAAAGCT
TGAAGAACGTGGGGTTGATTTAGACCATCTGCTGGAGATGGAGGAAAAAGATATTGGAGCGCTTATTCGTTATGCGCATGGTGGACGGTTGGTCAAGCAA
CACTTGGGATACTTCCCCCAAATCCAGCTTTCAGCTACAATAAGCCCAATAACCAGAACTGTCTTGAAGGTGGATTTGCTTGTGACTCCTGATATCATGT
GGAAAGAGCGATTTCATGGCACTGCATTACGTTGGTGGATTCTCGTTGAGGATTCTGAGAATGATCACATCTACCACTCGGAGCTTTTCACTCTAACAAA
GCGGATGGCTCGTGGAGAACCTCAGAAGCTTTCATTCACAGTGCCTATTTTTGAACCACATCCACCGCAATACTACATACGGGCAGTCTCTGATTCATGG
TTGCATGCTGAATCTTTTCACACTGTGACTTTCAAGAATCTACAGTTGCCAGAGGCTCGTACCTCTCACACAGAACTTCTAGATCTGAAACCTCTTCCTG
TGACTGCTCTTGGGAATGAGACTTTCCTATCCCTGTACAAATTTTCACATTTTAATCCTATACAGACGCAGATTTTCCATGTACTGTACCACTCAGACAA
CAACGTCCTTTTAGGAGCACCTACTGGAAGTGGGAAAACTATTTCTGCTGAACTTGCTATGCTTAGACTTTTCAGTACTCAGCCTGATATGAAGGTGATT
TATATAGCACCTCTAAAGGCTATAGTCAGGGAAAGAATGCACGATTGGCAAAAGCATCTCGTCCCTTCACTGGGGAAAAAGATGGTCGAAATGACTGGAG
ATTTTACTCCAGATTTGACGGCCCTCTTGTCAGCAGACATTATTATATCTACTCCTGAAAAGTGGGATGGCATTAGTCGTAACTGGCAGACTCGAAGCTA
TGTTACCAAGGTAGGACTTATGATTCTGGATGAGATTCACCTGCTGGGTGCTGATCGTGGACCCATCCTTGAGGTAATAGTCTCAAGAATGAGATATATC
TCTTCACAAACGGAGCGTGCCGTGCGTTTTGTAGGTTTGTCAACAGCATTGGCAAATGCAAGGTTTCTTGCTAGCTTAAACATTAAAGCATGCACTGGTG
AAGGGATGTATTGTTTACATGACAAAAGTGTACTTGACCATGGATATCCTGGAAAATACTACTGCCCAAGAATGAACAGCATGAATAAACCAGCCTATGC
TGCCATCTCCTCACATTCACCTACAAAACCAGTACTCATATTTGTTTCTTCTCGTCGTCAAACAAGGCTCACTGCCCTTGACCTTATTCAGTATGCAGCT
TCTGATGAACATCCAATACAGTTTCTGAGTATGGAAGAAGACGACCTTCAAATGGTTCTTTCACAGGTTATGGACCAGAATTTGAGGCACACATTGCAGT
TCGGCATAGGTCTGCATCATGCAGGGCTAAATGACAAAGATAGATCTCTGGTTGAGGAACTTTTTGCAAACAACAAGATCCAGGTACTGGTTTGCACTAG
TACGTTGGCATGGGGAGTAAATCTTCCTGCTCATTTGGTTATTATCAAGGGAACAGAATATTTTGATGGAAAATCAAAGAGATATGTAGATTTCCCAATC
ACAGACATCCTACAGATGATGGGTCGTGCTGGTCGGCCACAGTTTGACCAACATGGGAAGGCAGTCATCCTCGTTCATGAGCCAAAAAAGAGCTTTTATA
AAAAGTTTCTATATGAACCGTTTCCTGTAGAGAGTAGTCTGAGGGAGCAGTTGCACAATCACTTCAATGCTGAGATAGCAACAGGCACAATCTGTCACAA
AGAGGATGCAGTTCATTACCTGACATGGACATACTTATTCCGTAGACTGACGGTTAATCCGGCCTACTATGGTTTAGAAAGCACAGAGGCAGAGATGTTA
AGTTCGTATTTGTCAAGATTAGTGCAGAACACTTTTGAGGATTTGGAAGATGGTGGATGCATCAAGATGGATGAGAACACTGTGGAGTGCACGATGTTGG
GTATGATTGCGTCGCAATATTACCTCAGCTACATGACCGTGTCAATGTTTGGGTCAAATATAGGGCCAGATACATCTCTGGAGGTGTTTCTTCACATTCT
ATCTGGAGCATCGGAATATGATGAACTGCCTGTGCGGCATAATGAGGAAAAATTTAATGAAGCTCTGTCAGAGAAAGTGCCATATATGGTGGATAAGAAT
AACCTTGATGATCCTCATGTCAAAGCAAACTTACTCTTTCAGGCACATTTTTCCCAGTTGGAATTGCCAATTAGTGACTATATTACTGACTTAAAGTCAG
TATTGGACCAAAGCATACGCATCATGCAAGCCATGATAGATGTATGTGCCAACAGTGGATGGCTTTCAAGCTCCATTACTTGTATGCATCTTATGCAAAT
GGTGATGCAGGGTTTATGGTTTGAGCGCGATTCTGCCTTGTGGATGCTGCCTTGCATGAACACAGATCTTTATGCCGTGCTGAACAGTCGAGGGATTTCC
AGTGTACATCAACTATTGGATATTCCCAAGTCGAGCTTGCAGGATTTGATTGGAAATTTCCCTTTCTCGAGGTTAAACAAGGATCTGCAACAATTTCCTC
GGATTCAAGTCAAACTGAAACTCCAGAAAAAAGGCACCGGTGGCAGTGCTTTGAATATTAGACTCGAGAAGATCAACTTCAGACAAAGCACGCCAAGGGC
TTTGGCTCCTAGGTTCCCCAAGGTAAAAGACGAGGCATGGTGGTTAGTTCTTGGCAATACAGCAACTTCTGAACTTTATGCTCTGAAGAGGGTTTCTTTC
ACCGATCGTTTGTCGACGCGCATGGACTTGCCACCATCTTTGACGTCTGTCCAGGGCTCGAAGTTGATCCTGGTTTCAGATTGTTACCTTGGTTTCGAAC
AAGAATACTCGATCGAAGACTTCCCAATGTCTCAATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10040524 pacid=23157794 polypeptide=Lus10040524 locus=Lus10040524.g ID=Lus10040524.BGIv1.0 annot-version=v1.0
MLLQIPRLTNSLREPFDVDQAYLHRKIILHNHFKPSKHGNSLNESDLARKIVIGWDEASTEVRQVYKQFIGAVVELIDAEVPSEEFREVALTAYRLFAFP
GAEKEEVSAGIIKRDKSELEQIVGHATNGSLQKVAALAERLSGLQSSNYGAAFALESRRDGSENDFEFGTDLLFQAPARFLVDVTLDDGELLGEESTAPF
STLRERLHDHRDHVHNHSTLGGNFNLSWLRNACDRIASESTSQLSRDDLAMAICRVLDSGRPGEEIADVLLDLVGDGAFEIVQDLISHRKELVDAIQHGF
SVLKSDKMASNNQSRMPSYGTQVTVQTESEKQIDKLRRKEEKRHRRGTEYGENDTSATTFSSLLEASERKTLLDDLVNSGPGTLQSAVTALPQGTVRKHF
KGYEEVIIPPTPTAAMKPGEKLVCAPTGAGKTNIAMISILHEIGQHFKDGYLHKNEFKIVYVAPMKALAAEVTSTFGRRLAPLNMIVRELTGDMQLSKNE
LEETQMIVTTPEKWDVITRKSSDMSLSMLVKLLIIDEVHLLNDDRGPVIEALVARTLRQVESTQTMIRIVGLSATLPNYMEVAQFLRVNPEAGLFFFDSS
YRPVPLAQQYIGISEQNFAARNELLNEVCYKKVVDSLKQGHQAMVFVHSRKDTAKTADKLVELARKYEEMELFCNDNHPQFGLIKKEVSKSRNKDLVQLF
ESAVGVHHAGMLRSDRGLTERLFSDGLLKVLVCTATLAWGVNLPAHTVVIKGTQLYDPKAGGWRDLGMLDVMQIFGRAGRPQFDKSGEGIIITSHEKLAY
YLRLLTSQLPIESQFISSLKDNLNAEVALGTVTNVKEASAWLGYTYLFIRMRKNPLVYGIGWDEVIADPSLGLKKRALITDAARALDKAKMMRFDEKSGN
FYCTELGRIASHFYIQYSSVETYNEMLRRHMSDSEVIEMVARSSEFENIVVREEEQNELESILRTSCPLEVKGGPSNKYGKISILIQLYISRGSIDTFSL
VSDAAYISASLARIMRALFEICLRKGWCEMTLFMLEYCKAVDRQMWPYHHPLRQFDRDLSAEILRKLEERGVDLDHLLEMEEKDIGALIRYAHGGRLVKQ
HLGYFPQIQLSATISPITRTVLKVDLLVTPDIMWKERFHGTALRWWILVEDSENDHIYHSELFTLTKRMARGEPQKLSFTVPIFEPHPPQYYIRAVSDSW
LHAESFHTVTFKNLQLPEARTSHTELLDLKPLPVTALGNETFLSLYKFSHFNPIQTQIFHVLYHSDNNVLLGAPTGSGKTISAELAMLRLFSTQPDMKVI
YIAPLKAIVRERMHDWQKHLVPSLGKKMVEMTGDFTPDLTALLSADIIISTPEKWDGISRNWQTRSYVTKVGLMILDEIHLLGADRGPILEVIVSRMRYI
SSQTERAVRFVGLSTALANARFLASLNIKACTGEGMYCLHDKSVLDHGYPGKYYCPRMNSMNKPAYAAISSHSPTKPVLIFVSSRRQTRLTALDLIQYAA
SDEHPIQFLSMEEDDLQMVLSQVMDQNLRHTLQFGIGLHHAGLNDKDRSLVEELFANNKIQVLVCTSTLAWGVNLPAHLVIIKGTEYFDGKSKRYVDFPI
TDILQMMGRAGRPQFDQHGKAVILVHEPKKSFYKKFLYEPFPVESSLREQLHNHFNAEIATGTICHKEDAVHYLTWTYLFRRLTVNPAYYGLESTEAEML
SSYLSRLVQNTFEDLEDGGCIKMDENTVECTMLGMIASQYYLSYMTVSMFGSNIGPDTSLEVFLHILSGASEYDELPVRHNEEKFNEALSEKVPYMVDKN
NLDDPHVKANLLFQAHFSQLELPISDYITDLKSVLDQSIRIMQAMIDVCANSGWLSSSITCMHLMQMVMQGLWFERDSALWMLPCMNTDLYAVLNSRGIS
SVHQLLDIPKSSLQDLIGNFPFSRLNKDLQQFPRIQVKLKLQKKGTGGSALNIRLEKINFRQSTPRALAPRFPKVKDEAWWLVLGNTATSELYALKRVSF
TDRLSTRMDLPPSLTSVQGSKLILVSDCYLGFEQEYSIEDFPMSQ
|
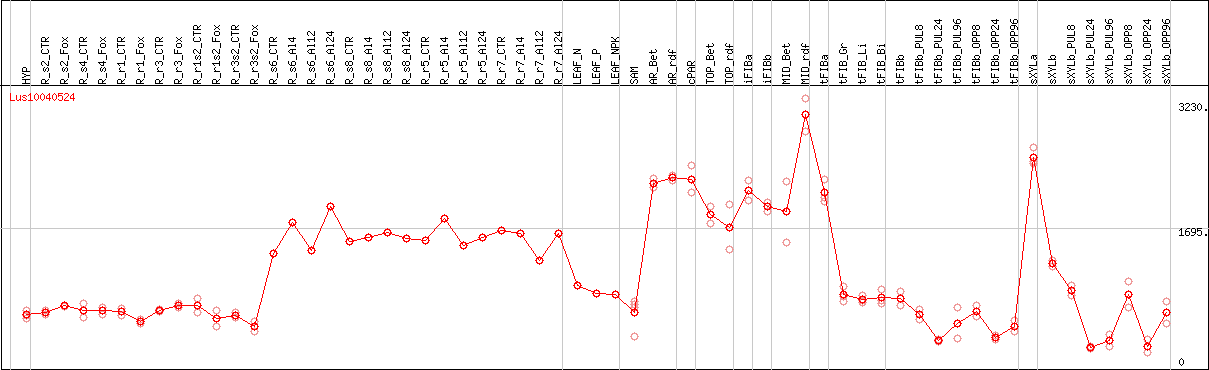
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040524 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
