External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G10940 218 / 9e-68
ASG2
ALTERED SEED GERMINATION 2, transducin family protein / WD-40 repeat family protein (.1.2)
AT4G35140 66 / 6e-13
Transducin/WD40 repeat-like superfamily protein (.1)
AT4G38480 64 / 2e-12
Transducin/WD40 repeat-like superfamily protein (.1)
AT3G45620 64 / 2e-12
Transducin/WD40 repeat-like superfamily protein (.1.2)
AT5G56130 49 / 4e-07
THO3, AtTEX1
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G67320 42 / 0.0001
HOS15
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
AT1G64610 40 / 0.0005
Transducin/WD40 repeat-like superfamily protein (.1.2)
AT2G43770 39 / 0.0005
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G58230 39 / 0.0008
MSI1, MEE70, ATMSI1
MATERNAL EFFECT EMBRYO ARREST 70, ARABIDOPSIS MULTICOPY SUPRESSOR OF IRA1, Transducin/WD40 repeat-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033290
62 / 2e-11
AT3G45620 557 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10034751
61 / 2e-11
AT3G45620 556 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10025088
61 / 3e-11
AT4G35140 563 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Lus10023993
61 / 3e-11
AT4G35140 557 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Lus10002860
43 / 3e-05
AT2G43770 617 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Lus10012230
43 / 4e-05
AT2G43770 620 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Lus10015407
43 / 4e-05
AT3G49180 169 / 3e-49
ROOT INITIATION DEFECTIVE 3, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10043473
43 / 5e-05
AT1G15440 1058 / 0.0
\(PERIODIC TRYPTOPHAN PROTEIN 2, periodic tryptophan protein 2 (.1.2)
Lus10027113
42 / 6e-05
AT5G56130 555 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G158300
224 / 6e-70
AT5G10940 1080 / 0.0
ALTERED SEED GERMINATION 2, transducin family protein / WD-40 repeat family protein (.1.2)
Potri.002G103132
162 / 9e-51
AT5G10940 390 / 2e-132
ALTERED SEED GERMINATION 2, transducin family protein / WD-40 repeat family protein (.1.2)
Potri.002G103216
95 / 1e-25
AT5G10940 91 / 4e-22
ALTERED SEED GERMINATION 2, transducin family protein / WD-40 repeat family protein (.1.2)
Potri.009G139000
64 / 2e-12
AT4G35140 602 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.004G178700
64 / 2e-12
AT4G35140 538 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.001G470900
45 / 7e-06
AT5G56130 586 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.019G096900
42 / 7e-05
AT2G43770 645 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.005G144400
39 / 0.0008
AT5G67320 671 / 0.0
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Lus10040537 pacid=23157918 polypeptide=Lus10040537 locus=Lus10040537.g ID=Lus10040537.BGIv1.0 annot-version=v1.0
ATGGACCAACACGGGTTTCACGACGGCAATTTCTACAATTTGCTGGAGAGCAGATGCTTGGACGTTCGACATGACATTGTTGGCAACTTTCAGATGCATT
CATCATTGATGCGGAGGCTCTCGCAAGAGAAGGAAATGGAAGGGCATCAGGGTTGCGTCAATGCTATAGCTTGGAATTCGAATGGTTCCCTCTTGATTTC
TGGGTCAGATGATACCAGGGTTAGGGTATTCAATTTGTCTCGATTAAGCGGAAGAGGTCCTGAGGATACTAGCATTGTTCCTTCTGCACTCTACCAATGT
CACTCTAGAAGAGTAAAGAAATTAGCTATTGAAGTTGGAAATCCGAATGTGGTATGGAGTGCCAGTGAAGATGGAACTTTGAGACAGCATGACCTTAGGG
AGTGTGCTTCTTGTCCACCAGCAGGATCTTCTCCTCAAGAATGCCGCAATATTTTGGTGAGTTTTGGCTGA
AA sequence
>Lus10040537 pacid=23157918 polypeptide=Lus10040537 locus=Lus10040537.g ID=Lus10040537.BGIv1.0 annot-version=v1.0
MDQHGFHDGNFYNLLESRCLDVRHDIVGNFQMHSSLMRRLSQEKEMEGHQGCVNAIAWNSNGSLLISGSDDTRVRVFNLSRLSGRGPEDTSIVPSALYQC
HSRRVKKLAIEVGNPNVVWSASEDGTLRQHDLRECASCPPAGSSPQECRNILVSFG
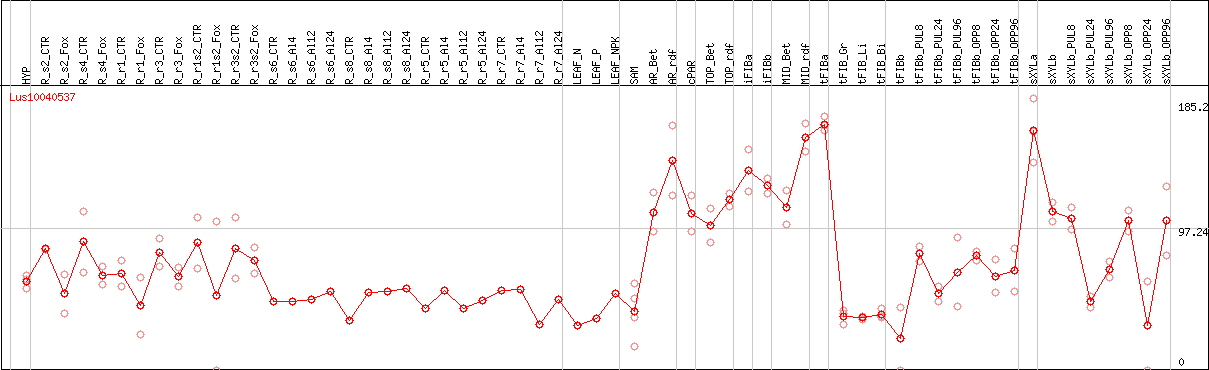
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040537 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.