External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G26150 55 / 2e-08
GATA
GATA22, CGA1, GNL
GNC-LIKE, GATA TRANSCRIPTION FACTOR 22, cytokinin-responsive gata factor 1 (.1)
AT5G56860 55 / 2e-08
GATA
GATA21, GNC
GATA, nitrate-inducible, carbon metabolism-involved, GATA TRANSCRIPTION FACTOR 21, GATA type zinc finger transcription factor family protein (.1)
AT5G26930 46 / 3e-06
GATA
GATA23
GATA transcription factor 23 (.1)
AT3G50870 45 / 3e-05
GATA
GATA18, HAN, MNP
MONOPOLE, HANABA TANARU, GATA TRANSCRIPTION FACTOR 18, GATA type zinc finger transcription factor family protein (.1)
AT4G36620 44 / 3e-05
GATA
GATA19, HANL2
hanaba taranu like 2, GATA transcription factor 19 (.1)
AT5G49300 43 / 4e-05
GATA
GATA16
GATA transcription factor 16 (.1)
AT2G18380 42 / 0.0001
GATA
GATA20, HANL1
hanaba taranu like 1, GATA transcription factor 20 (.1)
AT3G16870 42 / 0.0002
GATA
GATA17
GATA transcription factor 17 (.1)
AT4G16141 41 / 0.0003
GATA
GATA type zinc finger transcription factor family protein (.1)
AT1G08010 41 / 0.0005
GATA
GATA11
GATA transcription factor 11 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006000
256 / 3e-83
AT4G26150 79 / 7e-16
GNC-LIKE, GATA TRANSCRIPTION FACTOR 22, cytokinin-responsive gata factor 1 (.1)
Lus10016849
44 / 3e-05
AT3G06740 154 / 6e-49
GATA transcription factor 15 (.1)
Lus10037721
44 / 4e-05
AT3G06740 153 / 2e-48
GATA transcription factor 15 (.1)
Lus10029863
43 / 0.0001
AT4G16141 90 / 1e-22
GATA type zinc finger transcription factor family protein (.1)
Lus10041746
43 / 0.0002
AT3G50870 93 / 8e-22
MONOPOLE, HANABA TANARU, GATA TRANSCRIPTION FACTOR 18, GATA type zinc finger transcription factor family protein (.1)
Lus10028301
42 / 0.0002
AT3G50870 135 / 4e-39
MONOPOLE, HANABA TANARU, GATA TRANSCRIPTION FACTOR 18, GATA type zinc finger transcription factor family protein (.1)
Lus10020684
41 / 0.0003
AT4G16141 94 / 9e-24
GATA type zinc finger transcription factor family protein (.1)
Lus10010027
39 / 0.0009
AT5G49300 87 / 8e-23
GATA transcription factor 16 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G229200
52 / 2e-07
AT4G26150 111 / 3e-28
GNC-LIKE, GATA TRANSCRIPTION FACTOR 22, cytokinin-responsive gata factor 1 (.1)
Potri.018G053600
52 / 2e-07
AT4G26150 75 / 4e-15
GNC-LIKE, GATA TRANSCRIPTION FACTOR 22, cytokinin-responsive gata factor 1 (.1)
Potri.005G020500
45 / 1e-05
AT3G06740 67 / 1e-14
GATA transcription factor 15 (.1)
Potri.008G213900
43 / 3e-05
AT3G06740 126 / 4e-38
GATA transcription factor 15 (.1)
Potri.010G001300
43 / 4e-05
AT3G06740 135 / 2e-41
GATA transcription factor 15 (.1)
Potri.002G199800
42 / 0.0001
AT5G49300 112 / 1e-32
GATA transcription factor 16 (.1)
Potri.005G122700
42 / 0.0002
AT3G50870 198 / 1e-62
MONOPOLE, HANABA TANARU, GATA TRANSCRIPTION FACTOR 18, GATA type zinc finger transcription factor family protein (.1)
Potri.007G024500
42 / 0.0002
AT3G50870 202 / 5e-64
MONOPOLE, HANABA TANARU, GATA TRANSCRIPTION FACTOR 18, GATA type zinc finger transcription factor family protein (.1)
Potri.014G124400
40 / 0.0005
AT5G49300 106 / 2e-30
GATA transcription factor 16 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0167
Zn_Beta_Ribbon
PF00320
GATA
GATA zinc finger
Representative CDS sequence
>Lus10040556 pacid=23157655 polypeptide=Lus10040556 locus=Lus10040556.g ID=Lus10040556.BGIv1.0 annot-version=v1.0
ATGATGAATTCAAATTATCAATATTCCTCCCACCAATTCCCTCCTTTCTCCATAGACCTTAACCAAGATCATCAGCACTACCTAAACGCTCATTACTTCA
CTACTGCTATTACAACAGCAACAACAACAACAAGTAGTTCACCATCTTCGTCTTCTTCTTCTCCTCTACCTAATTATCATATCTTCATCAACCCTCCTCC
TCCTGCTACCCAGTACTCTGGCGAAGATACCGATCCCGGTGGACACTTAGACTACTCATCCTGGGGGCTTCCTCCTCCAGCAGCAGCAGCAGCAGCAGCT
TCCTTCCACCAGTATCATCAACCTCCCCAGATCCTTCTCCAAAAGGGCGGCTCATCATGTATGTATGATCAGGAGATGGTGGAAGTAGTAGATTATAATC
ATAACGGATCGCTAGACAAGATGGAGAAAGAAATGAGTTATGATGTAGTGTCGCCCAAGTTGAGGATCATGAATCAGAAGAAGAAGAAGCTGACGATCAA
GACAAATGTTAAAAAACCCAATAACTTAAACGACGTCCAGTCGCCGTCAACTTTACAGCTCCAGCCTGCTGCTATGACTGACAACACCACCAATACTACT
ACTATGACTACAACAACTAGTAGCAATTCTCCAACGGCGGGTAGTCCTACGGTTAGGGTTTGCTCTGATTGCAACACCACCAAGACTCCTCTCTGGCGTA
GCGGACCCCAAGGCCCTAAGGATAAGATTGATATGATGCACTGTTGGGCGAGTGTGATGAAGGAGTGTGGTAGGAAATCATCGGTACTGGCAAATTGGGC
CAGGAGGAAGAGGAGGAGAGTCACGAGATTGGATGTAGCATCTTAG
AA sequence
>Lus10040556 pacid=23157655 polypeptide=Lus10040556 locus=Lus10040556.g ID=Lus10040556.BGIv1.0 annot-version=v1.0
MMNSNYQYSSHQFPPFSIDLNQDHQHYLNAHYFTTAITTATTTTSSSPSSSSSSPLPNYHIFINPPPPATQYSGEDTDPGGHLDYSSWGLPPPAAAAAAA
SFHQYHQPPQILLQKGGSSCMYDQEMVEVVDYNHNGSLDKMEKEMSYDVVSPKLRIMNQKKKKLTIKTNVKKPNNLNDVQSPSTLQLQPAAMTDNTTNTT
TMTTTTSSNSPTAGSPTVRVCSDCNTTKTPLWRSGPQGPKDKIDMMHCWASVMKECGRKSSVLANWARRKRRRVTRLDVAS
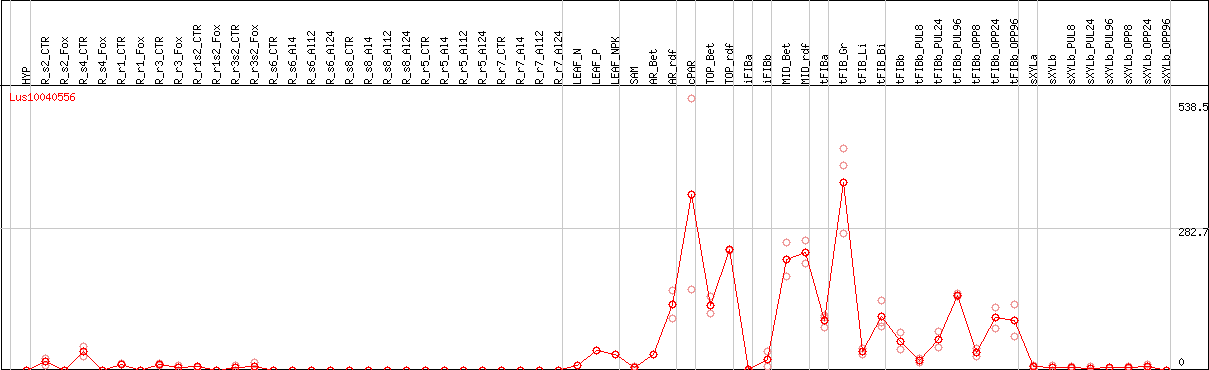
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040556 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.