External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G09270 63 / 7e-13
ATGSTU8
glutathione S-transferase TAU 8 (.1)
AT2G29420 61 / 5e-12
GST25, ATGSTU7
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
AT1G10360 56 / 2e-10
GST29, ATGSTU18
GLUTATHIONE S-TRANSFERASE 29, glutathione S-transferase TAU 18 (.1)
AT2G29490 55 / 6e-10
GST19, ATGSTU1
GLUTATHIONE S-TRANSFERASE 19, glutathione S-transferase TAU 1 (.1)
AT1G53680 54 / 2e-09
ATGSTU28
glutathione S-transferase TAU 28 (.1)
AT1G78370 52 / 9e-09
ATGSTU20
glutathione S-transferase TAU 20 (.1)
AT2G29470 52 / 1e-08
GST21, ATGSTU3
GLUTATHIONE S-TRANSFERASE 21, glutathione S-transferase tau 3 (.1)
AT1G78340 50 / 6e-08
ATGSTU22
glutathione S-transferase TAU 22 (.1)
AT2G29440 49 / 7e-08
GST24, ATGSTU6
GLUTATHIONE S-TRANSFERASE 24, glutathione S-transferase tau 6 (.1)
AT2G29460 48 / 2e-07
GST22, ATGSTU4
GLUTATHIONE S-TRANSFERASE 22, glutathione S-transferase tau 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016471
150 / 1e-46
AT2G29420 234 / 8e-78
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10040761
128 / 6e-38
AT2G29420 233 / 1e-77
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10016469
120 / 8e-35
AT2G29420 230 / 2e-76
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10040764
114 / 2e-32
AT2G29420 229 / 6e-76
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10016470
99 / 1e-26
AT2G29420 166 / 6e-52
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10016467
77 / 3e-18
AT2G29420 207 / 2e-67
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Lus10021103
64 / 2e-13
AT3G09270 250 / 1e-84
glutathione S-transferase TAU 8 (.1)
Lus10017208
64 / 4e-13
AT3G09270 212 / 3e-69
glutathione S-transferase TAU 8 (.1)
Lus10040727
62 / 2e-12
AT2G29420 216 / 6e-71
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G024200
63 / 9e-13
AT2G29420 188 / 4e-60
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Potri.010G070900
61 / 3e-12
AT2G29420 221 / 9e-73
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Potri.013G072400
61 / 4e-12
AT3G09270 161 / 2e-49
glutathione S-transferase TAU 8 (.1)
Potri.016G023340
56 / 3e-10
AT2G29420 194 / 3e-62
GLUTATHIONE S-TRANSFERASE 25, glutathione S-transferase tau 7 (.1)
Potri.016G023200
55 / 6e-10
AT3G09270 189 / 1e-60
glutathione S-transferase TAU 8 (.1)
Potri.010G061401
53 / 1e-09
AT3G09270 113 / 7e-32
glutathione S-transferase TAU 8 (.1)
Potri.010G061200
52 / 5e-09
AT3G09270 202 / 8e-66
glutathione S-transferase TAU 8 (.1)
Potri.011G140800
52 / 6e-09
AT1G78380 318 / 1e-111
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.001G436600
52 / 9e-09
AT1G78380 318 / 2e-111
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
Potri.001G431700
51 / 1e-08
AT1G78380 307 / 4e-107
GLUTATHIONE TRANSFERASE 8, A. THALIANA GLUTATHIONE S-TRANSFERASE TAU 19, glutathione S-transferase TAU 19 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0497
GST_C
PF00043
GST_C
Glutathione S-transferase, C-terminal domain
Representative CDS sequence
>Lus10040763 pacid=23157925 polypeptide=Lus10040763 locus=Lus10040763.g ID=Lus10040763.BGIv1.0 annot-version=v1.0
ATGGGATGCTTTTTGCAGGTGTTGCTAACGGCGTTTAAGGCTTACATAGGGAAGGGGGACAAAGAACAAGAGGACGCGATCGAGGAATTCACACAGCAAG
TGAAGTATCTGGAGGGGGAGCTGAACGGGAAGGAGTACTTCGGAGGGGAAAGGATCGGGTTTGTGGACATTGCAGTGTTCTCCATCTTGTACTGGCTTGG
AGCGTTGGAGGAAGTTACGGAAGCAGATCTGTTGACCAAGGAGAAGTTCCCTGTATTGTTCAAATTGATGGAGAAGCTCCAAGATGATAACGATGTTGTG
ATGGGGATGCCTGGCCTGCGAGGGACAAGCATCTGGATTACGTCCGAGCTGCTCGTGCTTTTCTATTGGAAGTTGTTGCTTCCAAAACAAATAGCTTAA
AA sequence
>Lus10040763 pacid=23157925 polypeptide=Lus10040763 locus=Lus10040763.g ID=Lus10040763.BGIv1.0 annot-version=v1.0
MGCFLQVLLTAFKAYIGKGDKEQEDAIEEFTQQVKYLEGELNGKEYFGGERIGFVDIAVFSILYWLGALEEVTEADLLTKEKFPVLFKLMEKLQDDNDVV
MGMPGLRGTSIWITSELLVLFYWKLLLPKQIA
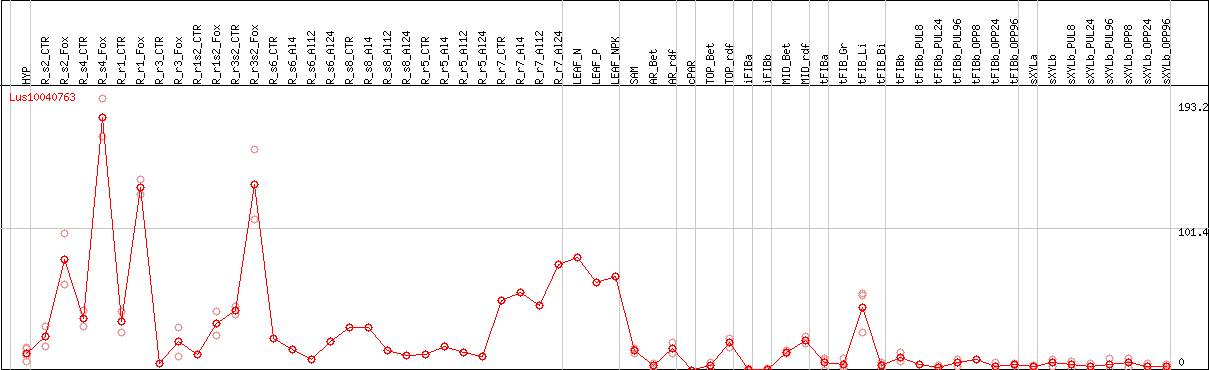
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040763 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.