External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G24330 221 / 5e-73
SDG34, ATXR6
SET DOMAIN PROTEIN 34, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 6 (.1)
AT5G09790 199 / 2e-64
PDE336, SDG15, ATXR5
SETDOMAIN GROUP 15, PIGMENT DEFECTIVE 336, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 5 (.1.2)
AT4G30860 40 / 0.0001
SDG4, ASHR3
ASH1-related 3, SET domain group 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016526
269 / 1e-92
AT5G24330 388 / 4e-136
SET DOMAIN PROTEIN 34, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 6 (.1)
Lus10007782
241 / 1e-81
AT5G24330 386 / 1e-135
SET DOMAIN PROTEIN 34, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 6 (.1)
Lus10038740
45 / 3e-06
AT2G44150 399 / 5e-137
SET DOMAIN-CONTAINING PROTEIN 7, histone-lysine N-methyltransferase ASHH3 (.1)
Lus10010121
40 / 0.0002
AT3G61740 1008 / 0.0
SET domain protein 14 (.1.2)
Lus10012626
40 / 0.0002
AT3G61740 1077 / 0.0
SET domain protein 14 (.1.2)
Lus10030262
39 / 0.0005
AT3G61740 742 / 0.0
SET domain protein 14 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G018300
230 / 1e-76
AT5G24330 480 / 3e-171
SET DOMAIN PROTEIN 34, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 6 (.1)
Potri.015G009632
229 / 3e-76
AT5G24330 483 / 1e-172
SET DOMAIN PROTEIN 34, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 6 (.1)
Potri.005G058000
207 / 5e-67
AT5G09790 481 / 2e-170
SETDOMAIN GROUP 15, PIGMENT DEFECTIVE 336, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 5 (.1.2)
Potri.007G109900
205 / 2e-66
AT5G09790 473 / 1e-167
SETDOMAIN GROUP 15, PIGMENT DEFECTIVE 336, ARABIDOPSIS TRITHORAX-RELATED PROTEIN 5 (.1.2)
Potri.007G147200
39 / 0.0005
AT2G44150 477 / 2e-169
SET DOMAIN-CONTAINING PROTEIN 7, histone-lysine N-methyltransferase ASHH3 (.1)
Potri.005G182100
38 / 0.0008
AT1G77300 807 / 0.0
LAZARUS 2, CAROTENOID CHLOROPLAST REGULATORY1, ASH1 HOMOLOG 2, histone methyltransferases(H3-K4 specific);histone methyltransferases(H3-K36 specific) (.1), histone methyltransferases(H3-K4 specific);histone methyltransferases(H3-K36 specific) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00856
SET
SET domain
Representative CDS sequence
>Lus10040794 pacid=23157903 polypeptide=Lus10040794 locus=Lus10040794.g ID=Lus10040794.BGIv1.0 annot-version=v1.0
ATGGTCGTGTTTGATCCACTAGAAGGGTTCACAGTAGAGGCAGACAGATCGATCAAGGACTGGACAATCATCACCGAGTATGTCGGAGACGTAGATTACT
TGAGGAACAGGGAAAACGACAATGGAGATAGCACAATGACGCTGCTTTCGGCTTCCGATCCTGGCAAGAGCCTGGTGGTGTGTCCTTATAGGAATGGGAA
TATAGCAAGATTCATCAACGGCATCAACAACCATACGCAGGAAGGTAGGAGGAAGCAGAACCTGAAATGTGTGAGGTTTGCTGTTGATGATGAGTGCCGG
GTGTTGCTGATTGCTAATAGGGATATAGCTAATGGGGAGAGGTTGTATTATGATTATAACGGTGATGAGCATGAGTACCCTACTGAGCATTTTGTTTGA
AA sequence
>Lus10040794 pacid=23157903 polypeptide=Lus10040794 locus=Lus10040794.g ID=Lus10040794.BGIv1.0 annot-version=v1.0
MVVFDPLEGFTVEADRSIKDWTIITEYVGDVDYLRNRENDNGDSTMTLLSASDPGKSLVVCPYRNGNIARFINGINNHTQEGRRKQNLKCVRFAVDDECR
VLLIANRDIANGERLYYDYNGDEHEYPTEHFV
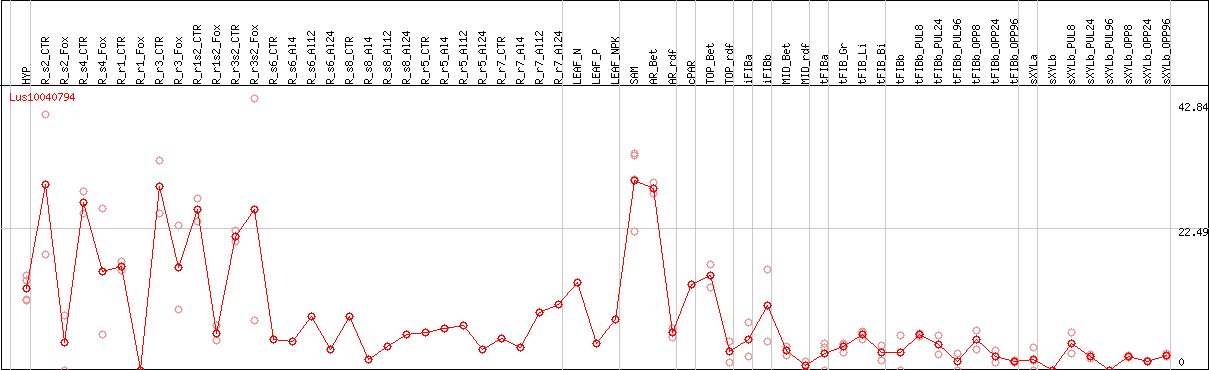
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040794 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.