External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G07900 129 / 2e-34
Mitochondrial transcription termination factor family protein (.1)
AT1G21150 111 / 5e-28
Mitochondrial transcription termination factor family protein (.1)
AT1G61970 96 / 4e-22
Mitochondrial transcription termination factor family protein (.1.2)
AT1G61980 90 / 4e-20
Mitochondrial transcription termination factor family protein (.1)
AT1G62110 90 / 5e-20
Mitochondrial transcription termination factor family protein (.1)
AT1G61960 89 / 1e-19
Mitochondrial transcription termination factor family protein (.1)
AT3G46950 81 / 3e-17
Mitochondrial transcription termination factor family protein (.1)
AT1G62120 81 / 6e-17
Mitochondrial transcription termination factor family protein (.1)
AT5G23930 78 / 6e-16
Mitochondrial transcription termination factor family protein (.1)
AT1G62085 77 / 7e-16
Mitochondrial transcription termination factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016551
485 / 2e-175
AT5G07900 144 / 3e-40
Mitochondrial transcription termination factor family protein (.1)
Lus10040462
283 / 4e-97
AT5G07900 93 / 1e-22
Mitochondrial transcription termination factor family protein (.1)
Lus10040817
278 / 4e-92
AT5G07900 225 / 6e-70
Mitochondrial transcription termination factor family protein (.1)
Lus10016550
269 / 1e-88
AT5G07900 219 / 9e-68
Mitochondrial transcription termination factor family protein (.1)
Lus10004957
255 / 4e-83
AT5G07900 238 / 4e-75
Mitochondrial transcription termination factor family protein (.1)
Lus10040819
228 / 7e-74
AT5G07900 130 / 2e-35
Mitochondrial transcription termination factor family protein (.1)
Lus10040820
221 / 7e-70
AT5G07900 221 / 2e-68
Mitochondrial transcription termination factor family protein (.1)
Lus10002883
155 / 2e-44
AT5G07900 196 / 9e-59
Mitochondrial transcription termination factor family protein (.1)
Lus10016553
105 / 9e-27
AT5G07900 171 / 7e-51
Mitochondrial transcription termination factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G022700
143 / 4e-40
AT5G07900 220 / 5e-68
Mitochondrial transcription termination factor family protein (.1)
Potri.015G038500
141 / 4e-39
AT5G07900 218 / 2e-67
Mitochondrial transcription termination factor family protein (.1)
Potri.012G046700
140 / 6e-39
AT5G07900 241 / 5e-76
Mitochondrial transcription termination factor family protein (.1)
Potri.008G216200
140 / 9e-39
AT5G07900 217 / 8e-67
Mitochondrial transcription termination factor family protein (.1)
Potri.004G012400
139 / 3e-38
AT5G07900 212 / 9e-65
Mitochondrial transcription termination factor family protein (.1)
Potri.014G133200
135 / 1e-36
AT5G07900 436 / 3e-152
Mitochondrial transcription termination factor family protein (.1)
Potri.015G038400
131 / 2e-35
AT5G07900 237 / 1e-74
Mitochondrial transcription termination factor family protein (.1)
Potri.004G012900
130 / 3e-35
AT5G07900 220 / 6e-68
Mitochondrial transcription termination factor family protein (.1)
Potri.012G046800
129 / 1e-34
AT5G07900 228 / 4e-71
Mitochondrial transcription termination factor family protein (.1)
Potri.004G013100
127 / 7e-34
AT5G07900 198 / 1e-59
Mitochondrial transcription termination factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02536
mTERF
mTERF
Representative CDS sequence
>Lus10040818 pacid=23157689 polypeptide=Lus10040818 locus=Lus10040818.g ID=Lus10040818.BGIv1.0 annot-version=v1.0
ATGGCGGCTCCCAAAGCAACAGCTTACCTCACTCCGATCAAGCAAAATACTCCTTCATCGTCGCCTGAGCTTTTTGAAGCCTTGAAGAATCGGCACCTCC
ATAGCGATAGATTACTACGTTTCCAAGTTCGAAACCATCCGCGACAGTTCTCGAACGAATCCGACGGCAATCAAACCACCACCTTCGGCCCGATTGTTTT
ATCTTACCTCATCAATGAATGCGGAATCCGTCCCGAAAAGGCAGCATCCGCCGTCAAATACGCCCAATTCAAAACCCCAGAAAAGGCCAATTCGGTCTTG
CATTTCTTCACAACCCTCGGCTTCTCAGCCGCCGACATCTCCAAAATCATCGGGGGGCGTCCTCGTCTGCTACTTTCCGATCCGGACAAACAATTCGCTC
CCCGGATTAGTTTCCTGGAATCGCAAGGGTTTTCTCCCTCCGACAGTGCTCAAGTCTTACTGAGAGGTCCTGAGATATTCTATTCCAGCGTAGCCAAACA
GCTGCTACCCAATTTTGAATTCATCAAGAGCGTCGTCCTTTCCAACGAGAAGGCTGTTCAAGCCATTAGGCGAAGTCCTTTGTTTTTGACTTGTAAGCTG
GAGACGTATTTGATCACGAGCATCAAGACTCTGAAGGAAAACGGGATTAGCGACTCGTTTATTGTACGGTTGCTTGTCTCTAGCCCCTTATTAACTATGT
TGACACCGCAGGCTAAATTATCCGAGGCTGTTGAGCGTGTGAAGGAAAATGGGATTGATCCTAACTCGGCAACGTTTGCTGTTGCTATGGAGACGATCAA
AGCGACTAAAGTGACGTGGGATGGTAAGGTTGATGCTTATAAGAAGTGGGGGGGTGGTCTGATGAAGTGA
AA sequence
>Lus10040818 pacid=23157689 polypeptide=Lus10040818 locus=Lus10040818.g ID=Lus10040818.BGIv1.0 annot-version=v1.0
MAAPKATAYLTPIKQNTPSSSPELFEALKNRHLHSDRLLRFQVRNHPRQFSNESDGNQTTTFGPIVLSYLINECGIRPEKAASAVKYAQFKTPEKANSVL
HFFTTLGFSAADISKIIGGRPRLLLSDPDKQFAPRISFLESQGFSPSDSAQVLLRGPEIFYSSVAKQLLPNFEFIKSVVLSNEKAVQAIRRSPLFLTCKL
ETYLITSIKTLKENGISDSFIVRLLVSSPLLTMLTPQAKLSEAVERVKENGIDPNSATFAVAMETIKATKVTWDGKVDAYKKWGGGLMK
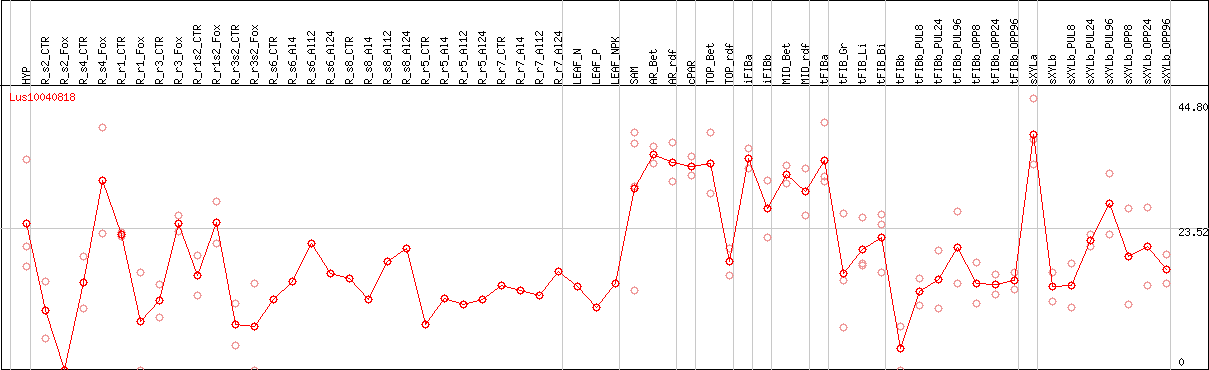
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10040818 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.