Lus10041008 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10041008 pacid=23179811 polypeptide=Lus10041008 locus=Lus10041008.g ID=Lus10041008.BGIv1.0 annot-version=v1.0
ATGAATGCCATCGCTGGGAGGGGACTTTCACCTCTTGGCCATGCTCTTAATAACTTCTCTGAGGTAAGGGATCGTGATCTGGTTGAAGCAAGGGGCATTC
CCTGTCTGATTGCAATGGCTGCAAAGCAGCCTATAGATATAAGTTCTGATAGTGATCTGGGGCTGGATGATGATGATGACGACTTGGACATTAGTGAATC
TCTAAGACAGGGTTCTAATAATAAGCGGATTCTTCCTCAGTGGGAGACAGGCACAAGTAGTGCAGGGGCCGTCAGACAAAATGGTAATGCTGGTAGCCTG
AGATCTGGTAACTCTAGGATTGCTAATGTTTCTGGTGGCGAGCAAGCTTTGAAGCGGACCCTTCCAATGTCTCTGGATGGAGCTGCACCAAATTACAAGA
TGAACAGGTTAGCTGATAATGCAGCTAGCAGCCAGACACATGGTGTTCATCGCAATGTCTACAATCCCTCAGTAGCAGGGAGGAGCCAGGAATACACTCA
CGGCCCCTACAGCCATGTTGGTAATGAAGACGTTATGATGTATGAAAAAAATGGAGGCAGGGCTCTTCCTCAGTCATGGACGCATGTTAAGTCTACCTTG
CCTGTGCATTTGGCTGGTTCAAGTGACCATGGATACGCTTCCATGGTAGGTGATGAAAGAACTGCTGAATGTGATGAGAGGTTAATCTATCAAGCAGCAC
TACAGGGTATTAATCAACCAAAGATGGAAGCCAACTTACCTGAAGGTTTATTATCTGTGCCTCTTCTGAAACACCAGAAAATTGCATTGGCGTGGATGCT
TCAGAAGGAGATGAGAAGCTTACACTGTTTAGGTGGAATTTTAGCTGATGATCAGGGTCTTGGGAAAACTATATCAATGATTGCCCTTGTACAGATGCAG
AAGTATTTGGAGTTGAAATCTAAGTCAGAAGATCAAAATGCAAGAAATACTGAAGCACTGAATTTGGATGACGATGATGATTGCAGCACTGTTGCCTCAG
ATAAAATGAAACAGTCAGCTGAAATTAAGGATACCGTAATAACTGAACAAGTAAATACTTCACAGGCACTGATTAGAAGGAGACCTGCTGCTGGTACATT
GGTTGTGTGCCCAGCAAGTGTTCTCCGCCAGTGGGCTAGGGAGTTGGATGATAAAGTTGCGGAAAAGTCTAAACTGTCCGTCTTGATATACCACGGAGGT
ACGAGAACAAAAGATCCTGTTGAATTGGCAAAGTTTGATGTCGTTATGACGACATACGCCATAGTAACTAATGAACTTCCCAAGCAACCTTTGGTTGATG
ATGATGAACTTGATGACAAAACTGCAGAGAAAAGTGCTCTGTCTTCTGCATTTTCAGTTGATAAGAAGGGGAAAAAGCTGCTTGGTACCAGCAAGAAGAG
GAAGAAGGGCACAAAGGGATCAGATTTCTCTTCTCTTGATTATGATTGTAGTCCACTTGCAAGGGTGGGTTGGTGTAGGGTGATTCTAGATGAAGCTCAA
ACAATAAAAAATCATCGAACCCAAGTGGCCAGGGCTTGCTGTAGCCTTCGTGCTAAGAGGAGATGGTGCCTATCTGGAACACCTATACAAAATTCTATTG
ATGATTTGTACAGCTATTTCAGATTTCTCAGATATGAACCATACTCGGGGTACAAATCATTTTGCAGTGCCATAAAGCTCCCAATATCTAGAAATTCACT
TCATGGCTACAAGAAGCTGAGAGCAGTTTTAGCAGCAGTAATGCTGCGCCGTACTAAAGGGACATTGATAAATGGGGAGCCTATAATCAGATTGCCTCCA
AAATCAATATGTTTGAGGAAGGTGGATTTCTCGATCGAGGAGCGGGCTTTCTACACCCAGCTAGAAGCTGATTCGCGTTCTAAGTTCAAGGCTTATGCTG
CTGCTGGCACAGTAAATCAAAACTATGCAAATATTCTTTTGATGCTTTTGCGCCTTCGGCAGGCTTGTGACCACCCAGTTCTTGTCAAAGGTTTTAATTC
CAACTCTGTACAAAAAGTATCAGTGGATATGGCAAATAAACTTCCAAAGGAGATGGTGATGGATCTCTTAAGTAGACTGACTACTTCTCTGACTATTTGC
CGTGTATGCACTGATCTGCCCGAGGACCCTGTTGTTACAATGTGCGGTCATGTTTTCTGTTATCAGTGTGTGTCAAATTACTTGGCCGGGGATGATAACA
CCTGCCCTGGTCCTGATTGCAAAGAACAGCTTGGTGCTGATATTGTCTTCTCGGAAGCTACGTTGAGAAGTTGCATGTCTAACAATGGTGATGCAGGTCG
TTCTCGATCCAACTTTGATGAGATAATGGAGAGCGAGTATAGCTCTTCAAAAATCAGAGCTATCCTTGAGATTCTTTTCTCACATAGTGAAATGAGTGAA
CCTGATGTTGTAGAGTGCACAGTTGTGAGCTCTGTTCCTAAAGGACCTATAAAAACGATAATTTTTTCACAATGGACTGGCATGTTGGACTTGGTTGAGC
GCTCGCTGAAAGAGACTGGCCTAGCGTATAGGAGACTTGATGGCACAATGAGTTTAAATGTGAGAGACAAGGCTGTCAAAGATTTCAATACAAATCCCGA
GGTAACGGTAATGCTGATGTCACTCAAGGCAGGAAATCTTGGTTTGAATATGGTTGCTGCATGTCACGTTATCCTTCTGGATCTTTGGTGGAATCCAAGC
ACAGAAGATCAGGCAGTTGACAGAGCACATAGAATCGGACAGACTCGGCCAGTTACTGTAACCCGGCTCACCATTGCAGACACTGTGGAGGATAGGATAC
TAGCTTTGCAGGATGAGAAGAGGAAAATGGTTTCGTCAGCTTTTGGTGAAGACCAAAGTGGGGGGACAGCTACACGGCTTACAGTGGAAGATCTAAGATT
TCTGTTCATGGACAGGGATGGGACCCGCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10041008 pacid=23179811 polypeptide=Lus10041008 locus=Lus10041008.g ID=Lus10041008.BGIv1.0 annot-version=v1.0
MNAIAGRGLSPLGHALNNFSEVRDRDLVEARGIPCLIAMAAKQPIDISSDSDLGLDDDDDDLDISESLRQGSNNKRILPQWETGTSSAGAVRQNGNAGSL
RSGNSRIANVSGGEQALKRTLPMSLDGAAPNYKMNRLADNAASSQTHGVHRNVYNPSVAGRSQEYTHGPYSHVGNEDVMMYEKNGGRALPQSWTHVKSTL
PVHLAGSSDHGYASMVGDERTAECDERLIYQAALQGINQPKMEANLPEGLLSVPLLKHQKIALAWMLQKEMRSLHCLGGILADDQGLGKTISMIALVQMQ
KYLELKSKSEDQNARNTEALNLDDDDDCSTVASDKMKQSAEIKDTVITEQVNTSQALIRRRPAAGTLVVCPASVLRQWARELDDKVAEKSKLSVLIYHGG
TRTKDPVELAKFDVVMTTYAIVTNELPKQPLVDDDELDDKTAEKSALSSAFSVDKKGKKLLGTSKKRKKGTKGSDFSSLDYDCSPLARVGWCRVILDEAQ
TIKNHRTQVARACCSLRAKRRWCLSGTPIQNSIDDLYSYFRFLRYEPYSGYKSFCSAIKLPISRNSLHGYKKLRAVLAAVMLRRTKGTLINGEPIIRLPP
KSICLRKVDFSIEERAFYTQLEADSRSKFKAYAAAGTVNQNYANILLMLLRLRQACDHPVLVKGFNSNSVQKVSVDMANKLPKEMVMDLLSRLTTSLTIC
RVCTDLPEDPVVTMCGHVFCYQCVSNYLAGDDNTCPGPDCKEQLGADIVFSEATLRSCMSNNGDAGRSRSNFDEIMESEYSSSKIRAILEILFSHSEMSE
PDVVECTVVSSVPKGPIKTIIFSQWTGMLDLVERSLKETGLAYRRLDGTMSLNVRDKAVKDFNTNPEVTVMLMSLKAGNLGLNMVAACHVILLDLWWNPS
TEDQAVDRAHRIGQTRPVTVTRLTIADTVEDRILALQDEKRKMVSSAFGEDQSGGTATRLTVEDLRFLFMDRDGTR
|
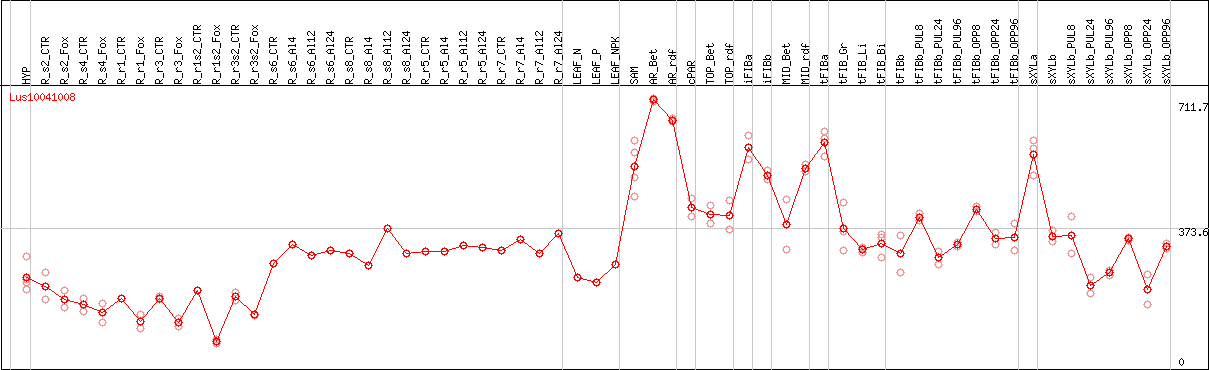
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041008 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
