External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G50370 237 / 4e-80
AtFYPP1
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
AT3G19980 235 / 2e-79
STPP, ATFYPP3, EMB2736
SERINE/THREONINE PROTEIN PHOSPHATASE, EMBRYO DEFECTIVE 2736, "flower-specific, phytochrome-associated protein phosphatase 3", flower-specific, phytochrome-associated protein phosphatase 3 (.1)
AT4G26720 137 / 7e-41
PPX-1, EP129, EP124, PPX1
PROTEIN PHOSPHATASE X-1, protein phosphatase X 1 (.1)
AT5G55260 134 / 1e-39
EP128, PPX-2, PPX2
PROTEIN PHOSPHATASE X -2, protein phosphatase X 2 (.1)
AT1G10430 117 / 3e-33
PP2A-2
protein phosphatase 2A-2 (.1)
AT1G69960 115 / 1e-32
PP2A
serine/threonine protein phosphatase 2A (.1)
AT1G59830 112 / 2e-31
PP2A-1
protein phosphatase 2A-2 (.1.2)
AT2G42500 106 / 2e-29
PP2A-3, PP2A-4
protein phosphatase 2A-3 (.1.2.3)
AT3G58500 107 / 3e-29
PP2A-4, EP7, PP2A-3
protein phosphatase 2A-4 (.1)
AT2G39840 87 / 2e-21
TOPP4
type one serine/threonine protein phosphatase 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013456
240 / 2e-81
AT3G19980 623 / 0.0
SERINE/THREONINE PROTEIN PHOSPHATASE, EMBRYO DEFECTIVE 2736, "flower-specific, phytochrome-associated protein phosphatase 3", flower-specific, phytochrome-associated protein phosphatase 3 (.1)
Lus10041015
240 / 2e-81
AT1G50370 570 / 0.0
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Lus10010156
237 / 4e-80
AT1G50370 617 / 0.0
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Lus10017361
237 / 4e-80
AT1G50370 617 / 0.0
flower- specific, phytochrome-associated protein phosphatase 1, Calcineurin-like metallo-phosphoesterase superfamily protein (.1)
Lus10016576
139 / 7e-41
AT5G55260 615 / 0.0
PROTEIN PHOSPHATASE X -2, protein phosphatase X 2 (.1)
Lus10013095
132 / 3e-39
AT5G55260 388 / 4e-137
PROTEIN PHOSPHATASE X -2, protein phosphatase X 2 (.1)
Lus10030810
118 / 2e-33
AT1G10430 617 / 0.0
protein phosphatase 2A-2 (.1)
Lus10013287
118 / 2e-33
AT1G10430 617 / 0.0
protein phosphatase 2A-2 (.1)
Lus10033122
118 / 2e-33
AT1G10430 606 / 0.0
protein phosphatase 2A-2 (.1)
PFAM info
Representative CDS sequence
>Lus10041012 pacid=23179521 polypeptide=Lus10041012 locus=Lus10041012.g ID=Lus10041012.BGIv1.0 annot-version=v1.0
ATGTGGAGTGATCCTGAAGACATAGAAACATGGGCAGTTAGTCCCAGAGGAGCAGGTTGGCTTTTTGGATCAAGGGTTACCTCTGAGTATAACCACATTA
ACAATCTTGACTTGGTTTGTCGGGCGCATCAACTTGTACAAGAAGGCCTCAAGTACATGTTTCAAGATAAGGGCTTAGTAACCGTCTGGTCTGCACCGAA
CTACTGTTACCGATGTGGGAATGTAGCTTCCATCCTGAGCTTCAATGAGAATATGGAGAGAGAAGTAAAGTTCTTCACCGAAACGGAGGAGAACAATCAG
ATGAGAGGACCAAGAACAGGAGTGCCCTATTTCCTATGA
AA sequence
>Lus10041012 pacid=23179521 polypeptide=Lus10041012 locus=Lus10041012.g ID=Lus10041012.BGIv1.0 annot-version=v1.0
MWSDPEDIETWAVSPRGAGWLFGSRVTSEYNHINNLDLVCRAHQLVQEGLKYMFQDKGLVTVWSAPNYCYRCGNVASILSFNENMEREVKFFTETEENNQ
MRGPRTGVPYFL
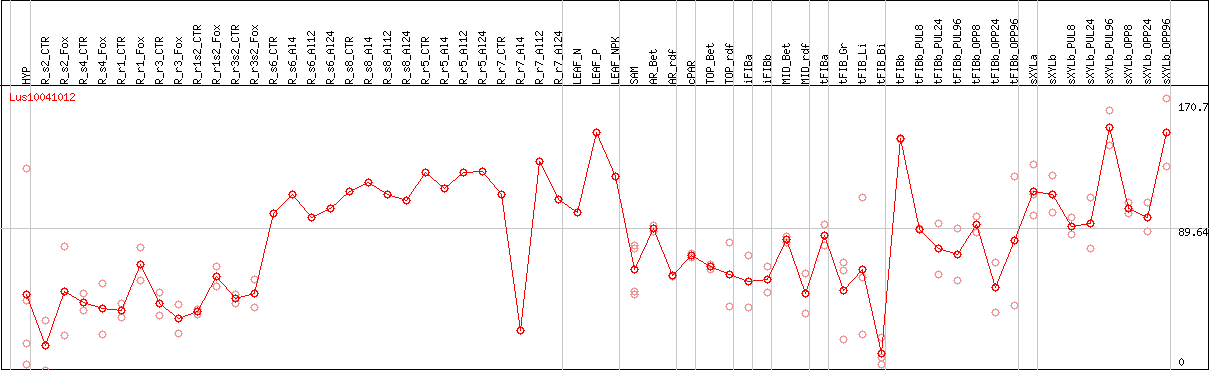
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041012 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.