Lus10041019 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10041019 pacid=23179660 polypeptide=Lus10041019 locus=Lus10041019.g ID=Lus10041019.BGIv1.0 annot-version=v1.0
ATGAGTAGCTTGGTTGAGAGGCTTAGAGTCCGATCTGATCGAAGGCCGATATACAACCTTGATGAATCAGATGATGAAGATTTTCTGTCCCGAAAAGCTG
GGAAATCAGCAGAAAATGCAGAGAGAATAGTCCGACCTGATGCGAAAGATAACCTATGTCAAGCATGTGGAGAAAGTGGCGAGCTTGTGAACTGTGAAAC
CTGCACTTATGCCTATCATGCGAAGTGTCTGCTTCCACCCCTAAAAGCTCCTCCTTCCAGCAGTTGGAGCTGTCCTGAATGTGTTAGTCCTCTAAATGAA
ATTGAGAAGATATTGGACTGTGAAATGCGTCCAACGGCAGCTGATGATAGTGATGCATCTACGCTGGGCTCAAAGCAAATTTTCGTTAAGCAATATCTGG
TGAAGTGGAAGCGGTTATCATATCTTCACTGCACTTGGGTTCCTGAGAAAGAGTTCGTGAAAGCATTCAAGTCAAACCCCCGTCTTAGGACTAAATTAAG
CAACTTCCATAGACAAGGTACCTCAACCAATTCTGAGGAGGAACCTGCAATCAGGCCAGAATGGACAACAGTTGATAGAATTCTTGCATGCAGGGGTGAG
GATGATGAAAAGGAATACCTTGTGAAGTATAAGGAACTTCCTTATGATGAATGTTACTGGGAATTTGAAACAGATATATCAGCATTTCAGCCAGAAATAG
AGAAGTTTAATATAATTCACTTAAAGTCCCACCGGTTAAATAAGCAAAAGAGTTTCCGTGATGCTCCTGATTCCAAGAAGAATTTGAAAGAATTCCAACA
GATTGAGCAAAGCCCTGAGTTTATCTCAGGAGGCTCATTGCATCCATATCAACTAGAAGGGGTAAACTTTCTGCGGGTTTTTTTTTTTAAGCAAACACAT
GTCATACTTGCCGATGAAATGGGTTTGGGGAAAACTATACAAAGTATTGCTTTTCTAGCATCTCTGTTCGAAGAGAACCTGTACCCCCATTTAGTGATTG
CCCCACTTTCAACATTGAGAAACTGGGAACGTGAATTTGCAACATGGGCACCTCAACTGAATGTCGTTATGTATGTTGGTTCTGCTCAAGGCCGTGCTGT
CATTAGGGAATATGAGTTTTACTACCCTAAAAATCACAAGAAGATGAAGAAAAAGAAATCTGGTCAACTTGTTAGTGAAAGCAAGCAAGATAGGATAAAG
TTTGATGTGCTCTTGACTTCCTATGAGATGGTTAACATGGATACAGCATCCCTAAAACCCATTAAATGGGAGTGCATGATTGTCGACGAAGGTCATCGCC
TGAAAAACAAGGATTCAAAACTTTTTATTTCTCTGAAGCAATATTCCAGTAATCACCGTGTGCTTTTGACGGGGACCCCTCTTCAGAATAATCTTGATGA
GCTATTCATGCTGATGCACTTCCTTGATGCTGGAAAGTTTTCAAGTTTGGAGGAGTTTCAAGAAGAGTTCAAGGATATAAATCAGGAGGAACAGATTTCT
CGGCTTCATAAAATGTTGGCTCCCCATCTCCTTAGAAGGGTGAAAAAGGATGTTATGACAGAGTTGCCTCCTAAAAAGGAACTCATTCTACGTGTTGATC
TAAGTAGCTTGCAAAAAGAATATTACAAGGCAATTCTAACTCGTAATTACCAAATTTTAACACGGCGTGGTGGTCCACAGATTTCACTTATCAACGTGGT
TATGGAATTGCGGAAGCTCTGCTGTCATCCTTATATGTTAGAAGGAGTTGAACCAGCAATAGACGATCCAACAGAATCTTTCAAGCAATTTGTGGAGTCA
TCTGGAAAGTTGCAACTACTTGACAAAATGATGGTGAAACTGAAAGAGCAAGGCCATAGAGTTCTTATTTATTCCCAGTTTCAGCACATGCTGGATTTAC
TTGAGGATTACTGCGTCTACAAGAAATGGCAATACGAACGAATTGATGGGAAAGTTGGTGGAGCTGAGAGACAAATTCGCATTGATCGGTTTAATTCAAA
AAATTCGTCCAGATTCTGTTTCCTTCTCTCTACTAGGGCTGGAGGACTGGGTATAAACCTTGCAACTGCTGATACAGTCATCATATACGACAGTGATTGG
AATCCGCATGCTGACCTCCAGGCTATGGCTAGGGCTCATCGACTTGGCCAAACTAACAAGGTCATGATTTATAGGCTTATAACACGAGGAAGTATTGAGG
AGCGCATGATGCAATTGACTAAGAAGAAAATGGTATTAGAACATCTTGTTGTTGGGAGGCTTAAGGCTCAAAATATAAACCAGGAAGAGTTGGATGACAT
TATCAGGTATGGCTCAAAGGAGCTATTTGCTGATGAGAATGATGAATTGGGGAAGTCTCGTCAAATTCATTATGATGACGGTGCAATTAACCGATTACTG
GACCGTGAACAAATTGGAGATGAAGAGGCAACATTGGACGATGAAGAAGAAAATGGATTTCTAAAGGCATTTAAGGTTGCAAACTTTGAGTATATTGATG
AAAAGGCGGAGGAAGAAGCTCCAAAACCTGCGTTGGAGACGATGAGCAATTCAGAGAAATTAAATTTCTGGGAGGAGCTCTTAAGAGATAAATTTGAAGA
GCAAAAGGGCGAGGAATCGAAATCTCTTGGAAAGGGAAAAAGAAGCCGCAAACAGATGGTGTCTGTTGAAGATGATGACCTTGCTGGTATGGAGGATTTA
AGCTCTGATGGAGAGGATGACAATTATGAAGCAGACTTAACCGATGGAGAAACTTCATCGGGAACTCAGGCTGGGAGAAAACCTTACAGAAAGAGAGCTC
GGGACAATTCTGAACCCATTCCTTTAATGGAAGGAGAAGGAAAGTCCTTCAAAGTACTTGGTTTCAATCAGAACCAGAGGGCTCTTTTTGTTCAGATAAT
GATGAGGTTTGGAATTGGGGACGAAAACTGGAAACAGTTTGTACCTCGCATGAAACAGAAGACTCTCGAGGAAATCCATAATTATGGGAAGATGTTCTTC
AGACATGTTATTGAAGAAATGAACGACTTACCACACTTCTCAGACGGTGTTCCTAAAGAAGGACTTCGAATAGCAGACGTATTGGTCAGAATTGCATCGT
TGAATATAATGAGGCAAAAGGTGAAGCTCGTCTCAGAAAATCCAACCATGCCGCTTTTTGAGGAGGACGTTTATCCACGCTTGCCAGGACTGAAAGGTGG
GAAATTTTGGACCGAAGAACATGATAAAACTTTACTCAAGGCAGTGATTAAGCAAGGCTATGGCCGTTGGCAAGCTATTCTTGATGACAAGGACGTGAAA
ATTCAGGAAGTTGTATCCCAAGAGCTGAATCTCCCCGTTATGAATCTCCCTACCCCTGGGCAAATTTTGTCTCAGGCACAAAACGGTGCTAGTTCTGGGA
TTACTGAAGGTCCAAGCACCCAGCAGGCCTCAGTAAATGCTACCGGGAATGAGATTGGCGCCAATGCTGCCCAGGGTATACCTCAAGGGCAATCTTACCA
AGAATCCATCCTGTACCATCTTAGAGATATGCAGAGAAGAATGGTTGAATTTGTTAAGAAGAGGGTGCTTCTTCTTGAGCGGGGGATGAATGTTGAAGTC
CAGAAAGAATACTATGCCAATCTAGATACTAAGCCAACAGTGAAAGATCCTAGTAGTGAAAGAAAGGGCACAAATCACTCAGCTTCAGTTTCGTTAGAGA
CCGACTCTGAAAAGATTAATCAACTGCCGCCGATGGAAGAAATCACATCTGAAGATACTGTTGCTGCCGGTTGTGTTGCTGACCCAGATCGATTAGACTT
GGTCGAGCTTCACAATAAGATGTGCAAGAGTGTTGAGAACATTGCTCCTGATTTGCTTACGACATCCTCAAGGAGCCAAATAACAAGTTTTGAACTGAAA
CAGAAGCTTCAACCATTGGAGACCATCTGTGAGCGTATGAATACTGTGCTCTCACCTTCTGAGCAGCCATTGCAAGATGTGGAGCAAAAAGGAGGCGCTG
AGCCACCCAGAAGTCTAGATGAGTCGTGCTCTACATCACTCGAACATAGCGAACATGCTCTTCCTAACAGTGTAGCAAAGATGGAGTTTGAAATCGGCAG
TTTGTTTGGGATGCCAAGTGTGGGTTCAGTGACTGTGCCGGACACGATTGATGTGGAAATGGAAGATAAATGGAAGGATGGTGCCCCTGCCGCAGATGAT
GATTTAACTAACAATGGCGGGCACCCGGGATTCCCATACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10041019 pacid=23179660 polypeptide=Lus10041019 locus=Lus10041019.g ID=Lus10041019.BGIv1.0 annot-version=v1.0
MSSLVERLRVRSDRRPIYNLDESDDEDFLSRKAGKSAENAERIVRPDAKDNLCQACGESGELVNCETCTYAYHAKCLLPPLKAPPSSSWSCPECVSPLNE
IEKILDCEMRPTAADDSDASTLGSKQIFVKQYLVKWKRLSYLHCTWVPEKEFVKAFKSNPRLRTKLSNFHRQGTSTNSEEEPAIRPEWTTVDRILACRGE
DDEKEYLVKYKELPYDECYWEFETDISAFQPEIEKFNIIHLKSHRLNKQKSFRDAPDSKKNLKEFQQIEQSPEFISGGSLHPYQLEGVNFLRVFFFKQTH
VILADEMGLGKTIQSIAFLASLFEENLYPHLVIAPLSTLRNWEREFATWAPQLNVVMYVGSAQGRAVIREYEFYYPKNHKKMKKKKSGQLVSESKQDRIK
FDVLLTSYEMVNMDTASLKPIKWECMIVDEGHRLKNKDSKLFISLKQYSSNHRVLLTGTPLQNNLDELFMLMHFLDAGKFSSLEEFQEEFKDINQEEQIS
RLHKMLAPHLLRRVKKDVMTELPPKKELILRVDLSSLQKEYYKAILTRNYQILTRRGGPQISLINVVMELRKLCCHPYMLEGVEPAIDDPTESFKQFVES
SGKLQLLDKMMVKLKEQGHRVLIYSQFQHMLDLLEDYCVYKKWQYERIDGKVGGAERQIRIDRFNSKNSSRFCFLLSTRAGGLGINLATADTVIIYDSDW
NPHADLQAMARAHRLGQTNKVMIYRLITRGSIEERMMQLTKKKMVLEHLVVGRLKAQNINQEELDDIIRYGSKELFADENDELGKSRQIHYDDGAINRLL
DREQIGDEEATLDDEEENGFLKAFKVANFEYIDEKAEEEAPKPALETMSNSEKLNFWEELLRDKFEEQKGEESKSLGKGKRSRKQMVSVEDDDLAGMEDL
SSDGEDDNYEADLTDGETSSGTQAGRKPYRKRARDNSEPIPLMEGEGKSFKVLGFNQNQRALFVQIMMRFGIGDENWKQFVPRMKQKTLEEIHNYGKMFF
RHVIEEMNDLPHFSDGVPKEGLRIADVLVRIASLNIMRQKVKLVSENPTMPLFEEDVYPRLPGLKGGKFWTEEHDKTLLKAVIKQGYGRWQAILDDKDVK
IQEVVSQELNLPVMNLPTPGQILSQAQNGASSGITEGPSTQQASVNATGNEIGANAAQGIPQGQSYQESILYHLRDMQRRMVEFVKKRVLLLERGMNVEV
QKEYYANLDTKPTVKDPSSERKGTNHSASVSLETDSEKINQLPPMEEITSEDTVAAGCVADPDRLDLVELHNKMCKSVENIAPDLLTTSSRSQITSFELK
QKLQPLETICERMNTVLSPSEQPLQDVEQKGGAEPPRSLDESCSTSLEHSEHALPNSVAKMEFEIGSLFGMPSVGSVTVPDTIDVEMEDKWKDGAPAADD
DLTNNGGHPGFPY
|
DESeq2's median of ratios [FLAX]
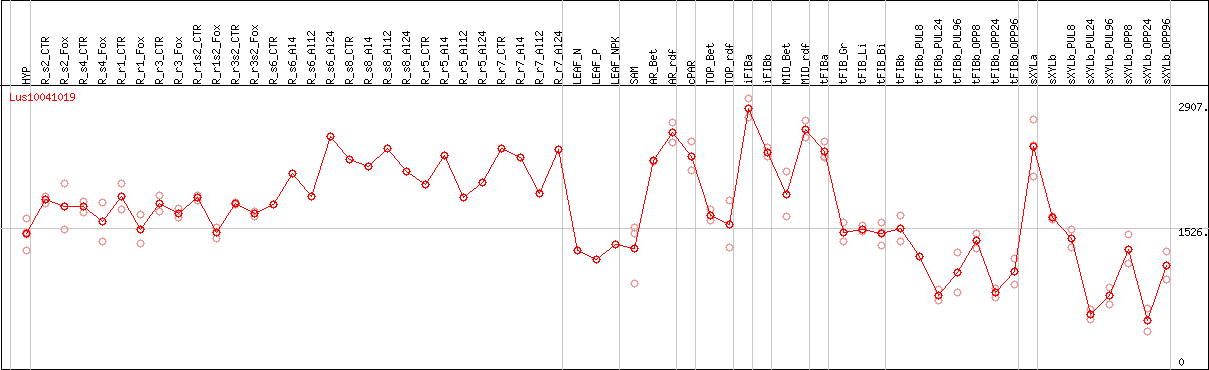
Coexpressed genes
Lus10041019 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
