External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G11100 105 / 1e-28
SYT4, NTMCTYPE2.2, ATSYTD ,NTMC2T2.2 ,NTMC2TYPE2.2
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G05500 95 / 8e-25
SYT5, NTMCTYPE2.1, ATSYTE ,NTMC2T2.1 ,NTMC2TYPE2.1
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006193
128 / 9e-37
AT5G11100 862 / 0.0
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10038117
98 / 8e-26
AT1G05500 888 / 0.0
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10008055
75 / 1e-17
AT1G05500 775 / 0.0
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G025000
119 / 1e-33
AT5G11100 824 / 0.0
synaptotagmin 4, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.006G063900
100 / 2e-26
AT1G05500 847 / 0.0
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.018G124000
99 / 3e-26
AT1G05500 877 / 0.0
synaptotagmin 5, ARABIDOPSIS THALIANA SYNAPTOTAGMIN HOMOLOG E, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
PFAM info
Representative CDS sequence
>Lus10041033 pacid=23179753 polypeptide=Lus10041033 locus=Lus10041033.g ID=Lus10041033.BGIv1.0 annot-version=v1.0
ATGTTGATGTTGGAGGTTTGGGATCATGACACTTTCGGAAAGGATAAAATTGGTAGAGTGATAATGACACTAACAAGAGTGATAATGGAAGGTGAGTTTC
AGGATTCATTTCCTCTGGATGGTGCCAAATCTGGGAAGCTTTTCTTGCATCTCAAGTGGACTCCCCAGCTTAAATTCCGTGACTCTTAA
AA sequence
>Lus10041033 pacid=23179753 polypeptide=Lus10041033 locus=Lus10041033.g ID=Lus10041033.BGIv1.0 annot-version=v1.0
MLMLEVWDHDTFGKDKIGRVIMTLTRVIMEGEFQDSFPLDGAKSGKLFLHLKWTPQLKFRDS
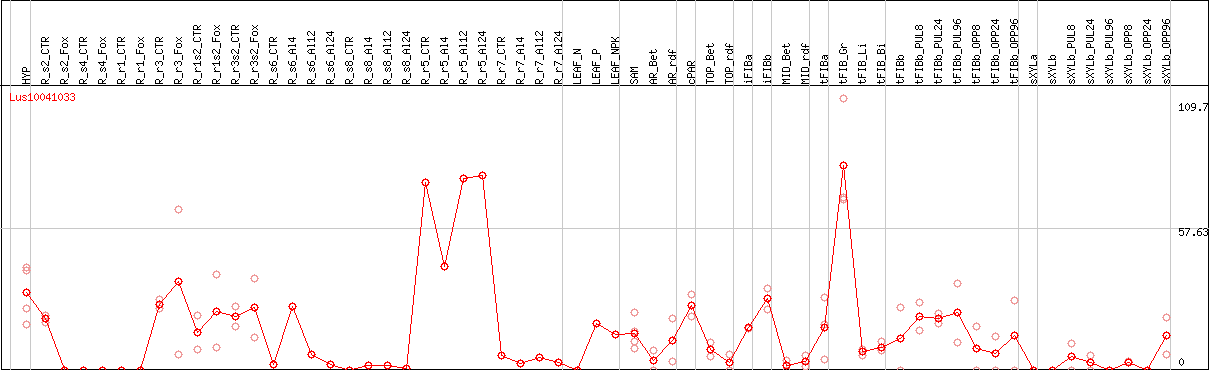
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041033 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.