External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G11730 302 / 3e-104
ATFP8, AtRABD1
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG D1, Ras-related small GTP-binding family protein (.1)
AT4G17530 278 / 9e-95
RAB1C, AtRab1C, AtRABD2c
RAB GTPase homolog 1C (.1)
AT1G02130 275 / 1e-93
ARA5, AtRABD2a, AtRab1B, Ara-5
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
AT5G47200 273 / 5e-93
AtRABD2b, AtRab1A
ARABIDOPSIS RAB GTPASE HOMOLOG D2B, RAB GTPase homolog 1A (.1)
AT3G46060 199 / 2e-63
ARA3, Ara-3, AtRABE1c, AtRab8A
RAB GTPase homolog 8A (.1.2.3)
AT5G59840 199 / 2e-63
Ras-related small GTP-binding family protein (.1)
AT3G53610 195 / 6e-62
ATRAB8, AtRab8B, AtRABE1a
RAB GTPase homolog 8 (.1.2.3)
AT3G09900 188 / 2e-59
AtRABE1e, AtRab8E
RAB GTPase homolog E1E (.1)
AT5G03520 185 / 6e-58
ATRAB-E1D, AtRab8C, AtRABE1d
ARABIDOPSIS RAB HOMOLOG E1D, RAB GTPase homolog 8C (.1.2)
AT3G07410 154 / 4e-46
AtRABA5b
RAB GTPase homolog A5B (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036442
356 / 3e-125
AT3G11730 350 / 2e-124
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG D1, Ras-related small GTP-binding family protein (.1)
Lus10010999
273 / 1e-92
AT5G47200 387 / 3e-139
ARABIDOPSIS RAB GTPASE HOMOLOG D2B, RAB GTPase homolog 1A (.1)
Lus10000595
271 / 6e-92
AT5G47200 387 / 3e-139
ARABIDOPSIS RAB GTPASE HOMOLOG D2B, RAB GTPase homolog 1A (.1)
Lus10037608
228 / 2e-75
AT1G02130 255 / 1e-87
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Lus10007698
196 / 1e-62
AT3G46060 378 / 1e-135
RAB GTPase homolog 8A (.1.2.3)
Lus10024423
196 / 2e-62
AT3G46060 376 / 7e-135
RAB GTPase homolog 8A (.1.2.3)
Lus10023430
196 / 4e-62
AT3G46060 364 / 3e-129
RAB GTPase homolog 8A (.1.2.3)
Lus10025309
195 / 5e-62
AT3G46060 374 / 1e-133
RAB GTPase homolog 8A (.1.2.3)
Lus10005890
194 / 9e-62
AT3G46060 396 / 2e-142
RAB GTPase homolog 8A (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G226600
345 / 3e-121
AT3G11730 364 / 4e-130
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG D1, Ras-related small GTP-binding family protein (.1)
Potri.003G004000
337 / 3e-118
AT3G11730 359 / 2e-128
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG D1, Ras-related small GTP-binding family protein (.1)
Potri.003G081800
278 / 6e-95
AT1G02130 391 / 9e-141
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Potri.002G138400
278 / 7e-95
AT1G02130 393 / 1e-141
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Potri.001G080400
278 / 1e-94
AT1G02130 387 / 2e-139
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Potri.014G049400
275 / 1e-93
AT1G02130 390 / 2e-140
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Potri.001G152800
270 / 1e-91
AT1G02130 382 / 2e-137
ARABIDOPSIS THALIANA RAB D2A, ARABIDOPSIS THALIANA RESPONSIVE TO ABSCISIC ACID 1B, ARABIDOPSIS RAS 5, RAS 5 (.1)
Potri.009G027900
201 / 2e-64
AT3G46060 341 / 8e-121
RAB GTPase homolog 8A (.1.2.3)
Potri.001G236100
197 / 8e-63
AT5G59840 333 / 1e-117
Ras-related small GTP-binding family protein (.1)
Potri.010G208900
197 / 1e-62
AT3G46060 349 / 5e-124
RAB GTPase homolog 8A (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00071
Ras
Ras family
Representative CDS sequence
>Lus10041117 pacid=23179689 polypeptide=Lus10041117 locus=Lus10041117.g ID=Lus10041117.BGIv1.0 annot-version=v1.0
ATGACTACTTTAAGACAAAGAAAGTGGTGGACTGGTGGGAGGAAAGCTCTTACTCTGCAGCCAGTTCCCGCTGCTTTGCTTTCTCAAAGAGGAAAGGAAC
TCAATTCCCTTCCATCTCCAGACGGAATTCATTCATTGAGAGCAAGAAACCCTATCATCTTCAATCATGGGCACCGAATACGACTATCTCTTCAAGCTTC
TGCTAATCGGCGACTCCTCCGTTGGAAAATCTTGCCTGCTGCTCCGATTTGCTGTAATTTTGACTTTCTGTTGACCCCGCTCTTCTTTCTCGTGTCTGAT
GAGTTTGGATTTAATCATCTAGGGTTTCCATTTTTGATTTTGCAGGATGATTCGTACGTTGACAGCTACATCAGTACTATAGGAGTTGATTTCAAAATCA
GAACTGTGGAGCTGGATGGAAAGACGGTCAAGCTTCAGATCTGGGATACTGCTGGTCAGGAGCGCTTTAGAACAATAACAAGCAGTTATTACCGAGGGGC
ACATGGAATCATCATTGTCTATGATGTTACTGACATGGACAGCTTCAACAATGTCAAACAATGGTTAAATGAGATTGACCGATATGCAAATGATACTGTA
TGCAAGCTTTTGGTTGGGAACAAATGCGATCTTGTTGAGAACAAAGTTGTCGATACGCAGACAGCAAAGGCGTTGGCCGATGAGCTAGGCATCCCTTTTC
TGGAGACCAGTGCCAAAGATTCAATAAATGTGGAACAAGCTTTCTTAACAATGGCTGCAGAAATTAAGAAAAAAATGGGTAATCAACCGACAGCTAGCAA
GGCGACCGGAACGGTTCAGATGAAAGGACAACCGATCCAGCAAAGCAACAACTGCTGTGGTTAA
AA sequence
>Lus10041117 pacid=23179689 polypeptide=Lus10041117 locus=Lus10041117.g ID=Lus10041117.BGIv1.0 annot-version=v1.0
MTTLRQRKWWTGGRKALTLQPVPAALLSQRGKELNSLPSPDGIHSLRARNPIIFNHGHRIRLSLQASANRRLLRWKILPAAPICCNFDFLLTPLFFLVSD
EFGFNHLGFPFLILQDDSYVDSYISTIGVDFKIRTVELDGKTVKLQIWDTAGQERFRTITSSYYRGAHGIIIVYDVTDMDSFNNVKQWLNEIDRYANDTV
CKLLVGNKCDLVENKVVDTQTAKALADELGIPFLETSAKDSINVEQAFLTMAAEIKKKMGNQPTASKATGTVQMKGQPIQQSNNCCG
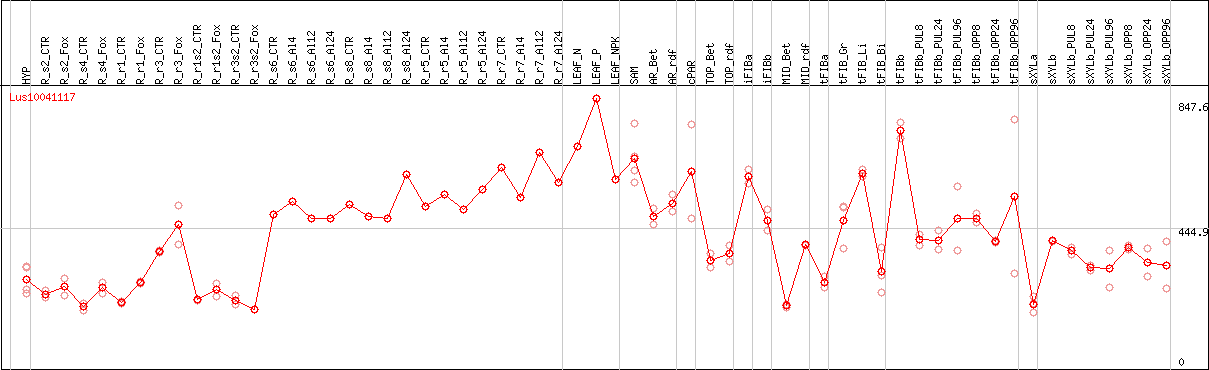
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041117 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.