External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G04190 501 / 3e-180
TPR3
tetratricopeptide repeat 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G62740 100 / 4e-23
Hop2
Hop2, stress-inducible protein, putative (.1)
AT4G12400 94 / 3e-21
Hop3
Hop3, stress-inducible protein, putative (.1.2)
AT1G12270 93 / 9e-21
Hop1
Hop1, stress-inducible protein, putative (.1)
AT3G04710 86 / 3e-18
TPR10
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
AT2G42810 79 / 4e-16
AtPP5, PP5.2, PP5, PAPP5
Arabidopsis thaliana protein phosphatase 5, protein phosphatase 5.2 (.1.2)
AT5G09420 73 / 4e-14
MTOM64, ATTOC64-V, AtmtOM64
outer membrane 64, ARABIDOPSIS THALIANA TRANSLOCON AT THE OUTER MEMBRANE OF CHLOROPLASTS 64-V, translocon at the outer membrane of chloroplasts 64-V (.1)
AT4G08320 71 / 2e-13
TPR8
tetratricopeptide repeat 8, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G53300 71 / 3e-13
TTL1
tetratricopetide-repeat thioredoxin-like 1 (.1)
AT3G17970 69 / 8e-13
ATTOC64-III
translocon at the outer membrane of chloroplasts 64-III (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021905
623 / 0
AT1G04190 515 / 0.0
tetratricopeptide repeat 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10004336
92 / 3e-20
AT1G62740 873 / 0.0
Hop2, stress-inducible protein, putative (.1)
Lus10024597
91 / 5e-20
AT1G62740 886 / 0.0
Hop2, stress-inducible protein, putative (.1)
Lus10032234
91 / 6e-20
AT1G62740 892 / 0.0
Hop2, stress-inducible protein, putative (.1)
Lus10028919
89 / 3e-19
AT1G62740 890 / 0.0
Hop2, stress-inducible protein, putative (.1)
Lus10042001
82 / 4e-17
AT3G17970 714 / 0.0
translocon at the outer membrane of chloroplasts 64-III (.1)
Lus10006559
81 / 2e-16
AT3G04710 581 / 0.0
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
Lus10009682
81 / 2e-16
AT3G17970 700 / 0.0
translocon at the outer membrane of chloroplasts 64-III (.1)
Lus10003260
80 / 2e-16
AT3G04710 587 / 0.0
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G082900
543 / 0
AT1G04190 540 / 0.0
tetratricopeptide repeat 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.008G156500
534 / 0
AT1G04190 540 / 0.0
tetratricopeptide repeat 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G119500
97 / 3e-22
AT4G12400 864 / 0.0
Hop3, stress-inducible protein, putative (.1.2)
Potri.003G113400
96 / 1e-21
AT4G12400 830 / 0.0
Hop3, stress-inducible protein, putative (.1.2)
Potri.013G042300
90 / 6e-20
AT3G04710 619 / 0.0
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
Potri.005G054800
86 / 2e-18
AT3G04710 611 / 0.0
tetratricopeptide repeat 10, ankyrin repeat family protein (.1.2.3)
Potri.012G046900
82 / 4e-17
AT3G17970 744 / 0.0
translocon at the outer membrane of chloroplasts 64-III (.1)
Potri.015G038600
81 / 1e-16
AT3G17970 747 / 0.0
translocon at the outer membrane of chloroplasts 64-III (.1)
Potri.001G205300
77 / 3e-15
AT5G09420 724 / 0.0
outer membrane 64, ARABIDOPSIS THALIANA TRANSLOCON AT THE OUTER MEMBRANE OF CHLOROPLASTS 64-V, translocon at the outer membrane of chloroplasts 64-V (.1)
Potri.014G141800
76 / 5e-15
AT2G42810 895 / 0.0
Arabidopsis thaliana protein phosphatase 5, protein phosphatase 5.2 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF00515
TPR_1
Tetratricopeptide repeat
Representative CDS sequence
>Lus10041189 pacid=23179842 polypeptide=Lus10041189 locus=Lus10041189.g ID=Lus10041189.BGIv1.0 annot-version=v1.0
ATGGCGGAAACAGCTCGAGGATCGGCTACAAATGGAACCCAGATGTCATTGAAAGATCAGGGGAACGAATTCTTCAAAGCCGGAAACTTCCTCAAAGCTG
CTGCTGTCTACACCCAGGCTATCAAGCTCGACCCTTCTAATCCGACTCTATATAGTAATCGTGCTGCTGCGTTTGTGCAACTGGTTAAGCTGAATAAAGC
ACTTGCTGATGCTGAGACCACTATCAAACTCAACCCACAGTGGGAGAAGGGTTATTTCAGGAAAGGGTGTGTATTGGAGGTCATGGAACGGTATGATGAT
GCCTTGGAATCCTTCCAAGAAGCTTTACAGTACAACCCACAAAGTGCAGAAGTATCTAGAAAGATTAAGAGGATAACTCAATTAGCAAGGGACAAAAAGA
GAGCTCAAGAGGTGGAGAGCATGAGATCAAATATTGATATCGCAAAGCACTTGAATCAGTTTAAAGCTGAAATGTCAGGGAAGCTTGAATCTGAAGAATT
GGAGAAAGAAGTGTTCTCTTTCATTGTTGATACAATGGAGACAGCTGTTAAGTCATGGCATGAAACTTCTAAAGTGGATCCAAGAGTCTTCTTTCTTTTG
GATAAGGAAAAGACACAAACTGATAAATATGCTCCAATGGTCAACATTGACAAGGCCTTTGAATCACCTCATACACACAGCAATTGCTTCTCATTCCTAA
GGCAGTATGCCGAAGAATCATACTCCAAAGCAGCTTGCTTAATTGTGGCCAAGAATATCATAGCTTACCCGCAGGTTTGGAAAGGTCAAGGATCAAGGAA
GTGGAAGCATGGCCAACTTGATGGCTTCTTCGTTCAGTTCGAGTCACCAACTTCGCGGCAATTGTGGTTTGTCGCTAGCACAACCGAAAGAGGGAAAACA
CTGTGCAGTGACCCAGAGAGCCTAGATATCAGTACACATGAAGTCATCCCGCGAATTTTCAAAGAGAAGTTGAGCACTTCCTAG
AA sequence
>Lus10041189 pacid=23179842 polypeptide=Lus10041189 locus=Lus10041189.g ID=Lus10041189.BGIv1.0 annot-version=v1.0
MAETARGSATNGTQMSLKDQGNEFFKAGNFLKAAAVYTQAIKLDPSNPTLYSNRAAAFVQLVKLNKALADAETTIKLNPQWEKGYFRKGCVLEVMERYDD
ALESFQEALQYNPQSAEVSRKIKRITQLARDKKRAQEVESMRSNIDIAKHLNQFKAEMSGKLESEELEKEVFSFIVDTMETAVKSWHETSKVDPRVFFLL
DKEKTQTDKYAPMVNIDKAFESPHTHSNCFSFLRQYAEESYSKAACLIVAKNIIAYPQVWKGQGSRKWKHGQLDGFFVQFESPTSRQLWFVASTTERGKT
LCSDPESLDISTHEVIPRIFKEKLSTS
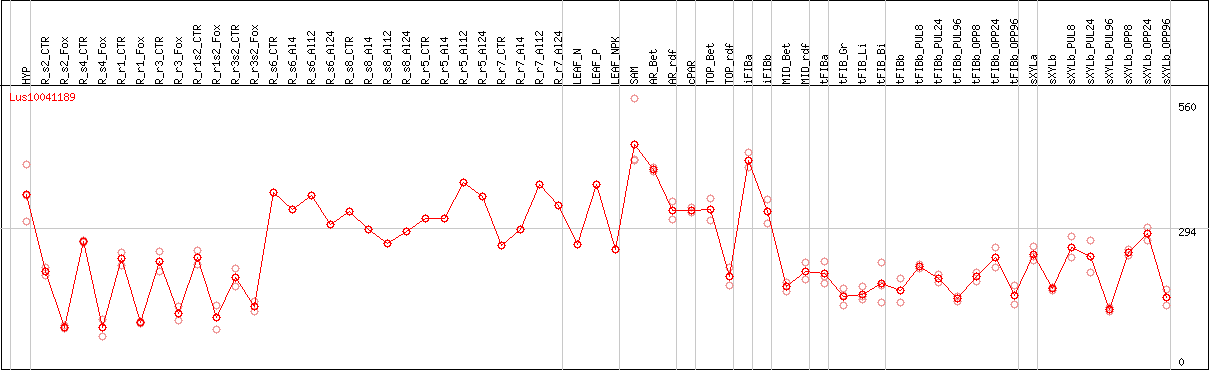
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041189 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.